Abstract
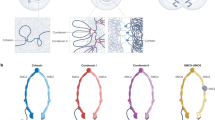
Structural maintenance of chromosome (SMC) complexes are central regulators of chromosome architecture that are essential in all domains of life. For decades, the structural biology field has been debating how these conserved protein complexes use their intricate ring-like structures to structurally organize DNA. Here, we review the contributions of single-molecule biophysical approaches to resolving the molecular mechanism of SMC protein function.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Uhlmann, F. SMC complexes: from DNA to chromosomes. Nat. Rev. Mol. Cell Biol. 17, 399–412 (2016).Excellent recent review of the current understanding of the mechanism of SMC complexes.
Nasmyth, K. & Haering, C.H. Cohesin: its roles and mechanisms. Annu. Rev. Genet. 43, 525–558 (2009).
Hirano, T. Condensins: universal organizers of chromosomes with diverse functions. Genes Dev. 26, 1659–1678 (2012).
Haering, C.H. & Gruber, S. SnapShot: SMC protein complexes part I. Cell 164, 326–326.e1 (2016).
Nolivos, S. & Sherratt, D. The bacterial chromosome: architecture and action of bacterial SMC and SMC-like complexes. FEMS Microbiol. Rev. 38, 380–392 (2014).
Britton, R.A., Lin, D.C. & Grossman, A.D. Characterization of a prokaryotic SMC protein involved in chromosome partitioning. Genes Dev. 12, 1254–1259 (1998).
Wang, Q., Mordukhova, E.A., Edwards, A.L. & Rybenkov, V.V. Chromosome condensation in the absence of the non-SMC subunits of MukBEF. J. Bacteriol. 188, 4431–4441 (2006).
Gruber, S. et al. Interlinked sister chromosomes arise in the absence of condensin during fast replication in B. subtilis. Curr. Biol. 24, 293–298 (2014).
Danilova, O., Reyes-Lamothe, R., Pinskaya, M., Sherratt, D. & Possoz, C. MukB colocalizes with the oriC region and is required for organization of the two Escherichia coli chromosome arms into separate cell halves. Mol. Microbiol. 65, 1485–1492 (2007).
Uhlmann, F., Lottspeich, F. & Nasmyth, K. Sister-chromatid separation at anaphase onset is promoted by cleavage of the cohesin subunit Scc1. Nature 400, 37–42 (1999).
Waizenegger, I.C., Hauf, S., Meinke, A. & Peters, J.M. Two distinct pathways remove mammalian cohesin from chromosome arms in prophase and from centromeres in anaphase. Cell 103, 399–410 (2000).
Merkenschlager, M. & Nora, E.P. CTCF and cohesin in genome folding and transcriptional gene regulation. Annu. Rev. Genomics Hum. Genet. 17, 17–43 (2016).
Ono, T. et al. Differential contributions of condensin I and condensin II to mitotic chromosome architecture in vertebrate cells. Cell 115, 109–121 (2003).
Frosi, Y. & Haering, C.H. Control of chromosome interactions by condensin complexes. Curr. Opin. Cell Biol. 34, 94–100 (2015).
Meyer, B.J. Targeting X chromosomes for repression. Curr. Opin. Genet. Dev. 20, 179–189 (2010).
Lehmann, A.R. et al. The rad18 gene of Schizosaccharomyces pombe defines a new subgroup of the SMC superfamily involved in DNA repair. Mol. Cell. Biol. 15, 7067–7080 (1995).
De Piccoli, G. et al. Smc5-Smc6 mediate DNA double-strand-break repair by promoting sister-chromatid recombination. Nat. Cell Biol. 8, 1032–1034 (2006).
Lindroos, H.B. et al. Chromosomal association of the Smc5/6 complex reveals that it functions in differently regulated pathways. Mol. Cell 22, 755–767 (2006).
Jeppsson, K. et al. The chromosomal association of the Smc5/6 complex depends on cohesion and predicts the level of sister chromatid entanglement. PLoS Genet. 10, e1004680 (2014).
Dekker, J. & Mirny, L. The 3D genome as moderator of chromosomal communication. Cell 164, 1110–1121 (2016).
Haering, C.H., Farcas, A.-M., Arumugam, P., Metson, J. & Nasmyth, K. The cohesin ring concatenates sister DNA molecules. Nature 454, 297–301 (2008).
Cuylen, S., Metz, J. & Haering, C.H. Condensin structures chromosomal DNA through topological links. Nat. Struct. Mol. Biol. 18, 894–901 (2011).
Kanno, T., Berta, D.G. & Sjögren, C. The Smc5/6 complex is an ATP-dependent intermolecular DNA linker. Cell Rep. 12, 1471–1482 (2015).
