Abstract
Male sterility enables hybrid crop breeding to increase yields and has been extensively studied. But thermo-sensitive female sterility, which is an ideal property that may enable full mechanization in hybrid rice breeding, has rarely been investigated due to the absence of such germplasm. Here we identify the spontaneous thermo-sensitive female sterility 1 (tfs1) mutation that confers complete sterility under regular/high temperature and partial fertility under low temperature as a point mutation in ARGONAUTE7 (AGO7). AGO7 associates with miR390 to form an RNA-Induced Silencing Complex (RISC), which triggers the biogenesis of small interfering RNAs (siRNAs) from TRANS-ACTING3 (TAS3) loci by recruiting SUPPRESSOR OF GENE SILENCING (SGS3) and RNA-DEPENDENT RNA POLYMERASE6 (RDR6) to TAS3 transcripts. These siRNAs are known as tasiR-ARFs as they act in trans to repress auxin response factor genes. The mutant TFS1 (mTFS1) protein is compromised in its ability to load the miR390/miR390* duplex and eject miR390* during RISC formation. Furthermore, tasiR-ARF levels are reduced in tfs1 due to the deficiency in RDR6 but not SGS3 recruitment by mTFS1 RISC under regular/high temperature, while low temperature partially restores mTFS1 function in RDR6 recruitment and tasiR-ARF biogenesis. A miR390 mutant also exhibits female sterility, suggesting that female fertility is controlled by the miR390-AGO7 module. Notably, the tfs1 allele introduced into various elite rice cultivars endows thermo-sensitive female sterility. Moreover, field trials confirm the utility of tfs1 as a restorer line in fully mechanized hybrid rice breeding.
Similar content being viewed by others
Introduction
Heterosis, or hybrid vigor, has been extensively exploited to increase rice productivity.1,2 Hybrid rice seeds can be produced using three-line or two-line schemes.1,2 During hybrid seed production, a restorer line is used as the pollen donor. However, the hybrid breeding systems suffer intrinsic problems. As the restorer line has to provide enough pollen for hybridization, it needs to be grown near the male sterile line. To avoid self-seeds from contaminating hybrid seeds, manual labor is needed to remove self-seeds from the restorer line before mechanically harvesting hybrid seeds, thus preventing full mechanization in agricultural practices. Theoretically, a thermo-sensitive female-sterile line is an ideal restorer line that acts as the pollen donor for a male-sterile line to produce hybrid seeds. However, such germplasm remains rare in nature.
In plants, small RNAs including microRNAs (miRNAs) and small interfering RNAs (siRNAs) produced by Dicer-like (DCL) proteins associate with Argonaute (AGO) family proteins to cause post-transcriptional or transcriptional gene silencing.3,4 In miRNA biogenesis, the processing of single-stranded precursor RNAs containing hairpin structures by DCL1 generates miRNA/miRNA* duplexes, which are loaded into AGO proteins to form pre-RISCs. A pre-RISC is then processed into a mature RISC through the ejection of the passenger strand. On the other hand, siRNAs are DCL products from long double-stranded RNAs (dsRNAs). One type of endogenous siRNAs is produced by the successive dicing of long dsRNAs and is known as phased siRNAs or phasiRNAs.3 The biogenesis of certain phasiRNAs involves a miRISC trigger. For example, AGO7 specifically loads miR390 to target noncoding TAS3 transcripts at two sites (3ʹ and 5ʹ sites), leading to their cleavage at the 3′ site.5,6,7 Interaction between AGO7 RISC and TAS3 RNA at the 5′ site recruits SGS3, which then promotes the recruitment of RDR6; cleavage of TAS3 RNA at the 3′ site removes the polyA tail to promote the synthesis of the complementary strand by RDR6.8,9,10 The resulting dsRNAs are then processed into 21-nt phasiRNAs by DCL proteins, and one species of the phasiRNAs can target auxin responsive factor (ARF) transcripts in trans and is known as trans-acting siRNA (tasiR-ARF).8,11 Through the repression of ARF family members, miR390 and tasiR-ARF regulate numerous developmental processes such as leaf polarity patterning,12,13 developmental timing8,14,15 and lateral root growth.16,17,18 Moreover, the miR390-TAS3-ARF pathway is highly conserved in land plants.19 ago7 mutants in plants including Arabidopsis,8,20 tomato21,22 maize,23 and rice24,25 display abnormal vegetative development. Maize and rice ago7 mutants exhibit strong defects in shoot apical meristem maintenance and leaf polarity and are often seedling lethal.23,24,25 It is still unknown whether AGO7 functions in plant reproduction and whether its activities are modulated by temperature in rice. Here, we report a unique spontaneous thermo-sensitive female sterility mutation in the TFS1 gene encoding AGO7 and elucidate how the mutation responds to temperature to control the female sterility-fertility conversion without causing any defects in vegetative or male reproductive development. Field trials further confirmed that tfs1 shows a female-sterility trait suitable for hybrid rice seed production system, which endows hybrid breeding in full mechanization.
