Abstract
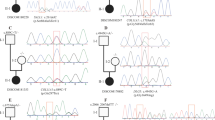
The mechanism underlying anorectal malformations (ARMs)-related VACTERL (vertebral defects, anal atresia, cardiac defects, tracheo–esophageal fistula, and renal and limb abnormalities) remains unclear. Copy number variation (CNV) contributed to VACTERL pathogenicity. Here, we report a novel CNV in 8p23 and 12q23.1 identified in a case of ARMs-related VACTERL association. This 12-year-old girl presented a cloaca (urethra, vagina, and rectum opening together and sharing a single tube length), an isolated kidney, and a perpetuation of the left superior vena cava at birth. Her intelligence, growth, and development were slightly lower than those of normal children of the same age. Array comparative genomic hybridization revealed a 9.6-Mb deletion in 8p23.1–23.3 and a 0.52-Mb duplication in 12q23.1 in her genome. Furthermore, we reviewed the cases involving CNVs in patients with VACTERL, 8p23 deletion, and 12q23.1 duplication, and our case was the first displaying ARMs-related VACTERL association with CNV in 8p23 and 12q23.1. These findings enriched our understanding between VACTERL association and the mutations of 8p23 deletion and 12q23.1 duplication.
Impact
-
This is a novel case of a Chinese girl with anorectal malformations (ARMs)-related VACTERL with an 8p23.1–23.3 deletion and 12q23.1 duplication. Cloaca malformation is presented with novel copy number variation in 8p23.1–23.3 deletion and 12q23.1 duplication.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 14 print issues and online access
$259.00 per year
only $18.50 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All of the material is owned by the authors and/or no permissions are required.
References
Tonni, G. et al. Clinical presentations and diagnostic imaging of vacterl association. Fetal Pediatr. Pathol. 42, 651–674 (2023).
Solomon, B. D. Vacterl/vater association. Orphanet J. Rare Dis. 6, 56 (2011).
Czeizel, A. & Ludányi, I. An aetiological study of the vacterl-association. Eur. J. Pediatr. 144, 331–337 (1985).
Lomas, F. E., Dahlstrom, J. E. & Ford, J. H. Vacterl with hydrocephalus: family with X-linked Vacterl-H. Am. J. Med. Genet. 76, 74–78 (1998).
Hilger, A. et al. Familial occurrence of the vater/vacterl association. Pediatr. Surg. Int 28, 725–729 (2012).
Reutter, H. & Ludwig, M. Vater/vacterl association: evidence for the role of genetic factors. Mol. Syndromol. 4, 16–19 (2013).
Jenkins, D. et al. Mutational analyses of Upiiia, Shh, Efnb2 and Hnf1beta in persistent Cloaca and associated kidney malformations. J. Pediatr. Urol. 3, 2–9 (2007).
Kause, F. et al. Hspa6: a new autosomal recessive candidate gene for the vater/vacterl malformation spectrum. Birth Defects Res. 111, 591–597 (2019).
Saisawat, P. et al. Whole-exome resequencing reveals recessive mutations in Trap1 in individuals with cakut and vacterl association. Kidney Int. 85, 1310–1317 (2014).
van de Putte, R. et al. A genetics-first approach revealed monogenic disorders in patients with arm and vacterl anomalies. Front. Pediatr. 8, 310 (2020).
Hilger, A. et al. De novo microduplications at 1q41, 2q37.3, and 8q24.3 in patients with vater/vacterl association. Eur. J. Hum. Genet. 21, 1377–1382 (2013).
LaBranche, J. T. N., Argiropoulos, B. & Thomas, M. A. 8p23.2p22 deletion: a case report of a large deletion encompassing 8p23.1 with additional clinical features. Clin. Dysmorphol. 29, 207–209 (2020).
Oh, K. S., Febres-Aldana, C. A., Pelaez, L. & Alexis, J. 8p23.1 microdeletion syndrome and obstructing myxomatous heart valve nodules. Autops. Case Rep. 10, e2020168 (2020).
Catusi, I. et al. 8p23.2-Pter microdeletions: seven new cases narrowing the candidate region and review of the literature. Genes 12, 652 (2021).
Schierz, I. A. M. et al. Clinical and genetic approach in the characterization of newborns with anorectal malformation. J. Matern. Fetal Neonatal Med. 35, 4513–4520 (2022).
Zhang, R. et al. Array-based molecular karyotyping in 115 vater/vacterl and vater/vacterl-like patients identifies disease-causing copy number variations. Birth Defects Res 109, 1063–1069 (2017).