Wilhelm, L. et al. SMC condensin entraps chromosomal DNA by an ATP hydrolysis dependent loading mechanism in Bacillus subtilis. eLife 4, 11202–11212 (2015).
Cuylen, S., Metz, J., Hruby, A. & Haering, C.H. Entrapment of chromosomes by condensin rings prevents their breakage during cytokinesis. Dev. Cell 27, 469–478 (2013).
Thadani, R., Uhlmann, F. & Heeger, S. Condensin, chromatin crossbarring and chromosome condensation. Curr. Biol. 22, R1012–R1021 (2012).
Cheng, T.M.K. et al. A simple biophysical model emulates budding yeast chromosome condensation. eLife 4, e05565 (2015).
Nasmyth, K. Disseminating the genome: joining, resolving, and separating sister chromatids during mitosis and meiosis. Annu. Rev. Genet. 35, 673–745 (2001).
Alipour, E. & Marko, J.F. Self-organization of domain structures by DNA-loop-extruding enzymes. Nucleic Acids Res. 40, 11202–11212 (2012).
Dolgin, E. DNA's secret weapon against knots and tangles. Nature 544, 284–286 (2017).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
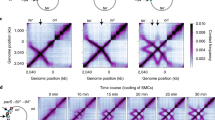
Wang, X., Brandão, H.B., Le, T.B.K., Laub, M.T. & Rudner, D.Z. Bacillus subtilis SMC complexes juxtapose chromosome arms as they travel from origin to terminus. Science 355, 524–527 (2017).
Sanborn, A.L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl. Acad. Sci. USA 112, E6456–E6465 (2015).
Goloborodko, A., Marko, J.F. & Mirny, L.A. Chromosome compaction by active loop extrusion. Biophys. J. 110, 2162–2168 (2016).
Goloborodko, A., Imakaev, M.V., Marko, J.F. & Mirny, L. Compaction and segregation of sister chromatids via active loop extrusion. eLife 5, e14864 (2016).
Diebold-Durand, M.-L. et al. Structure of full-length SMC and rearrangements required for chromosome organization. Mol. Cell 67, 334–347.e5 (2017).
Kimura, K. & Hirano, T. ATP-dependent positive supercoiling of DNA by 13S condensin: a biochemical implication for chromosome condensation. Cell 90, 625–634 (1997).
Kimura, K., Rybenkov, V.V., Crisona, N.J., Hirano, T. & Cozzarelli, N.R. 13S condensin actively reconfigures DNA by introducing global positive writhe: implications for chromosome condensation. Cell 98, 239–248 (1999).
Bazett-Jones, D.P., Kimura, K. & Hirano, T. Efficient supercoiling of DNA by a single condensin complex as revealed by electron spectroscopic imaging. Mol. Cell 9, 1183–1190 (2002).
Kim, H. & Loparo, J.J. Multistep assembly of DNA condensation clusters by SMC. Nat. Commun. 7, 10200 (2016).
Ha, T. Single-molecule methods leap ahead. Nat. Methods 11, 1015–1018 (2014).
Murayama, Y. & Uhlmann, F. Biochemical reconstitution of topological DNA binding by the cohesin ring. Nature 505, 367–371 (2014).
Tanaka, T., Fuchs, J., Loidl, J. & Nasmyth, K. Cohesin ensures bipolar attachment of microtubules to sister centromeres and resists their precocious separation. Nat. Cell Biol. 2, 492–499 (2000).
Ganji, M., Kim, S.H., van der Torre, J., Abbondanzieri, E. & Dekker, C. Intercalation-based single-molecule fluorescence assay to study DNA supercoil dynamics. Nano Lett. 16, 4699–4707 (2016).
Greene, E.C., Wind, S., Fazio, T., Gorman, J. & Visnapuu, M.L. DNA curtains for high-throughput single-molecule optical imaging. Methods Enzymol. 472, 293–315 (2010).
Juette, M.F. et al. The bright future of single-molecule fluorescence imaging. Curr. Opin. Chem. Biol. 20, 103–111 (2014).
Gligoris, T. & Löwe, J. Structural insights into ring formation of cohesin and related Smc complexes. Trends Cell Biol. 26, 680–693 (2016).
Bai, X.C., McMullan, G. & Scheres, S.H. How cryo-EM is revolutionizing structural biology. Trends Biochem. Sci. 40, 49–57 (2015).
Ando, T. et al. A high-speed atomic force microscope for studying biological macromolecules. Proc. Natl. Acad. Sci. USA 98, 12468–12472 (2001).