Results
tfs1 is a thermo-sensitive female-sterile mutant
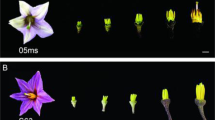
tfs1 was identified in an elite cultivar 4266 (wild type) during paddy field production as a strictly sterile germplasm with normal vegetative growth under regular/high growth temperature (>25 °C) (Fig. 1a‒c). The mutant also exhibited an open-hull phenotype (Fig. 1b). The fertility and open hull phenotype were partially recovered (more than 68% seed setting in one panicle) when tfs1 plants were subjected to low-temperature growth conditions (23 °C) before the booting stage (Fig. 1d; Supplementary information, Table S1). The test of day length/temperature combinations indicated that tfs1 responded to temperature but not day length (Supplementary information, Table S2). Nonetheless, seed setting of both 4266 and tfs1 was reduced at 22 °C, probably because of their indica background (Supplementary information, Table S2). Under regular/high temperature, F1 progeny from the cross between tfs1 (as the pollen donor, ♂) and 4266 (as the pollen acceptor, ♀) were obtained and showed normal fertility (Supplementary information, Fig. S1e and Table S3). However, pollination of tfs1 with pollen from 4266 failed to yield any progeny (Supplementary information, Fig. S1e). This finding demonstrated that tfs1 exhibited female-specific sterility. Consistent with this, I2-KI staining showed that tfs1 pollen grains were viable as in 4266 (Supplementary information, Fig. S1a, b and Table S4). Moreover, normal anther exertion and dehiscence with abundant pollen were observed in tfs1 during the time of flowering (~9:00 to 13:00). The tfs1 pistils appeared to be largely normal (Supplementary information, Fig. S1c) and tfs1 ovules also had normal 7-cell, 8-nucleus embryo sacs (Fig. 1e, 0HAP).
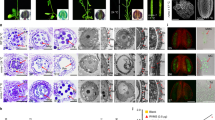
a Plants of 4266 and tfs1 grown at regular/high temperature (> 25 °C). b Florets of tfs1 and 4266 plants grown under regular/high temperature. Scale bar, 5 mm. c Grain development in tfs1 and 4266 plants grown under regular/high temperature. DAP, day after pollination. Scale bar, 1 mm. d Panicles of 4266 and tfs1 under regular/high and low temperature (LT, 23 °C). Scale bars, 2 cm. e Post-pollination embryo development in 4266 and tfs1 plants grown under regular/high temperature. A, antipodal cells; PN, polar nuclei; PT, pollen tube; E, egg; S, synergids; Em, embryo; SN, sperm nucleus. HAP, hours after pollination. Scale bars, 50 μm. The white dotted line shows the location of the normal pollen tube entering the embryo sac. The number of normal pollen tube entering the embryo sac is indicated at the upper left corner at 2 HAP. f Floret development at four stages in 4266 and tfs1 plants grown under regular/high temperature. Stage 1 is characterized by caryopsis primordia differentiation; stages 2 and 3 are marked by pistil and stamen formation, and stage 4 is when the floret starts to grow. Red arrows, lemma primordia; white arrows, palea primordia; stars, stamen primordia; the pistil primordium is in the center flanked by the stamen primordia. Scale bars, 400 μm.
Pollen grains attach to the stigma and germinate to generate pollen tubes after pollination. Thereafter, the pollen tube enters the stigma and then the transmission tract of the style, and eventually reaches the micropyle at the base of ovary.26 To investigate the behavior of pollen tube growth, we observed a large number of self-pollinated pistils. The aniline blue staining showed normal pollen germination and sufficient pollen tube growth to reach the micropyle in both tfs1 and 4266 ovules (Supplementary information, Fig. S1d). However, reaching the micropyle does not guarantee its subsequent success in entering the embryo sac because the rice micropyle is not a hole but is a solid structure consisting of transmitting tissues. Abundant communications with the surrounding sporophytic maternal tissues guide pollen tube growth to enter the embryo sac after the pollen tube reaches the micropyle. To further investigate the pollen tube growth behavior after reaching the micropyle, we observed fertilized pistils stained by eosin Y under confocal microscope. At 2 h after pollination (HAP), the pollen tube (white dotted line) entered the embryo sac and the sperm nucleus reached the central cell (with two polar nuclei) in 4266 (Fig. 1e). By contrast, the pollen tube in tfs1 failed to enter the embryo sac, indicating blocked pollen tube growth after reaching the micropyle (Fig. 1e; Supplementary information, Table S4). The sperm nucleus or the polar nuclei were invisible at 1 day after pollination (DAP) and an embryo was visible at 7 DAP (Fig. 1e) in 4266, whereas the polar nuclei remained visible without the sperm nucleus from 0 HAP to 7 DAP in tfs1 (Fig. 1e). Therefore, blocked pollen tube growth after reaching the micropyle resulted in the failure of double fertilization and seed set in tfs1. These observations, together with results from reciprocal crosses (Supplementary information, Fig. S1e) and the fertility of mutant pollen suggested that the pollen tube growth arrest in tfs1 was caused by a maternal defect and that the mutation did not affect the function of the male gametophyte. Besides, low temperature partially rescued the blockage of pollen tube growth after reaching the micropyle in tfs1 (Supplementary information, Fig. S1f).
Since the mutant also exhibited an open-hull phenotype, we used scanning electron microscopy to study floret development in four stages.27 The hull is derived from lemma and palea primordia. In comparison to 4266, the lemma primordium of tfs1 failed to expand towards the palea, leaving a gap in the hull at stage 4 (Fig. 1f). The anther and pistil primordia were morphologically similar between tfs1 and 4266 (Fig. 1f).