Moreno, O. M. et al. Phenotypic characteristics and copy number variants in a cohort of colombian patients with vacterl association. Mol. Syndromol. 11, 271–283 (2020).
Brosens, E. et al. Vacterl association etiology: the impact of de novo and rare copy number variations. Mol. Syndromol. 4, 20–26 (2013).
Solomon, B. D. et al. Clinical geneticists’ views of vacterl/vater association. Am. J. Med. Genet. A 158a, 3087–3100 (2012).
Reddy, K. S. A paternally inherited terminal deletion, Del(8)(P23.1)Pat, detected prenatally in an amniotic fluid sample: a review of deletion 8p23.1 cases. Prenat. Diagn. 19, 868–872 (1999).
Baynam, G., Goldblatt, J. & Walpole, I. Deletion of 8p23.1 with features of cornelia de lange syndrome and congenital diaphragmatic hernia and a review of deletions of 8p23.1 to 8pter? A further locus for cornelia de lange syndrome. Am. J. Med. Genet. A 146a, 1565–1570 (2008).
Barber, J. C. et al. Inside the 8p23.1 duplication syndrome; eight microduplications of likely or uncertain clinical significance. Am. J. Med. Genet. A 167a, 2052–2064 (2015).
Ballarati, L. et al. Genotype-phenotype correlations in a new case of 8p23.1 deletion and review of the literature. Eur. J. Med. Genet. 54, 55–59 (2011).
& Cicenia, M. et al. 8p23.1 deletion: look out for left ventricular hypertrabeculation and not only congenital heart diseases. single-center experience and literature revision. Am. J. Med. Genet. A 188, 883–895 (2022).
Stoll, C., Alembik, Y., Dott, B. & Roth, M. P. Associated malformations in patients with anorectal anomalies. Eur. J. Med. Genet. 50, 281–290 (2007).
Thomas, D. F. M. The embryology of persistent cloaca and urogenital sinus malformations. Asian J. Androl. 22, 124–128 (2020).
Hendren, W. H. Cloaca, the most severe degree of imperforate anus: experience with 195 cases. Ann. Surg. 228, 331–346 (1998).
Southard, A. E., Edelmann, L. J. & Gelb, B. D. Role of copy number variants in structural birth defects. Pediatrics 129, 755–763 (2012).
Goldmuntz, E. et al. Microdeletions and microduplications in patients with congenital heart disease and multiple congenital anomalies. Congenit. Heart Dis. 6, 592–602 (2011).
Dworschak, G. C. et al. Genome-wide mapping of copy number variations in patients with both anorectal malformations and central nervous system abnormalities. Birth Defects Res. A Clin. Mol. Teratol. 103, 235–242 (2015).
Brosens, E. et al. Copy number variations in 375 patients with oesophageal atresia and/or tracheoesophageal fistula. Eur. J. Hum. Genet. 24, 1715–1723 (2016).
Harrison, S. M., Seideman, C. & Baker, L. A. DNA copy number variations in patients with persistent cloaca. J. Urol. 191, 1543–1546 (2014).
Ramos, F. J., McDonald-McGinn, D. M., Emanuel, B. S. & Zackai, E. H. Tricho-rhino-phalangeal syndrome type Ii (Langer-Giedion) with persistent cloaca and prune Belly sequence in a girl with 8q interstitial deletion. Am. J. Med. Genet. 44, 790–794 (1992).
Fagan, K., Wilkinson, I., Allen, M. & Brownlea, S. The coagulation factor Vii regulator is located on 8p23.1. Hum. Genet. 79, 365–367 (1988).
Kumar, V., Roy, S. & Kumar, G. An interesting and unique case of 8p23.3p23.1 deletion and 8p23.1p11.1 interstitial duplication syndrome. J. Pediatr. Genet. 7, 125–129 (2018).
Molck, M. C., Monteiro, F. P., Simioni, M. & Gil-da-Silva-Lopes, V. L. 8p23.1 interstitial deletion in a patient with congenital cardiopathy, neurobehavioral disorders, and minor signs suggesting 22q11.2 deletion syndrome. J. Dev. Behav. Pediatr. 36, 544–548 (2015).
Páez, M. T. et al. Two patients with atypical interstitial deletions of 8p23.1: mapping of phenotypical traits. Am. J. Med. Genet. A 146a, 1158–1165 (2008).
Wu, B. L. et al. Distal 8p deletion (8)(P23.1): an easily missed chromosomal abnormality that may be associated with congenital heart defect and mental retardation. Am. J. Med. Genet. 62, 77–83 (1996).
Smith, S., Giriat, I., Schmitt, A. & de Lange, T. Tankyrase, a poly(adp-ribose) polymerase at human telomeres. Science 282, 1484–1487 (1998).
