Abstract
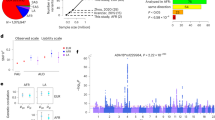
Different substance dependences have common effects on reward pathway and molecular adaptations, however little is known regarding their shared genetic factors. We aimed to identify the risk genetic variants that are shared for substance dependence (SD). First, promising genome-wide significant loci were identified from 3296 patients (521 alcoholic/1026 heroin/1749 methamphetamine) vs 2859 healthy controls and independently replicated using 1954 patients vs 1904 controls. Second, the functional effects of promising variants on gene expression, addiction characteristics, brain structure (gray and white matter), and addiction behaviors in addiction animal models (chronic administration and self-administration) were assessed. In addition, we assessed the genetic correlation among the three SDs using LD score regression. We identified and replicated three novel loci that were associated with the common risk of heroin, methamphetamine addiction, and alcoholism: ANKS1B rs2133896 (Pmeta = 3.60 × 10−9), AGBL4 rs147247472 (Pmeta = 3.40 × 10−12), and CTNNA2 rs10196867 (Pmeta = 4.73 × 10−9). Rs2133896 in ANKS1B was associated with ANKS1B gene expression and had effects on gray matter of the left calcarine and white matter of the right superior longitudinal fasciculus in heroin dependence. Overexpression of anks1b gene in the ventral tegmental area decreased addiction vulnerability for heroin and methamphetamine in self-administration rat models. Our findings could shed light on the root cause for substance dependence and will be helpful for the development of cost-effective prevention strategies for general addiction disorders.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
United Nations publication. World Drug Report. 2018; https://www.unodc.org/wdr2018/.
Vanyukov MM, Tarter RE, Kirisci L, Kirillova GP, Maher BS, Clark DB. Liability to substance use disorders: Common mechanisms and manifestations. Neurosci Biobehav Rev. 2003;27:507–15.
Nestler EJ. Is there a common molecular pathway for addiction? Nat Neurosci. 2005;8:1445–9.
Krueger RF, Hicks BM, Patrick CJ, Carlson SR, Iacono WG, McGue M. Etiologic connections among substance dependence, antisocial behavior, and personality: Modeling the externalizing spectrum. J Abnorm Psychol. 2002;111:411–24.
Uhl GR. Molecular genetic underpinnings of human substance abuse vulnerability: likely contributions to understanding addiction as a mnemonic process. Neuropharmacology. 2004;47(Suppl 1):140–7.
Palmer RHC, Brick L, Nugent NR, Bidwell LC, McGeary JE, Knopik VS, et al. Examining the role of common genetic variants on alcohol, tobacco, cannabis and illicit drug dependence: genetics of vulnerability to drug dependence. Addiction. 2015;110:530–7.
Agrawal A, Neale MC, Prescott CA, Kendler KS. Cannabis and other illicit drugs: comorbid use and abuse/dependence in males and females (vol 34, pg 217, 2004). Behav Genet. 2004;34:557–557.
Kendler KS, Jacobson KC, Prescott CA, Neale MC. Specificity of genetic and environmental risk factors for use and abuse/dependence of cannabis, cocaine, hallucinogens, sedatives, stimulants, and opiates in male twins. Am J Psychiatry. 2003;160:687–95.
Buhler KM, Gine E, Echeverry-Alzate V, Calleja-Conde J, de Fonseca FR, Lopez-Moreno JA. Common single nucleotide variants underlying drug addiction: more than a decade of research. Addict Biol. 2015;20:845–71.
Uhl GR, Drgon T, Johnson C, Fatusin OO, Liu QR, Contoreggi C, et al. “Higher order” addiction molecular genetics: Convergent data from genome-wide association in humans and mice. Biochem Pharmacol. 2008;75:98–111.
Li MD, Burmeister M. New insights into the genetics of addiction. Nat Rev Genet. 2009;10:225–31.
Reyes-Gibby CC, Yuan C, Wang J, Yeung SCJ, Shete S. Gene network analysis shows immune-signaling and ERK1/2 as novel genetic markers for multiple addiction phenotypes: alcohol, smoking and opioid addiction. Bmc Syst Biol. 2015;9:25.