The TFS1 trait locus encodes AGO7
To identify the locus responsible for the mutant phenotype in tfs1, map-based cloning was performed. An F2 population from the cross between tfs1(♂) and Nipponbare (Nip) (♀) segregated 3:1 (fertile to sterile plants, the abnormal hull phenotype strictly co-segregated with female sterility) in paddy fields, indicating that female sterility was controlled by a single, recessive, and nuclear locus (Supplementary information, Fig. S2a). Preliminary mapping and a modified MutMap method28 revealed one single nucleotide polymorphism (SNP) between 19 and 20 centiMorgan (cM) on chromosome 3 to be tightly linked to tfs1. The SNP was a C-to-A substitution in the genic region of AGO7, leading to a Leu-to-Ile substitution at amino acid 603 in linker 2 (Fig. 2a; Supplementary information, Fig. S2b). Therefore, we hypothesized that the SNP located in AGO7 was the candidate tfs1 mutation. Genetic complementation further confirmed the identity of this gene (Fig. 2b; Supplementary information, Fig. S2c and Table S1).
a Positional cloning of the TFS1 gene (red dot). The chromosomal locations and the numbers of recombinants are indicated by the numbers above and below the chromosome, respectively. Pink triangle indicates mutation position (a C-to-A substitution) that was identified by MutMap after initial mapping and is located in TFS1 in the 2.16 Mb candidate region. b Panicles of a complementation line grown under regular/high temperature (> 25 °C). Scale bars, 2 cm. c Abundance of miR390 and miR390* in 4266 and tfs1 under regular/high temperature. Data are means ± SD (n = 3). **P < 0.01 as determined by two-tailed Student’s t-test. d Abundance of tasiR-ARFs from TAS3 loci under regular/high temperature (Supplementary information, Table S2). e Transcript levels of ARF15 in 4266, tfs1 and TFS1:HA-mTFS1/tfs1 under regular/high temperature. Levels were relative to UBQ5. Data are means ± SD (n = 3). f Pistil, anther, and floret of Nip and the mir390-3 mutant under regular/high temperature. Scale bars, 1 mm. g Panicles of mir390-3 under regular/high temperature. Scale bars, 1 cm. h RNA gel blotting showing the abundance of miR390 and siRNA-1 (from TAS3b) under regular/high and low temperature (LT, 23 °C) in Nip and mir390-3. U6 was the internal control. The numbers represent relative abundance, with the levels in Nip set at 1. i Post-pollination embryo development in Nip and mir390-3 under regular/high temperature. A, antipodal cells; PN, polar nuclei; PT, pollen tube; E, egg; S, synergids; Em, embryo; SN, sperm nucleus. HAP, hours after pollination. DAP, days after pollination. Scale bars, 50 μm. The white dotted line shows the location of the normal pollen tube entering the embryo sac. The number of ovules with normal pollen tubes that have entered the embryo sac at 2 HAP is indicated at the upper left corner.
tfs1 is defective in tasiR-ARF biogenesis
To unveil the molecular basis for the tfs1 phenotype, we performed small RNA sequencing with tfs1 and 4266 grown under regular/high temperature (Supplementary information, Fig. S3a, b). We observed significantly higher levels of both miR390 and miR390* in tfs1 than in 4266 (Fig. 2c). The increase in miR390 and miR390* levels was rescued by the pTFS1:TFS1 genomic DNA (Supplementary information, Fig. S3c). Despite the increased miR390 abundance, the levels of TAS3-derived siRNAs, including tasiR-ARFs and non-tasiR-ARF siRNAs, were reduced in tfs1 as compared to 4266 (Fig. 2d; Supplementary information, Fig. S3d middle panel and Table S5), which resulted in increased transcript levels of the tasiR-ARF target gene ARF15 (LOC_Os05g48870) (Fig. 2e). Importantly, tasiR-ARF-resistant OsARF15 mutant lines were sterile, which phenocopied the tfs1 mutant.29,30 We performed in situ hybridization to determine whether the changes in small RNA accumulation occurred in floret primordia, embryo sacs, and anthers. An increase in miR390 and miR390* signals and a reduction in the signals of a TAS3-derived, non-tasiR-ARF siRNA (siRNA-1) (Supplementary information, Figs. S4 and S5a) were observed in tfs1 florets and embryo sacs under regular/high temperature. However, no signals were detected in anthers in either tfs1 or 4266 (Supplementary information, Fig. S5b).
Strong AGO7 mutations in rice abolish tasiR-ARF biogenesis, resulting in severe phenotypes such as seedling lethality.25 However, the tfs1 plants exhibited normal vegetative development with the only phenotypes being open hulls and female sterility. This weak phenotype may be due to the presence of sufficient levels of tasiR-ARFs to support vegetative development. To determine whether weak alleles in another component of the tasiR-ARF biogenesis pathway produce a similar phenotype, we constructed mutations in MIR390 using CRISPR/Cas9. Two mutations, mir390-1 and mir390-2, were only obtained in a heterozygous state, and one homozygous mutant, mir390-3, was generated (Supplementary information, Fig. S5c). The three mutants all had normal pollen viability under regular/high temperature (Supplementary information, Fig. S5d). The two heterozygous lines were partially sterile (Supplementary information, Fig. S5e), while mir390-3 was sterile with open hulls and normal-looking pistils, resembling the phenotype of tfs1 under regular/high temperature (Fig. 2f, g). We further performed reciprocal crosses to survey the female fertility of the mir390-3 mutant. F1 progeny was obtained when the mir390-3 mutant was crossed as a pollen donor with wild-type lines. However, no seeds were obtained when the mir390-3 mutant was crossed as the female parent with pollen grains from wild-type lines. This suggested that the mir390-3 mutant was female sterile. The mir390-3 mutant showed reduced levels of siRNA-1, indicating a mild defect in tasiR-ARF biogenesis (Fig. 2h). Notably, low temperature was unable to restore female fertility or levels of siRNA-1 in mir390-3 (Fig. 2h). Moreover, blocked pollen tube growth after reaching the micropyle was also observed in mir390-3 at 2HAP (Fig. 2i). Taken together, the results on tfs1 and mir390-3 mutants suggest that female fertility is controlled by the miR390-AGO7 module.