Khelifa, H. B. et al. Microarray analysis of 8p23.1 deletion in new patients with atypical phenotypical traits. J. Pediatr. Genet. 4, 187–193 (2015).
Trimborn, M. et al. Mutations in microcephalin cause aberrant regulation of chromosome condensation. Am. J. Hum. Genet. 75, 261–266 (2004).
Hemmat, M. et al. Cma analysis identifies homozygous deletion of Mcph1 in 2 brothers with primary microcephaly-1. Mol. Cytogenet. 10, 33 (2017).
Tabarés-Seisdedos, R. & Rubenstein, J. L. Chromosome 8p as a potential hub for developmental neuropsychiatric disorders: implications for schizophrenia, autism and cancer. Mol. Psychiatry 14, 563–589 (2009).
Geckinli, B. B. et al. Clinical report of a patient with de novo trisomy 12q23.1q24.33. Genet. Couns. 26, 393–400 (2015).
Cheung, A. H., Stewart, R. J. & Marsden, P. A. Endothelial Tie2/Tek ligands angiopoietin-1 (Angpt1) and angiopoietin-2 (Angpt2): regional localization of the human genes to 8q22.3-Q23 and 8p23. Genomics 48, 389–391 (1998).
Garg, V. et al. Gata4 mutations cause human congenital heart defects and reveal an interaction with Tbx5. Nature 424, 443–447 (2003).
Verhoeven, K. et al. Slowed conduction and thin myelination of peripheral nerves associated with mutant rho guanine-nucleotide exchange factor 10. Am. J. Hum. Genet. 73, 926–932 (2003).
Shimizu, A. et al. A novel giant gene Csmd3 encoding a protein with Cub and Sushi multiple domains: a candidate gene for benign adult familial myoclonic epilepsy on human chromosome 8q23.3-Q24.1. Biochem. Biophys. Res. Commun. 309, 143–154 (2003).
Myung, J. K. et al. Well-differentiated liposarcoma of the oesophagus: clinicopathological, immunohistochemical and array Cgh analysis. Pathol. Oncol. Res. 17, 415–420 (2011).
Poniah, P. et al. Genome-wide copy number analysis reveals candidate gene loci that confer susceptibility to high-grade prostate cancer. Urol. Oncol. 35, 545.e541–545.e511 (2017).
Rutherford, S., Hampton, G. M., Frierson, H. F. & Moskaluk, C. A. Mapping of candidate tumor suppressor genes on chromosome 12 in adenoid cystic carcinoma. Lab. Investig. 85, 1076–1085 (2005).
Dong, J. et al. Genome-wide association study identifies a novel susceptibility locus at 12q23.1 for lung squamous cell carcinoma in Han Chinese. PLoS Genet. 9, e1003190 (2013).
Vijayakrishnan, J. et al. A genome-wide association study identifies risk loci for childhood acute lymphoblastic leukemia at 10q26.13 and 12q23.1. Leukemia 31, 573–579 (2017).
Veluchamy, A. et al. Association of genetic variant at chromosome 12q23.1 with neuropathic pain susceptibility. JAMA Netw. Open 4, e2136560 (2021).
Dong, Y. et al. Genome-wide scan for hypertension linkage to chromosome 12q23.1 - Q23.3 in a Chinese family. Indian J. Med. Res. 137, 935–941 (2013).
Borg, K. et al. Molecular analysis of a constitutional complex genome rearrangement with 11 breakpoints involving chromosomes 3, 11, 12, and 21 and a approximately 0.5-Mb submicroscopic deletion in a patient with mild mental retardation. Hum. Genet. 118, 267–275 (2005).
Garcia-Barceló, M. M. et al. Identification of a Hoxd13 mutation in a vacterl patient. Am. J. Med. Genet. A 146a, 3181–3185 (2008).
Aguinaga, M., Zenteno, J. C., Pérez-Cano, H. & Morán, V. Sonic hedgehog mutation analysis in patients with vacterl association. Am. J. Med. Genet. A 152a, 781–783 (2010).
Agochukwu, N. B. et al. Analysis of Foxf1 and the Fox gene cluster in patients with vacterl association. Eur. J. Med. Genet. 54, 323–328 (2011).
Hernández-García, A. et al. Contribution of Lpp copy number and sequence changes to Esophageal Atresia, Tracheoesophageal Fistula, and Vacterl Association. Am. J. Med. Genet. A 158a, 1785–1787 (2012).
Kelley, R. I. & Hennekam, R. C. The Smith-Lemli-Opitz syndrome. J. Med. Genet. 37, 321–335 (2000).
