Abstract
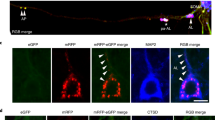
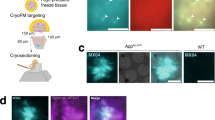
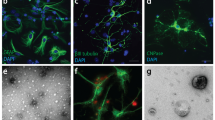
Alzheimer’s disease (AD) is characterized by the presence of neuritic plaques in which dystrophic neurites (DNs) are typical constituents. We recently showed that DNs labeled by antibodies to the tubular endoplasmic reticulum (ER) protein reticulon-3 (RTN3) are enriched with clustered tubular ER. However, multi-vesicle bodies are also found in DNs, suggesting that different populations of DNs exist in brains of AD patients. To understand how different DNs evolve to surround core amyloid plaques, we monitored the growth of DNs in AD mouse brains (5xFAD and APP/PS1ΔE9 mice) by multiple approaches, including two-dimensional and three-dimensional (3D) electron microscopy (EM). We discovered that a pre-autophagosome protein ATG9A was enriched in DNs when a plaque was just beginning to develop. ATG9A-positive DNs were often closer to the core amyloid plaque, whereas RTN3 immunoreactive DNs were mostly located in the outer layers of ATG9A-positive DNs. Proteins such as RAB7 and LC3 appeared in DNs at later stages during plaque growth, likely accumulated as a part of large autophagy vesicles, and were distributed relatively furthest from the core amyloid plaque. Reconstructing the 3D structure of different morphologies of DNs revealed that DNs in AD mouse brains were constituted in three layers that are distinct by enriching different types of vesicles, as validated by immune-EM methods. Collectively, our results provide the first evidence that DNs evolve from dysfunctions of pre-autophagosomes, tubular ER, mature autophagosomes, and the ubiquitin proteasome system during plaque growth.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Braak H, Braak E. Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol. 1991;82:239–59.
Price JL, Morris JC. Tangles and plaques in nondemented aging and “preclinical” Alzheimer’s disease. Ann Neurol. 1999;45:358–68.
Selkoe DJ. Alzheimer’s disease is a synaptic failure. Science. 2002;298:789–91. https://doi.org/10.1126/science.1074069
Trojanowski JQ, Lee VM. Brain degeneration linked to “fatal attractions” of proteins in Alzheimer’s disease and related disorders. J Alzheimers Dis. 2001;3:117–9.
Fukumoto H, et al. Amyloid beta protein deposition in normal aging has the same characteristics as that in Alzheimer’s disease. Predominance of A beta 42(43) and association of A beta 40 with cored plaques. Am J Pathol. 1996;148:259–65.
Joachim CL, Morris JH, Selkoe DJ. Diffuse senile plaques occur commonly in the cerebellum in Alzheimer’s disease. Am J Pathol. 1989;135:309–19.
Dickson DW, et al. Ubiquitin immunoelectron microscopy of dystrophic neurites in cerebellar senile plaques of Alzheimer’s disease. Acta Neuropathol. 1990;79:486–93.
Lenders MB, et al. Dystrophic neuropeptidergic neurites in senile plaques of Alzheimer’s disease precede formation of paired helical filaments. Acta Neurol Belg. 1989;89:279–85.
Onorato M, et al. Alteration of neuritic cytoarchitecture in Alzheimer disease. Prog Clin Biol Res. 1989;317:781–9.
Dickson TC, King CE, McCormack GH, Vickers JC. Neurochemical diversity of dystrophic neurites in the early and late stages of Alzheimer’s disease. Exp Neurol. 1999;156:100–10. S0014-4886(98)97010-8 [pii] https://doi.org/10.1006/exnr.1998.7010
Masliah E, et al. An antibody against phosphorylated neurofilaments identifies a subset of damaged association axons in Alzheimer’s disease. Am J Pathol. 1993;142:871–82.
Vickers JC, et al. Dystrophic neurite formation associated with age-related beta amyloid deposition in the neocortex: clues to the genesis of neurofibrillary pathology. Exp Neurol. 1996;141:1–11. S0014-4886(96)90133-8 [pii] https://doi.org/10.1006/exnr.1996.0133
Hu X, et al. Transgenic mice overexpressing reticulon 3 develop neuritic abnormalities. EMBO J. 2007;26:2755–67. doi:7601707 [pii] https://doi.org/10.1038/sj.emboj.7601707
Sharoar MG, et al. Dysfunctional tubular endoplasmic reticulum constitutes a pathological feature of Alzheimer’s disease. Mol Psychiatry. 2015. https://doi.org/10.1038/mp.2015.181 mp2015181 [pii].