Defective miR390/miR390* duplex loading and miR390* ejection in tfs1
To understand how the tfs1 allele causes reduced tasiR-ARF biogenesis, we first investigated the effects of the mutation on the expression of TFS1 and the subcellular localization of the TFS1 protein. Both GFP-TFS1 and GFP-mTFS1 proteins were found to be localized in the cytoplasm (Supplementary information, Fig. S6a), and TFS1 transcript levels were similar in 4266 and tfs1 under both regular/high and low temperature (Supplementary information, Fig. S6b).
Given the observed higher levels of miR390* in tfs1, next, we detected whether the levels of miR390 and miR390* bound by TFS1 were increased in tfs1 compared to 4266. We sequenced mTFS1- and TFS1-associated small RNAs from the HA-tagged transgenic lines of pTFS1:HA-mTFS1/tfs1 and pTFS1:HA-TFS1/4266. Sequencing results suggested that miR390 was bound by both TFS1 and mTFS1 as expected (Supplementary information, Fig. S6c, d). miR390* was more enriched in the mTFS1 immunoprecipitate than that of TFS1 (Supplementary information, Fig. S6d). miR390 was also at a higher level in the mTFS1 immunoprecipitate, although the difference was not statistically significant (Supplementary information, Fig. S6d).
In addition, we detected the unwinding of the miR390/miR390* duplex from mTFS1 pre-RISCs in vitro. We utilized translationally active lysates from the evacuolated protoplasts of tobacco BY-2 cultured cells31 to examine the effects of the tfs1 mutation on miR390-AGO7 RISC assembly (Fig. 3a). Both dsRNAs and single-stranded RNAs (ssRNAs) were co-immunoprecipitated with TFS1 and mTFS1, when miR390 was labeled with 32P. This suggested that both pre-RISCs containing miR390/miR390* and mature RISCs containing only miR390 were assembled by TFS1 and mTFS1(Fig. 3b). However, the abundance of the mTFS1 pre-RISCs containing dsRNA was 0.6-fold that of the TFS1 pre-RISCs (Fig. 3b, c), suggesting that mTFS1 was less efficient in loading miR390/miR390* than TFS1. The abundance of mature mTFS1 RISCs containing miR390 was only 0.3-fold that of mature TFS1 RISCs (Fig. 3b, c). The ratio of ss/ds RNA in TFS1 (0.9) was over two-fold that in mTFS1 (0.4) (Fig. 3c), indicating that mTFS1 was also defective in miR390* ejection.
a–c Biochemical assays for miR390-TFS1 or miR390-mTFS1 RISC assembly and miR390* ejection. The workflow is illustrated in a. Mature RISC (ss) only contains miR390, not miR390* (b). The ability to eject miR390* is shown by the ss/ds ratio (b and c at 25 °C). In b and c the upper panel is a native RNA gel and the lower panel is a western blot (WB) showing the levels of immunoprecipitated TFS1 or mTFS1. The numbers represent the relative abundances of pre-RISCs (ds) and RISCs (ss) normalized to protein abundance. Mock, no mRNA. d Protein half-life measurements in rice protoplasts. The upper panel shows protein abundance. The middle panel shows miR390 abundance in the GFP-TFS1 and GFP-mTFS1 immunoprecipitates. The bottom panel reflects protein input stained with Coomassie blue (CBB). The numbers below the protein and RNA blots indicate abundance relative to 0 h. e Slicer assays using HA-TFS1 or HA-mTFS1 immunoprecipitates from extracts of plants grown under regular/high temperature. The RNA substrates were TAS3b or TAS3b-5ʹ3ʹm. TAS3b-5ʹ3ʹm was non-cleavable. The top panel shows the full-length RNA and the two cleavage products as indicated. The second panel shows TFS1 or mTFS1 protein levels as determined by anti-HA WB. The third panel shows miR390 abundance in the immunoprecipitates. CBB staining (bottom panel) shows input proteins. Numbers indicate the relative abundance normalized to protein input. f Detection of interactions between TFS1 (or mTFS1) and SGS3 or between TFS1 (or mTFS1) and RDR6. Total extracts of TFS1: HA-TFS1/4266 and TFS1: HA-mTFS1/tfs1 grown under regular/high temperature were immunoprecipitated with anti-HA, and the immunoprecipitates were then probed by anti-HA (TFS1), anti-SGS3 and anti-RDR6 antibodies. The numbers represent relative abundance, with the level of TFS1 set at 1. g Diagram of part of the tasiR-ARF biogenesis pathway showing the RDR6 product and its detection with a TAS3b sense probe (in blue). h RNA gel blotting was used to detect and quantify RDR6 products in 4266 and tfs1 grown under regular/high temperature. U6 was the internal control. The negative control RNA was similarly produced by in vitro transcription without using biotin-UTP (the two lanes represent two replicates). The TAS3b full-length size control was performed with two replicates. The numbers represent relative abundance, with the levels in 4266 set at 1. All experiments were repeated at least three times with similar results.