Bulik-Sullivan BK, Loh PR, Finucane HK, Ripke S, Yang J. Schizophrenia Working Group of the Psychiatric Genomics C et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet. 2015;47:291–5.
Liu M, Jiang Y, Wedow R, Li Y, Brazel DM, Chen F, et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat Genet. 2019;51:237–44.
Koob GF, Volkow ND. Neurocircuitry of addiction. Neuropsychopharmacology. 2010;35:217–38.
Xue Y-X, Wu P, Shi HS, Xue LF, Chen C, Zhu WL, et al. A memory retrieval-extinction procedure to prevent drug craving and relapse. Science. 2012;336:241–5.
Zhang Y, Xue Y, Meng S, Luo Y, Liang J, Li J, et al. Inhibition of lactate transport erases drug memory and prevents drug relapse. Biol Psychiatry. 2016;79:928–39.
Hoseth EZ, Krull F, Dieset I, Morch RH, Hope S, Gardsjord ES, et al. Exploring the Wnt signaling pathway in schizophrenia and bipolar disorder. Transl Psychiatry. 2018;8:55.
Grupp LA. An investigation of intravenous ethanol self-administration in rats using a fixed-ratio schedule of reinforcement. Physiol Psychol. 1981;9:359–63.
Crowley TJ. The reinforcers for drug abuse: why people take drugs. Compr Psychiatry. 1972;13:51–62.
Tsuang MT, Lyons MJ, Meyer JM, Doyle T, Eisen SA, Goldberg J, et al. Co-occurrence of abuse of different drugs in men: the role of drug-specific and shared vulnerabilities. Arch Gen Psychiatry. 1998;55:967–72.
Ghersi E, Vito P, Lopez P, Abdallah M, D’Adamio L. The intracellular localization of amyloid beta protein precursor (A beta PP) intracellular domain associated protein-1 (AIDA-1) is regulated by A beta PP and alternative splicing. J Alzheimers Dis. 2004;6:67–78.
Jordan BA, Fernholz BD, Boussac M, Xu C, Grigorean G, Ziff EB, et al. Identification and verification of novel rodent postsynaptic density proteins. Mol Cell Proteom. 2004;3:857–71.
Jordan BA, Fernholz BD, Khatri L, Ziff EB. Activity-dependent AIDA-1 nuclear signaling regulates nucleolar numbers and protein synthesis in neurons. Nat Neurosci. 2007;10:427–35.
Luykx JJ, Bakker SC, Lentjes E, Neeleman M, Strengman E, Mentink L, et al. Genome-wide association study of monoamine metabolite levels in human cerebrospinal fluid. Mol psychiatry. 2014;19:228–34.
Enga RM, Rice AC, Weller P, Subler MA, Lee D, Hall CP, et al. Initial characterization of behavior and ketamine response in a mouse knockout of the post-synaptic effector gene Anks1b. Neurosci Lett. 2017;641:26–32.
Cross-Disorder Group of the Psychiatric Genomics C, Smoller JW, Craddock N, Kendler K, Lee PH, Neale BM, et al. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet. 2013;381:1371–9.
McClay JL, Adkins DE, Aberg K, Stroup S, Perkins DO, Vladimirov VI, et al. Genome-wide pharmacogenomic analysis of response to treatment with antipsychotics. Mol psychiatry. 2011;16:76–85.
McClay JL, Adkins DE, Aberg K, Bukszar J, Khachane AN, Keefe RSE, et al. genome-wide pharmacogenomic study of neurocognition as an indicator of antipsychotic treatment response in schizophrenia. Neuropsychopharmacolog. 2011;36:616–26.
Garriock HA, Kraft JB, Shyn SI, Peters EJ, Yokoyama JS, Jenkins GD, et al. A genomewide association study of citalopram response in major depressive disorder. Biol psychiatry. 2010;67:133–8.
Jia T, Macare C, Desrivieres S, Gonzalez DA, Tao C, Ji X, et al. Neural basis of reward anticipation and its genetic determinants. Proc Natl Acad Sci USA. 2016;113:3879–84.
Yalachkov Y, Kaiser J, Naumer MJ. Functional neuroimaging studies in addiction: multisensory drug stimuli and neural cue reactivity. Neurosci Biobehav Rev. 2012;36:825–35.