Friedman JR, Webster BM, Mastronarde DN, Verhey KJ, Voeltz GK. ER sliding dynamics and ER-mitochondrial contacts occur on acetylated microtubules. J Cell Biol. 2010;190:363–75. https://doi.org/10.1083/jcb.200911024 jcb.200911024 [pii]
Noda T. Autophagy in the context of the cellular membrane-trafficking system: the enigma of Atg9 vesicles. Biochem Soc Trans. 2017;45:1323–31. https://doi.org/10.1042/BST20170128BST20170128 [pii]
Bordi M, et al. Autophagy flux in CA1 neurons of Alzheimer hippocampus: increased induction overburdens failing lysosomes to propel neuritic dystrophy. Autophagy. 2016;12:2467–83. https://doi.org/10.1080/15548627.2016.1239003
Lee S, Sato Y, Nixon RA. Lysosomal proteolysis inhibition selectively disrupts axonal transport of degradative organelles and causes an Alzheimer’s-like axonal dystrophy. J Neurosci. 2011;31:7817–30. https://doi.org/10.1073/JNEUROSCI.6412-10.201131/21/7817 [pii]
Klionsky DJ, et al. A unified nomenclature for yeast autophagy-related genes. Dev Cell. 2003;5:539–45. S1534-5807(03)00296-X [pii]
Mizushima N. Autophagy: process and function. Genes Dev. 2007;21:2861–73. doi:21/22/2861 [pii] https://doi.org/10.1101/gad.1599207
Shintani T, Klionsky DJ. Autophagy in health and disease: a double-edged sword. Science. 2004;306:990–5. doi:306/5698/990 [pii] https://doi.org/10.1126/science.1099993
Klionsky DJ. Autophagy: from phenomenology to molecular understanding in less than a decade. Nat Rev Mol Cell Biol. 2007;8:931–7. doi:nrm2245 [pii] https://doi.org/10.1038/nrm2245
Sadleir KR, et al. Presynaptic dystrophic neurites surrounding amyloid plaques are sites of microtubule disruption, BACE1 elevation, and increased Abeta generation in Alzheimer’s disease. Acta Neuropathol. 2016;132:235–56. https://doi.org/10.1007/s00401-016-1558-910.1007/s00401-016-1558-9 [pii]
Tammineni P, Cai Q. Defective retrograde transport impairs autophagic clearance in Alzheimer disease neurons. Autophagy. 2017;13:982–4. https://doi.org/10.1080/15548627.2017.1291114
Sanchez-Varo R, et al. Abnormal accumulation of autophagic vesicles correlates with axonal and synaptic pathology in young Alzheimer’s mice hippocampus. Acta Neuropathol. 2012;123:53–70. https://doi.org/10.1007/s00401-011-0896-x
Gowrishankar S, et al. Massive accumulation of luminal protease-deficient axonal lysosomes at Alzheimer’s disease amyloid plaques. Proc Natl Acad Sci USA. 2015;112:E3699–3708. https://doi.org/10.1073/10.1073/pnas.15103291121510329112 [pii]
Nixon RA, et al. Extensive involvement of autophagy in Alzheimer disease: an immuno-electron microscopy study. J Neuropathol Exp Neurol. 2005;64:113–22.
Nixon RA. Autophagy, amyloidogenesis and Alzheimer disease. J Cell Sci. 2007;120:4081–91. doi:120/23/4081 [pii] https://doi.org/10.1242/jcs.019265
He W, et al. Reticulon family members modulate BACE1 activity and amyloid-beta peptide generation. Nat Med. 2004;10:959–65. https://doi.org/10.1038/nm1088nm1088 [pii]
Shi Q, et al. Impact of RTN3 deficiency on expression of BACE1 and amyloid deposition. J Neurosci. 2014;34:13954–62. https://doi.org/10.1073/JNEUROSCI.1588-14.201434/42/13954 [pii]
Schindelin J, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82. https://doi.org/10.1038/nmeth.2019nmeth.2019 [pii]
Fiala JC, Harris KM. Extending unbiased stereology of brain ultrastructure to three-dimensional volumes. J Am Med Inform Assoc. 2001;8:1–16.
Parzych KR, Klionsky DJ. An overview of autophagy: morphology, mechanism, and regulation. Antioxid Redox Signal. 2014;20:460–73. https://doi.org/10.1089/ars.2013.5371
Mari M, et al. An Atg9-containing compartment that functions in the early steps of autophagosome biogenesis. J Cell Biol. 2010;190:1005–22. https://doi.org/10.1083/jcb.200912089jcb.200912089 [pii]
Yamamoto H, et al. Atg9 vesicles are an important membrane source during early steps of autophagosome formation. J Cell Biol. 2012;198:219–33. https://doi.org/10.1083/jcb.201202061jcb.201202061 [pii]
Karanasios E, et al. Autophagy initiation by ULK complex assembly on ER tubulovesicular regions marked by ATG9 vesicles. Nat Commun. 2016;7:12420. https://doi.org/10.1038/ncomms12420ncomms12420 [pii]
Jager S, et al. Role for Rab7 in maturation of late autophagic vacuoles. J Cell Sci. 2004;117:4837–48. https://doi.org/10.1242/jcs.01370jcs.01370 [pii]
Borchelt DR, et al. Familial Alzheimer’s disease-linked presenilin 1 variants elevate Abeta1-42/1-40 ratio in vitro and in vivo. Neuron. 1996;17:1005–13. doi:S0896-6273(00)80230-5 [pii]
Feng Y, Klionsky DJ. Autophagic membrane delivery through ATG9. Cell Res. 2017;27:161–2. https://doi.org/10.1038/cr.2017.4cr20174 [pii]
Orsi A, et al. Dynamic and transient interactions of Atg9 with autophagosomes, but not membrane integration, are required for autophagy. Mol Biol Cell. 2012;23:1860–73. https://doi.org/10.1091/mbc.E11-09-0746mbc.E11-09-0746 [pii]
Noda T, et al. Apg9p/Cvt7p is an integral membrane protein required for transport vesicle formation in the Cvt and autophagy pathways. J Cell Biol. 2000;148:465–80.