Interestingly, the reduced efficiency of RISC formation by mTFS1 observed in vitro was contrary to the more enrichment of miR390/miR390* in the mTFS1 immunoprecipitate and the higher levels of miR390 and miR390* in tfs1 in vivo. We hypothesized that this inconsistency was caused by the longer half-live of mTFS1 than TFS1. To test our hypothesis, we examined the half-lives of TFS1 and mTFS1 using rice protoplasts transformed with 35S:GFP-TFS1 or 35S:GFP-mTFS1. In the presence of the translation inhibitor cycloheximide, the abundance of GFP-mTFS1 decreased much more slowly than that of GFP-TFS1 over time, suggesting that mTFS1 was more stable than TFS1 (Fig. 3d). We also examined the abundance of GFP-TFS1- and GFP-mTFS1-associated miR390 by immunoprecipitation of GFP-TFS1 or GFP-mTFS1 followed by RNA gel blot analysis in cycloheximide-treated protoplasts. The abundance of miR390 decreased in a manner similar to that of TFS1 or mTFS1 over the time course (Fig. 3d), demonstrating that the turnover of miR390 was positively correlated with that of TFS1 or mTFS1 proteins over the time course. Thus, the higher levels of miR390 and miR390* in mTFS1 immunoprecipitate probably reflected the increased stability of the mTFS1 pre-RISCs or mature RISCs in vivo.
Defective recruitment of RDR6 by mTFS1
Given the potentially higher levels of mTFS1 RISCs in vivo, the question remains as to why the mTFS1 RISCs produced lower levels of tasiR-ARFs. We first considered the possibility that mTFS1 RISCs were defective in cleaving TAS3 transcripts to produce 5ʹ fragments, from which tasiR-ARFs are produced. We thereby used TFS1 and mTFS1 immunoprecipitates from the HA-tagged transgenic lines of pTFS1:HA-mTFS1/tfs1 and pTFS1:HA-TFS1/4266, respectively, to perform a slicer assay on an in vitro transcribed TAS3b transcript. Consistent with previous findings, more mTFS1 and its associated miR390 were immunoprecipitated than TFS1-miR390 (Fig. 3e, the second panel). More 5ʹ TAS3b fragment was produced by mTFS1 (Fig. 3e, the top panel) in vitro, which was consistent with levels of the 5ʹ TAS3b fragment in tfs1 in vivo, as observed by RNA sequencing (Supplementary information, Fig. S3d, the third panel). These results suggested that in tfs1 plants, mTFS1 RISCs were able to effectively cleave TAS3b RNA. To further investigate why mTFS1 RISCs produced lower levels of tasiR-ARFs, we next examined the steps in tasiRNA biogenesis after TAS3b slicing, that is, the recruitment of SGS3 and RDR6 to AGO7.5,7,8,31,32 IP was performed with HA-tagged transgenic lines described above, and SGS3 and RDR6 were detected in the IP. The levels of SGS3 associated with TFS1 and mTFS1 were similar, but mTFS1 was less efficient in recruiting RDR6 than TFS1 (Fig. 3f). We also detected the antisense RNA generated by RDR6 from TAS3b RNA using a sense RNA probe. Indeed, the antisense RNA was present at much lower levels in tfs1 than in 4266 under high/regular temperature (Fig. 3g, h), indicating reduced RDR6 activity in tfs1. To determine whether the stability of RDR6 was affected in tfs1, we examined the decay of RDR6 upon inhibition of protein synthesis in protoplasts. The abundance of RDR6 decreased to similar degrees over time in 4266 and tfs1 (Supplementary information, Fig. S6e, f), suggesting that RDR6 stability was not affected in tfs1. Next, we tested whether phasiRNAs other than tasiR-ARFs were affected in tfs1 by small RNA sequencing. Reads in our small RNA sequencing data were mapped to previously reported PHAS loci or RDR6-dependent PHAS loci.33 The abundance of 21-nt phasiRNAs in tfs1 showed no global difference from that in 4266 (Supplementary information, Fig. S6g). The RDR6-dependent 21-nt phasiRNAs also displayed similar abundance between 4266 and tfs1 (Supplementary information, Fig. S6g). However, low temperature reduced the levels of 21-nt phasiRNAs in both 4266 and tfs1 (Supplementary information, Fig. S6g). These results indicate that the mutation in AGO7 did not affect the accumulation of phasiRNAs triggered by other mechanisms such as miR2118.
In summary, the mTFS1 protein differed from TFS1 in several ways; specifically, it exhibited: (1) reduced efficiency in miR390/miR390* loading, (2) reduced miR390* ejection, (3) increased stability, and (4) reduced ability to recruit RDR6, with the last defect being accountable for the decrease in tasiR-ARF biogenesis in the tfs1 mutant.
Low temperature partially restores mTFS1 function
Next, we determined the temperature-induced changes in small RNA abundance using RNA gel blotting and RT-qPCR assays. Increased abundance of miR390 and miR390* was detected in tfs1 compared with 4266 under regular/high temperature. In contrast, miR390* levels were reduced in tfs1 under low temperature (Fig. 4a, b). Two non-tasiR-ARF siRNAs from the TAS3 loci showed lower levels in tfs1 at regular/high temperature; low temperature did not affect their levels in 4266 but substantially increased their levels in tfs1 (Fig. 4b). The abundance of the two tasiR-ARFs was also increased in tfs1 at low temperature compared with regular/high temperature (Fig. 4a). Additionally, the expression level of target gene ARF15 in tfs1 was much higher than that in 4266 at regular/high temperature, but the difference was reduced at low temperature (Fig. 4a). In addition, we found that TAS3-derived siRNAs were more abundant in tfs1 under low temperature, and some siRNAs reached levels similar to those in 4266 as revealed by small RNA sequencing (Supplementary information, Fig. S7a, b and Table S6). The overall increase of TAS3-derived siRNAs in the mutant at low temperature was indicative of higher miR390-mTFS1 activity at low temperature.