Katan, A.J. & Dekker, C. High-speed AFM reveals the dynamics of single biomolecules at the nanometer scale. Cell 147, 979–982 (2011).
Niki, H. et al. E.coli MukB protein involved in chromosome partition forms a homodimer with a rod-and-hinge structure having DNA binding and ATP/GTP binding activities. EMBO J. 11, 5101–5109 (1992).The first electron micrographs of SMC proteins (MukB) showing the globular ATPase domains and coiled coils.
Melby, T.E., Ciampaglio, C.N., Briscoe, G. & Erickson, H.P. The symmetrical structure of structural maintenance of chromosomes (SMC) and MukB proteins: long, antiparallel coiled coils, folded at a flexible hinge. J. Cell Biol. 142, 1595–1604 (1998).
Matoba, K., Yamazoe, M., Mayanagi, K., Morikawa, K. & Hiraga, S. Comparison of MukB homodimer versus MukBEF complex molecular architectures by electron microscopy reveals a higher-order multimerization. Biochem. Biophys. Res. Commun. 333, 694–702 (2005).
Soh, Y.M. et al. Molecular basis for SMC rod formation and its dissolution upon DNA binding. Mol. Cell 57, 290–303 (2015).
Mascarenhas, J. et al. Dynamic assembly, localization and proteolysis of the Bacillus subtilis SMC complex. BMC Cell Biol. 6, 28 (2005).
Hirano, M., Anderson, D.E., Erickson, H.P. & Hirano, T. Bimodal activation of SMC ATPase by intra- and inter-molecular interactions. EMBO J. 20, 3238–3250 (2001).
Fuentes-Perez, M.E., Gwynn, E.J., Dillingham, M.S. & Moreno-Herrero, F. Using DNA as a fiducial marker to study SMC complex interactions with the atomic force microscope. Biophys. J. 102, 839–848 (2012).
Kamada, K., Miyata, M. & Hirano, T. Molecular basis of SMC ATPase activation: role of internal structural changes of the regulatory subcomplex ScpAB. Structure 21, 581–594 (2013).
Kamada, K., Su'etsugu, M., Takada, H., Miyata, M. & Hirano, T. Overall shapes of the SMC-ScpAB complex are determined by balance between constraint and relaxation of its structural parts. Structure 25, 603–616 e4 (2017).
Bahng, S., Hayama, R. & Marians, K.J. MukB-mediated catenation of DNA is ATP and MukEF independent. J. Biol. Chem. 291, 23999–24008 (2016).
Badrinarayanan, A., Reyes-Lamothe, R., Uphoff, S., Leake, M.C. & Sherratt, D. J. In vivo architecture and action of bacterial structural maintenance of chromosome proteins. Science 338, 528–531(2012).
Anderson, D.E., Losada, A., Erickson, H.P. & Hirano, T. Condensin and cohesin display different arm conformations with characteristic hinge angles. J. Cell Biol. 156, 419–424 (2002).
Haering, C.H., Löwe, J., Hochwagen, A. & Nasmyth, K. Molecular architecture of SMC proteins and the yeast cohesin complex. Mol. Cell 9, 773–788 (2002).
Elbatsh, A.M.O. et al. Cohesin releases DNA through asymmetric ATPase-driven ring opening. Mol. Cell 61, 575–588 (2016).
Huis in 't Veld, P.J. et al. Characterization of a DNA exit gate in the human cohesin ring. Science 346, 968–972 (2014).
Kulemzina, I. et al. A reversible association between Smc coiled coils is regulated by lysine acetylation and is required for cohesin association with the DNA. Mol. Cell 63, 1044–1054 (2016).
Sun, M., Nishino, T. & Marko, J.F. The SMC1-SMC3 cohesin heterodimer structures DNA through supercoiling-dependent loop formation. Nucleic Acids Res. 41, 6149–6160 (2013).
Bazett-Jones, D.P. & Hendzel, M.J. Electron spectroscopic imaging of chromatin. Methods 17, 188–200 (1999).
Eeftens, J.M. et al. Condensin Smc2-Smc4 dimers are flexible and dynamic. Cell Rep. 14, 1813–1818 (2016).High-speed AFM study showing that condensin dimers dynamically adopt different conformations.
Gruber, S. et al. Evidence that loading of cohesin onto chromosomes involves opening of its SMC hinge. Cell 127, 523–537 (2006).
Nasmyth, K. Cohesin: a catenase with separate entry and exit gates? Nat. Cell Biol. 13, 1170–1177 (2011).
De Vlaminck, I. & Dekker, C. Recent advances in magnetic tweezers. Annu. Rev. Biophys. 41, 453–472 (2012).