Wang X, Pathak S, Stefaneanu L, Yeh FC, Li S, Fernandez-Miranda JC. Subcomponents and connectivity of the superior longitudinal fasciculus in the human brain. Brain Struct Funct. 2016;221:2075–92.
Starnawska A, Tan Q, McGue M, Mors O, Borglum AD, Christensen K, et al. Epigenome-wide association study of cognitive functioning in middle-aged monozygotic twins. Front Aging Neurosci. 2017;9:413.
Johnson C, Drgon T, Liu QR, Zhang PW, Walther D, Li CY, et al. Genome wide association for substance dependence: convergent results from epidemiologic and research volunteer samples. BMC Med Genet. 2008;9:113.
Drgon T, Zhang PW, Johnson C, Walther D, Hess J, Nino M, et al. Genome wide association for addiction: replicated results and comparisons of two analytic approaches. PLoS ONE. 2010;5:e8832.
Abe K, Chisaka O, van Roy F, Takeichi M. Stability of dendritic spines and synaptic contacts is controlled by alpha N-catenin. Nat Neurosci. 2004;7:357–63.
Park C, Falls W, Finger JH, Longo-Guess CM, Ackerman SL. Deletion in Catna2, encoding alpha N-catenin, causes cerebellar and hippocampal lamination defects and impaired startle modulation. Nat Genet. 2002;31:279–84.
Lesch KP, Timmesfeld N, Renner TJ, Halperin R, Roser C, Nguyen TT, et al. Molecular genetics of adult ADHD: converging evidence from genome-wide association and extended pedigree linkage studies. J Neural Transm. 2008;115:1573–85.
Terracciano A, Esko T, Sutin AR, de Moor MH, Meirelles O, Zhu G, et al. Meta-analysis of genome-wide association studies identifies common variants in CTNNA2 associated with excitement-seeking. Transl Psychiatry. 2011;1:e49.
Hall FS, Drgonova J, Jain S, Uhl GR. Implications of genome wide association studies for addiction: are our a priori assumptions all wrong? Pharmacol Ther. 2013;140:267–79.
Acknowledgements
This work was supported by grants from the National Basic Research Program of China (2015CB553503), the National Natural Science Foundation of China (U180220091, 81821092, 81601165), the National Key Research and Development Program of China (2017YFC0803608, 2017YFC0803609, 2016YFC0800908), Beijing Municipal Science & Technology Commission (Z181100001518005 and Z161100002616006), and Youth Elite Scientists Sponsorship Program by CASR (CSTQT2017002). We are grateful to Beijing Compass Biotechnology Company for technical assistance with the microarray experiments.
Author information
Authors and Affiliations
Contributions
JS, YS, and SC designed the study and obtained financial support; YS, FW, WY, HS, ZN, and XC conducted cohort recruitment, collected biological samples, and phenotypic data. JL performed the genotype microarray experiments. SC and YS performed genetic data processing, statistical, and bioinformatics analysis. ZL performed the brain imaging analysis. LZ, YZ, and YC performed the animal and in vitro experiments. YS and SC drafted the manuscript. JS and LL supervised the experiments and data analysis. All authors critically reviewed the manuscript and approved the final version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Sun, Y., Chang, S., Liu, Z. et al. Identification of novel risk loci with shared effects on alcoholism, heroin, and methamphetamine dependence. Mol Psychiatry 26, 1152–1161 (2021). https://doi.org/10.1038/s41380-019-0497-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-019-0497-y
This article is cited by
-
SNP-based and haplotype-based genome-wide association on drug dependence in Han Chinese
BMC Genomics (2024)
-
BGWAS: Bayesian variable selection in linear mixed models with nonlocal priors for genome-wide association studies
BMC Bioinformatics (2023)
-
BICOSS: Bayesian iterative conditional stochastic search for GWAS
BMC Bioinformatics (2022)
-
Infant inhibited temperament in primates predicts adult behavior, is heritable, and is associated with anxiety-relevant genetic variation
Molecular Psychiatry (2021)
-
Sex-specific nicotine sensitization and imprinting of self-administration in rats inform GWAS findings on human addiction phenotypes
Neuropsychopharmacology (2021)