a Levels of miR390, miR390*, tasiR-ARF-3, tasiR-ARF-7, and ARF15 RNA as measured by RT-qPCR (n = 3). Small RNA levels shown are relative to U6, and ARF15 levels relative to UBQ5. Data are presented as means ± SD. LT, low temperature. Different letters indicate significant differences according to one-way ANOVA (P < 0.05). b The abundances of miR390, miR390*, and two non-tasiR-ARF siRNAs (derived from TAS3b, Supplementary information, Table S4) in 4266, tfs1, 4266-LT, and tfs1-LT. U6 was the internal control. The numbers represent relative abundance, with the levels in 4266 set at 1. c RISC assembly assays at different temperatures. The assays were performed as in Fig. 3a. The upper panel is a native RNA gel showing double-stranded RNA (ds) and single-stranded RNA (ss). WB shows the immunoprecipitated TFS1 or mTFS1. The numbers represent relative abundance of RISC containing ss miR390, normalized to protein abundance of TFS1 or mTFS1.The relative abundance in TFS1 was set as 1 under each temperature condition. d TAS3 RNA cleavage. The assays were performed as in Supplementary information, Fig. S7d, with RISC formation carried out at the indicated temperatures. The upper panel shows the full-length TAS3 RNA and the 5ʹ cleavage fragment. WB shows protein levels of TFS1 and mTFS1. The numbers represent relative abundances of the 5ʹ cleavage fragment normalized to TFS1 or mTFS1 levels. e Detection of interactions between TFS1-LT (or mTFS1-LT) and SGS3 or RDR6 was performed as in Fig. 3f. The numbers represent relative abundance, with the levels of both TFS1and TFS1-LT set at 1. LT, low temperature. f The abundance of the RDR6 product from TAS3b in 4266 and tfs1 determined by RNA gel blotting under regular/high and low temperatures (LT). The assays were performed as in Fig. 3g. U6 was the internal control. The numbers represent relative abundance, with the levels in 4266 set at 1. All experiments were repeated at least three times with similar results.
To investigate the effects of temperature on RISC assembly, we performed in vitro TFS1 and mTFS1 RISC assembly assays in BYL at three temperatures (20 °C, 25 °C, and 30 °C). The ratios of mTFS1 RISC to TFS1 RISC were higher at both 20 °C and 25 °C than at 30 °C (Fig. 4c; Supplementary information, Fig. S7c), suggesting that lower temperatures alleviated RISC formation defects in mTFS1. The slicing activities were too low at 20 °C for quantification, but the relative slicing activities of mTFS1- and TFS1-RISCs were similar to their relative levels at 25 °C and 30 °C (Fig. 4d; Supplementary information, Fig. S7d).
Since we had shown that RDR6 recruitment, rather than slicing was the limiting factor for tasiR-ARF biogenesis in tfs1 at regular/high temperature, we further examined the recruitment of RDR6 by mTFS1 at low temperature. Co-IP showed that, compared to TFS1, mTFS1 was better at precipitating RDR6 at low temperature than at regular/high temperature (Fig. 4e). The antisense RNA product of RDR6 was at much higher levels in tfs1 at low temperature than at regular/high temperature (Fig. 4f). Therefore, we speculate that better RDR6 recruitment by mTFS1 RISCs together with improvements in mTFS1 RISC formation accounts for increased tasiR-ARF biogenesis under low temperature.
mTFS1 allows for fully mechanized hybrid rice breeding
To examine whether other ago7 mutations in linker 2 tend to cause thermo-sensitive female sterility without disrupting the shoot apical meristem, a mutant (ago7-5) with the deletion of the 605th amino acid (serine) in linker 2 was obtained by CRISPR/Cas9. The ago7-5 plants were also sterile with open hulls, resembling the phenotype of tfs1 under regular/high temperature (Fig. 5a). The molecular changes of ago7-5 were similar to those observed in the tfs1 plants under regular/high temperature (Fig. 5b; Supplementary information, Fig. S8a). However, low temperature did not restore fertility to ago7-5, which was consistent with the molecular changes observed under low temperature (Fig. 5b; Supplementary information, Fig. S8a). Different day lengths also failed to restore fertility to ago7-5 plants Supplementary information, Table S2. Besides, we constructed a Leu603Ala mutation in AGO7 in one step (see Materials and Methods) in the 4266 background, and the homozygous mutants were fertile and morphologically identical to 4266 under regular/high temperature (Fig. S8b). Taken together, among three linker 2 mutants, the Leu603Ile natural allele is unique in causing temperature-sensitive female sterility.
a Panicles of ago7-5 grown under regular/high temperature. Scale bars, 5 mm. b The abundance of miR390, miR390*, and two non-tasiR-ARF siRNAs from TAS3b in Nip, ago7-5, Nip-LT and ago7-5-LT. U6 was the internal control. The numbers represent relative abundance, with the levels in Nip set at 1. LT, low temperature (23 °C). All experiments were repeated at least three times with similar results. Panicles from transgenic plants carrying mTFS1 in MH63 (c) and Nip (d) under regular/high and low temperature (LT). Scale bars, 2 cm. e Diagram showing the advantages of using the female-sterile line in hybrid seed breeding under regular/high temperature. Seeds of the female-sterile restorer line were either planted separately from those of the male-sterile line as in traditional hybrid breeding (left part) or mixed with seeds of the male-sterile line at a ratio of 1:6 prior to planting (right part). The lack of self-seed production from the restorer line (female-sterile lines) enables direct harvesting of hybrid seeds from the field.