Strick, T.R., Kawaguchi, T. & Hirano, T. Real-time detection of single-molecule DNA compaction by condensin I. Curr. Biol. 14, 874–880 (2004.)Pioneering magnetic-tweezers study showing that condensin can compact DNA.
Eeftens, J.M., Bisht, S., Kerssemakers, J., Haering, C. & Dekker, C. Real-time detection of condensin-driven DNA compaction reveals a multistep binding mechanism. bioRxiv http://www.doi.org/10.1101/149138 (2017).
Cui, Y., Petrushenko, Z.M. & Rybenkov, V.V. MukB acts as a macromolecular clamp in DNA condensation. Nat. Struct. Mol. Biol. 15, 411–418 (2008).
Petrushenko, Z.M., Cui, Y., She, W. & Rybenkov, V.V. Mechanics of DNA bridging by bacterial condensin MukBEF in vitro and in singulo. EMBO J. 29, 1126–1135 (2010).
Pyle, A.M. Translocation and unwinding mechanisms of RNA and DNA helicases. Annu. Rev. Biophys. 37, 317–336 (2008).
Singleton, M.R., Dillingham, M.S. & Wigley, D.B. Structure and mechanism of helicases and nucleic acid translocases. Annu. Rev. Biochem. 76, 23–50 (2007).
Yang, W. Lessons learned from UvrD helicase: mechanism for directional movement. Annu. Rev. Biophys. 39, 367–385 (2010).
Seidel, R., Bloom, J.G., Dekker, C. & Szczelkun, M.D. Motor step size and ATP coupling efficiency of the dsDNA translocase EcoR124I. EMBO J. 27, 1388–1398 (2008).
Stigler, J., Çamdere, G.Ö., Koshland, D.E. & Greene, E.C. Single-molecule imaging reveals a collapsed conformational state for DNA-bound cohesin. Cell Rep. 15, 988–998 (2016).One of three studies demonstrating diffusion of cohesin on stretched DNA and their ability to move across roadblocks.
Davidson, I.F. et al. Rapid movement and transcriptional re-localization of human cohesin on DNA. EMBO J. 35, 2671–2685 (2016).
Ladurner, R. et al. Cohesin's ATPase activity couples cohesin loading onto DNA with Smc3 acetylation. Curr. Biol. 24, 2228–2237 (2014).
Murayama, Y. & Uhlmann, F. Chromosome segregation: how to open cohesin without cutting the ring? EMBO J. 32, 614–616 (2013).
Kanke, M., Tahara, E., Huis In't Veld, P.J. & Nishiyama, T. Cohesin acetylation and Wapl-Pds5 oppositely regulate translocation of cohesin along DNA. EMBO J. 35, 2686–2698 (2016).
Terakawa, T. et al. The condensin complex is a mechanochemical motor that translocates along DNA. Science http://dx.doi.org/10.1126/eaan6516 (2017).
Mc Intyre, J. et al. In vivo analysis of cohesin architecture using FRET in the budding yeast Saccharomyces cerevisiae. EMBO J. 26, 3783–3793 (2007).
Shintomi, K., Takahashi, T.S. & Hirano, T. Reconstitution of mitotic chromatids with a minimum set of purified factors. Nat. Cell Biol. 17, 1014–1023 (2015).
Gruber, S. & Errington, J. Recruitment of condensin to replication origin regions by ParB/SpoOJ promotes chromosome segregation in B. subtilis. Cell 137, 685–696 (2009).
Sullivan, N.L., Marquis, K.A. & Rudner, D.Z. Recruitment of SMC by ParB-parS organizes the origin region and promotes efficient chromosome segregation. Cell 137, 697–707 (2009).
Schalbetter, S.A. et al. SMC complexes differentially compact mitotic chromosomes according to genomic context. Nat. Cell Biol. 19, 1071–1080 (2017).
Acknowledgements
We thank C. Haering, S. Bisht, A. Katan, J. Ryu, S. Pud, Y. Kabiri, and A. Japaridze for discussions. This work was funded by the ERC Advanced Grant SynDiv (ERC-ADG-2014 669598) and the Netherlands Organization for Scientific Research (NWO/OCW) as part of the Frontiers of Nanoscience program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Eeftens, J., Dekker, C. Catching DNA with hoops—biophysical approaches to clarify the mechanism of SMC proteins. Nat Struct Mol Biol 24, 1012–1020 (2017). https://doi.org/10.1038/nsmb.3507
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3507
This article is cited by
-
The condensin holocomplex cycles dynamically between open and collapsed states
Nature Structural & Molecular Biology (2020)