Jenkins, D. et al. De novo Uroplakin Iiia heterozygous mutations cause human renal adysplasia leading to severe kidney failure. J. Am. Soc. Nephrol. 16, 2141–2149 (2005).
Dworschak, G. C. et al. De novo 13q deletions in two patients with mild anorectal malformations as part of vater/vacterl and vater/vacterl-like association and analysis of Efnb2 in patients with anorectal malformations. Am. J. Med. Genet. A 161a, 3035–3041 (2013).
Ronzoni, L. et al. 2q33.1q34 deletion in a girl with brain anomalies and anorectal malformation. Cytogenet. Genome Res. 150, 23–28 (2016).
Smigiel, R. et al. Oesophageal atresia with tracheoesophageal fistula and anal atresia in a patient with a de novo microduplication in 17q12. Eur. J. Med. Genet. 57, 40–43 (2014).
Duan, F., Zhai, Y. & Kong, X. Clinical and genetic analysis of a child with X-linked hypohidrotic ectodermal dysplasia.Zhonghua Yi Xue Yi Chuan Xue Za Zhi 38, 469–471 (2021).
Schramm, C. et al. De novo microduplication at 22q11.21 in a patient with vacterl association. Eur. J. Med. Genet. 54, 9–13 (2011).
Ueda, H. et al. Combination of Miller-Dieker syndrome and vacterl association causes extremely severe clinical presentation. Eur. J. Pediatr. 173, 1541–1544 (2014).
Bhagat, M. Vacterl association-type anomalies in a male neonate with a Y-chromosome abnormality. Oxf. Med. Case Rep. 2015, 164–166 (2015).
Kang, J., Mao, M., Zhang, Y., Ai, F. F. & Zhu, L. Congenital anal atresia with rectovestibular fistula, scoliosis, unilateral renal agenesis, and finger defect (vacterl association) in a patient with partial bicornuate uterus and distal vaginal atresia: a case report. Medicines 97, e12822 (2018).
Tumini, S. et al. Yq microdeletion in a patient with vacterl association and shawl scrotum with bifid scrotum: a real pathogenetic association or a coincidence? Cytogenet. Genome Res. 158, 121–125 (2019).
He, X. et al. Reduced anogenital distance, hematuria and left renal hypoplasia in a patient with 13q33.1-34 deletion: case report and literature review. BMC Pediatr. 20, 327 (2020).
Chung, B. et al. From vacterl-h to heterotaxy: variable expressivity of Zic3-related disorders. Am. J. Med. Genet. A 155a, 1123–1128 (2011).
Puvabanditsin, S. et al. Vater/vacterl association and caudal regression with Xq25-Q27.3 microdeletion: a case report. Fetal Pediatr. Pathol. 35, 133–141 (2016).
Husain, M. et al. Phenotypic diversity of patients diagnosed with vacterl association. Am. J. Med. Genet. A 176, 1830–1837 (2018).
Umaña, L. A., Magoulas, P., Bi, W. & Bacino, C. A. A male newborn with vacterl association and fanconi anemia with a fancb deletion detected by array comparative genomic hybridization (Acgh). Am. J. Med. Genet. A 155a, 3071–3074 (2011).
Akcakaya, N. H. et al. De novo 8p23.1 deletion in a patient with absence epilepsy. Epileptic Disord. 19, 217–221 (2017).
Shi, S., Lin, S., Chen, B. & Zhou, Y. Isolated chromosome 8p23.2‑Pter deletion: novel evidence for developmental delay, intellectual disability, microcephaly and neurobehavioral disorders. Mol. Med. Rep. 16, 6837–6845 (2017).
Wagner-Mahler, K. et al. Is interstitial 8p23 microdeletion responsible of 46, Xy gonadal dysgenesis? One case report from birth to puberty. Mol. Genet Genom. Med 7, e558 (2019).
Lo Bianco, M. et al. Deciphering the invdupdel(8p) genotype-phenotype correlation: our opinion. Brain Sci. 10, 451 (2020).
Hao, D. et al. Inherited unbalanced translocation (4p16.3p15.32 duplication/8p23.3p23.2deletion) in the four generation pedigree with intellectual disability/developmental delay. Mol. Cytogenet. 14, 35 (2021).
Zhang, B., Cui, W., Zhang, Z., Li, J. & Shang, Q. Phenotypic and genetic analysis of a boy with Inv Dup Del(8p). Zhonghua Yi Xue Yi Chuan Xue Za Zhi 38, 581–584 (2021).