To test whether the tfs1 point mutation can cause the temperature-responsive sterility trait in both indica and japonica subspecies, a prerequisite for its application in hybrid seed production, we introduced mTFS1 through genetic introgression into Minghui63 (MH63) and Nip (Supplementary information, Fig. S9a). Individuals carrying the tfs1 mutation among the progeny of both genetic introgressions were sterile, with abnormal hulls but normal pollen viability (Supplementary information, Fig. S9b, c). Furthermore, the fertility was partially restored when the plants were grown at low temperature (23 °C) (Supplementary information, Fig. S9d).
As introgression takes time, we developed a one-step approach to create thermo-sensitive female-sterile lines using genome editing. The strategy was to knock out the endogenous TFS1 gene and simultaneously introduce the mutant gene (mTFS1) into any rice cultivar by a vector containing both knock-out and knock-in elements (see Materials and Methods). MH63 and Nip were transformed with the engineered plasmid (Supplementary information, Fig. S9e). Female-sterile individuals resembling tfs1 were found among the transgenic lines (Fig. 5c, d; Supplementary information, Table S7) at reasonable frequencies in the T0 generation (13/50 in MH63, 6/24 in Nip). These transgenic plants had normal pistils and pollen viability (Supplementary information, Fig. S9b, c). Female fertility was also partially recovered when the sterile lines were transferred to low temperature (23 °C) (Fig. 5c, d; Supplementary information, Table S7). The introgression and genome editing experiments together revealed that the thermo-sensitive female sterility trait was manifested in both indica and japonica rice varieties.
In a traditional hybrid seed production scheme, the wild-type restorer plants need to be grown separately from the male-sterile plants such that they are removed from the field prior to large-scale mechanized harvesting of hybrid seeds (Fig. 5e, left). In contrast, if a female-sterile line is used as the restorer following traditional breeding method (Fig. 5e, left), there is no need to remove the restore line because of the lack of self-seed production from the restorer line. Besides, seeds of male- and female-sterile lines can be mixed at a certain ratio for mechanized planting in the field (Fig. 5e, right), which also enables direct harvest of hybrid seeds from the field. To further test the utility of tfs1 in mechanized hybrid rice production, field trials were carried out under regular/high temperature during the summer growing season in Hong Kong and Hunan, China (Figs. 5e and 6). Three commercial, thermo-sensitive male-sterile lines (4155s, C815s, and Y58s) used in two-line systems, and a male-sterile line Huachuan utilized in three-line systems, were used as the female parents. mTFS1/MH63, mTFS1/Nip, and tfs1 served separately as the pollen donors for C815s and 4155s in field trials in Hong Kong and tfs1 served as the pollen donor for Y58s and Huachuan in field trials in Hunan (Fig. 6). In these field trials, seeds of the female-sterile restorer line were either planted separately from those of the male-sterile line as in traditional hybrid breeding (Fig. 5e, left) or mixed with seeds of the male-sterile line at a ratio of 1:6 prior to planting (Fig. 5e, right). In the Hong Kong field trials, hybrid panicles exhibited more than 30% seed set (Supplementary information, Table S8). In both cases performed in Hunan, more than 40% seed set was found in the hybrid panicles (Supplementary information, Table S8). Moreover, hybrid seeds of the field experiment (F1 seeds) were able to produce fully filled panicles, indicating the potential of tfs1 in commercial applications (Fig. 6). In order to obtain high seed sets of hybrid panicles for commercial hybrid breeding, more efforts should be invested to improve the receptibility between female and male lines. Taken together, these natural germplasms are applicable to mechanized hybrid rice production.
The top panel: 4155 s (♀) was crossed with tfs1, mTFS1/MH63 and mTFS1/Nip, respectively. Y58s (♀) was crossed with tfs1. Huachuan (♀) was crossed with tfs1. Panicles of the parents and the hybrid panicles are shown. Seed set rate was showed in the top of panicles. Scale bars, 2 cm. The lower panel, parents and hybrids from the cross between C815s (♀) and tfs1(♂). Whole plants and panicles are shown (with enlarged figures on the right). ♀, female parental line; ♂, male parental line.
Discussion
We identified the first spontaneous allele tfs1 conferring thermo-sensitive female sterility in rice, that is, female sterility under regular/high temperature and female fertility under low temperature. Unlike other rice ago7 mutants24,25 with severe defects in vegetative growth, tfs1 is normal in vegetative development as well as reproductive organ development and is male fertile, which satisfies the requirements for serving as a male parent. Pollen tube growth was blocked after reaching the micropyle in tfs1 plant, suggesting that AGO7 is required for the regulation of pollen tube growth in the maternal parent. Notably, tfs1 can be introduced into other elite rice cultivars. Field trials further demonstrated its utility in fully mechanized hybrid rice breeding, including mechanization at both transplanting and harvesting stages.
AGO7 together with its associated miR390 mediates the cleavage of TAS3 transcripts, with the 5′ cleavage products subsequently converted to dsRNA by RDR6 for tasiR-ARF biogenesis. In tfs1, the tasiR-ARF biogenesis defect correlates with female sterility: at regular/high temperature, reduced ability to recruit RDR6 by mTFS1 causes decreased tasiR-ARF levels, and the mutant plants are female sterile; at low temperature, tasiR-ARF biogenesis is partially restored and so is female fertility (Fig. 7). The reduction in tasiR-ARF biogenesis likely underlies the female sterility of tfs1, as we show that a weak mir390 allele generated by CRISPR/Cas9 also exhibits female sterility.