Yamada T, et al. Endothelial nitric-oxide synthase antisense (NOS3AS) gene encodes an autophagy-related protein (APG9-like2) highly expressed in trophoblast. J Biol Chem. 2005;280:18283–90. doi:M413957200 [pii] https://doi.org/10.1074/jbc.M413957200
Reggiori F, Tooze SA. Autophagy regulation through Atg9 traffic. J Cell Biol. 2012;198:151–3. https://doi.org/10.1083/jcb.201206119jcb.201206119 [pii]
Young AR, et al. Starvation and ULK1-dependent cycling of mammalian Atg9 between the TGN and endosomes. J Cell Sci. 2006;119:3888–3900. doi:jcs.03172 [pii] https://doi.org/10.1242/jcs.03172
Imai K, et al. Atg9A trafficking through the recycling endosomes is required for autophagosome formation. J Cell Sci. 2016;129:3781–91. jcs.196196 [pii] https://doi.org/10.1242/jcs.196196
Rao A, Simmons D, Sorkin A. Differential subcellular distribution of endosomal compartments and the dopamine transporter in dopaminergic neurons. Mol Cell Neurosci. 2011;46:148–58. https://doi.org/10.1016/j.mcn.2010.08.016S1044-7431(10)00207-1 [pii]
Wang ZX, Tan L, Yu JT. Axonal transport defects in Alzheimer’s disease. Mol Neurobiol. 2015;51:1309–21. https://doi.org/10.1007/s12035-014-8810-x
Morfini GA, et al. Axonal transport defects in neurodegenerative diseases. J Neurosci. 2009;29:12776–86. https://doi.org/10.1073/JNEUROSCI.3463-09.200929/41/12776 [pii]
Roy S, Zhang B, Lee VM, Trojanowski JQ. Axonal transport defects: a common theme in neurodegenerative diseases. Acta Neuropathol. 2005;109:5–13. https://doi.org/10.1007/s00401-004-0952-x
Reggiori F, et al. Early stages of the secretory pathway, but not endosomes, are required for Cvt vesicle and autophagosome assembly in Saccharomyces cerevisiae. Mol Biol Cell. 2004;15:2189–204. https://doi.org/10.1091/mbc.e03-07-0479E03-07-0479 [pii]
Zoppino FC, Militello RD, Slavin I, Alvarez C, Colombo MI. Autophagosome formation depends on the small GTPase Rab1 and functional ER exit sites. Traffic. 2010;11:1246–61. https://doi.org/10.1111/j.1600-0854.2010.01086.xTRA1086 [pii]
Bader CA, Shandala T, Ng YS, Johnson IR, Brooks DA. Atg9 is required for intraluminal vesicles in amphisomes and autolysosomes. Biol Open. 2015;4:1345–55. https://doi.org/10.1242/bio.013979bio.013979 [pii]
Yamaguchi J, et al. Atg9a deficiency causes axon-specific lesions including neuronal circuit dysgenesis. Autophagy. 2018;14:764–77. https://doi.org/10.1080/15548627.2017.1314897
Acknowledgements
We thank Dr. Graham Kidd (Cleveland Clinic Lerner Research Institute) for capturing 3D EM images and Dr. Maya Yankova (UConn Health EM core facility) for immuno-EM. This work is partially supported by National Institute of Health (NIH) grants to RY (AG025493, NS074256, RFAG058261, and AG046929), and an award from the Alzheimer’s Association to MGS (AARF-17–504724).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sharoar, M.G., Hu, X., Ma, XM. et al. Sequential formation of different layers of dystrophic neurites in Alzheimer’s brains. Mol Psychiatry 24, 1369–1382 (2019). https://doi.org/10.1038/s41380-019-0396-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-019-0396-2
This article is cited by
-
Integrated proteomics reveals autophagy landscape and an autophagy receptor controlling PKA-RI complex homeostasis in neurons
Nature Communications (2024)
-
Neuritic Plaques — Gateways to Understanding Alzheimer’s Disease
Molecular Neurobiology (2024)
-
Transformation of non-neuritic into neuritic plaques during AD progression drives cortical spread of tau pathology via regenerative failure
Acta Neuropathologica Communications (2023)
-
Attenuation of Alzheimer’s brain pathology in 5XFAD mice by PTH1-34, a peptide of parathyroid hormone
Alzheimer's Research & Therapy (2023)
-
Abnormal accumulation of extracellular vesicles in hippocampal dystrophic axons and regulation by the primary cilia in Alzheimer’s disease
Acta Neuropathologica Communications (2023)