Slavotinek, A. et al. Fryns syndrome phenotype caused by chromosome microdeletions at 15q26.2 and 8p23.1. J. Med. Genet. 42, 730–736 (2005).
Chen, C. P. et al. Prenatal diagnosis of de novo partial trisomy 13q (13q22 –> Qter) and partial monosomy 8p (8p23.3 –> Pter) associated with holoprosencephaly, premaxillary agenesis, hexadactyly, and a hypoplastic left heart. Prenat. Diagn. 25, 334–336 (2005).
Simioni, M. et al. Insertional translocation of 15q25-Q26 into 11p13 and duplication at 8p23.1 characterized by high resolution arrays in a boy with congenital malformations and aniridia. Am. J. Med. Genet. A 158a, 2905–2910 (2012).
Zeng, Y. et al. Prenatal diagnosis and genetic analysis of a special case with complex structural rearrangements of chromosome 8. Zhonghua Yi Xue Yi Chuan Xue Za Zhi 40, 1181–1184 (2023).
Winberg, J. et al. Pathogenic copy number variants are detected in a subset of patients with gastrointestinal malformations. Mol. Genet. Genom. Med. 8, e1084 (2020).
Meng, X. & Jiang, L. Prenatal detection of chromosomal abnormalities and copy number variants in fetuses with congenital gastrointestinal obstruction. BMC Preg. Childbirth 22, 50 (2022).
Thiem, C. E. et al. Re-sequencing of candidate genes Foxf1, Hspa6, Haao, and Kynu in 522 individuals with vater/vacterl, vacter/vacterl-like association, and isolated anorectal malformation. Birth Defects Res. 114, 478–486 (2022).
Reinicke, T., Costantino, C. L., Anderson, D. J., Tran, J. & Griggs, C. A network of anomalies prompting vacterl workup in a trisomy 21 newborn. Cureus 14, e21290 (2022).
Kotani, T. et al. Prenatal diagnosis of distal 13q deletion syndrome in a fetus with esophageal atresia: a case report and review of the literature. J. Med. Case Rep. 16, 481 (2022).
Barber, J. C. et al. 8p23.1 duplication syndrome; a novel genomic condition with unexpected complexity revealed by array Cgh. Eur. J. Hum. Genet 16, 18–27 (2008).
Pope, K. et al. Dextrocardia, atrial septal defect, severe developmental delay, facial anomalies, and supernumerary ribs in a child with a complex unbalanced 8;22 translocation including partial 8p duplication. Am. J. Med. Genet. A 158a, 641–647 (2012).
Chen, C. P. et al. Molecular cytogenetic characterization of Inv Dup Del(8p) in a fetus associated with ventriculomegaly, hypoplastic left heart, polyhydramnios and intestinal obstruction. Taiwan J. Obstet. Gynecol. 55, 415–418 (2016).
Tokutake, H. & Chiba, S. A case report of respiratory syncytial virus-infected 8p inverted duplication deletion syndrome with low natural killer cell activity. Tohoku J. Exp. Med. 257, 347–352 (2022).
Grassi, M. S. et al. Cytogenomics investigation of infants with congenital heart disease: experience of a Brazilian Center. Arq. Bras. Cardiol. 118, 61–67 (2022).
Montenegro, M. M. et al. Expanding the phenotype of 8p23.1 deletion syndrome: eight new cases resembling the clinical spectrum of 22q11.2 microdeletion. J. Pediatr. 252, 56–60.e52 (2023).
Hong, N. et al. The transcription factor Sox7 modulates endocardiac cushion formation contributed to atrioventricular septal defect through Wnt4/Bmp2 signaling. Cell Death Dis. 12, 393 (2021).
Funding
This work was supported by the National Key Research and Development Program (2021YFC2701003,2021YFC2701104) and the National Natural Science Foundation of China (Grant no. 82171649) by Zhengwei Yuan, the National Natural Science Foundation of China (Grant no. 82070531) by H.J., and the Basic Research project of the Education Department of Liaoning Province (grant number LJKMZ20221195) by Z.Y.
Author information
Authors and Affiliations
Contributions
Y.L. wrote the manuscript. Y.L. and P.L. reviewed the manuscript. W.W., H.J., and Z.Y. revised and completed the manuscript. Z.Y. wrote and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The protocol for this study was approved by the Animal Ethics Committee of Shengjing Hospital of China Medical University (No. 2021PS183K).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Y., Liu, P., Wang, W. et al. A novel genotype-phenotype between persistent-cloaca-related VACTERL and mutations of 8p23 and 12q23.1. Pediatr Res 95, 1246–1253 (2024). https://doi.org/10.1038/s41390-023-02928-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-023-02928-0