AGO7 together with its associated miR390 mediates the cleavage of TAS3 transcripts, and recruits SGS3 and RDR6 to the 5′ cleavage product. RDR6 then converts TAS3 transcripts into dsRNA, which is processed by Dicer to produce siRNAs including tasiR-ARFs. In tfs1, despite a reduced efficiency in RISC formation, the higher stability of mTFS1 results in a higher level of RISC. The mTFS1 RISC can cleave TAS3 RNA but cannot efficiently recruit RDR6, resulting in a tasiR-ARF biogenesis defect. The tasiR-ARF biogenesis defect correlates with female sterility: at regular/high temperature, reduced ability of recruiting RDR6 by mTFS1 causes decreased tasiR-ARF levels, and the mutant plants are female sterile; at low temperature, tasiR-ARF biogenesis is partially restored and so is female fertility.
Previous studies showed that highly complementary targets induce the degradation of miRISCs. For example, human AGO2 ubiquitylation followed by degradation leads to the turnover of associated miRNAs.34,35 There was also evidence for such a mechanism in plants that the ubiquitin ligase HAWAIIAN SKIRT is required for target-directed miRNA degradation.36,37 The higher stability of mTFS1 correlated with the slower turnover of miR390, suggesting that the pre- or mature RISCs of mTFS1 are more stable than those of TFS1. However, the higher stability of the mTFS1 RISCs may not be related to target-induced degradation, as mTFS1 RISCs are normal in slicer activity while target-directed miRNA degradation in plants often involves non-cleavable targets.38,39
Our results indicate that the tasiR-ARF biogenesis defect in tfs1 is likely due to compromised RDR6 but not SGS3 recruitment by mTFS1, resulting in reduced levels of antisense TAS3 RNA produced by RDR6 and, consequently, lower levels of the dsRNA precursors of siRNAs. How the Leu603Ile mutation in AGO7 compromises RDR6 recruitment in a temperature dependent manner requires further investigation. A recent study showed that the release of AGO7 RISC from TAS3 precursors is important for dsRNA biogenesis.10 It is possible that increased stability of mTFS1 causes mTFS1-RISC to be slower at dissociation from TAS3 precursors, leading to reduced levels of dsRNA precursors to siRNAs. It was shown that AGO7 RISC recruits RDR6 in part by stalling ribosomes.9 It is also possible that the mutant RISC adopts a different conformation that compromises ribosome stalling and hence RDR6 recruitment.
AGO proteins contain N, PAZ, MID, and PIWI domains and form an N-PAZ and MID-PIWI bi-lobal structure.40,41,42,43,44,45 AGO RISC is thought to undergo substantial conformational changes during target RNA/DNA recognition, including open-close and torsional movements of the two lobes.46,47,48 Linker 2, which is located between the two lobes, participates in RISC targeting by interacting with the guide‒target duplex.42,49 We show that two residues, Leu603 and Ser605, are required for female fertility in rice but dispensable for other developmental processes that require AGO7 while a Leu603Ala substitution does not cause any phenotypes. The lack of strong developmental defects probably reflects the weak effects of these mutations in the biogenesis of tasiR-ARFs. Given that linker 2 probably participates in miRNA‒target interactions, it is possible that the Leu603Ile substitution weakens the miR390‒TAS3 interaction, leading to reduced RDR6 recruitment. The Leu603Ile substitution leads to even weaker effects at low temperature, resulting in thermo-sensitive female sterility.
Materials and Methods
Plant strains and growth conditions
4266 is a hybrid variety containing 70% indica and 30% japonica backgrounds. tfs1 is a spontaneous mutant derived from 4266. The rice plants examined under natural field conditions (> 23.5 °C) were grown at the Baiyun Base of Guangdong Academy of Agricultural Sciences, Guangzhou, China and the gene garden area of The Chinese University of Hong Kong, Hong Kong, China. For low temperature conditions (23 °C), both 4266 and tfs1 were grown in the greenhouse and growth chambers (with a relative humidity 75%) at The Chinese University of Hong Kong, Hong Kong, China. The small-scale field trial was performed under regular/high temperature (> 25 °C) during the summer growing seasons in Hong Kong and Changsha, Hunan, respectively. The field trials were conducted according to the two diagrams in Fig. 5e: (1) regular planting according to the current agricultural practice of planting 2 rows of the male parent with every 12 rows of the female parent (Fig. 5e, the left panel); (2) mixed planting, with the ratios of the female line to the male line being 6–7: 1 (Fig. 5e, the right panel). The temperatures during rice growing seasons in the field are listed in Supplementary information, Table S9.
Pollen viability determination
Florets were harvested in the morning, about 2 h before anthesis. The pollen grains were stained by a 1% I2-KI solution50 and observed under a light microscope.
Embryo sac and embryo development observation
Fixation: Pistils (2 h before anthesis) were fixed in 2.5% glutaraldehyde in PBS (pH 7.2) for at least 24 h at 4 °C, then washed by PBS three times. The pistils were dehydrated sequentially in 30%, 50%, and 70% ethanol, each with 30 min incubation. The pistils were dissected in 70% ethanol under a stereoscope to remove the stigma.
Staining: Samples were incubated sequentially in 50%, 30% ethanol, distilled water, and 2% aluminum potassium sulfate, each for 20 min. The dye used for staining was 10 mg/L Eosin B dissolved in 4% sucrose and staining was done overnight at room temperature. The samples were washed with 2% aluminum potassium sulfate once and with distilled water twice and then dehydrated in an increasing ethanol series from 30% to 90%. After that, the samples were incubated in 50% methyl salicylate in ethanol for 1 h and then placed in pure methyl salicylate for clearing.
Embryo sac imaging: The samples were observed by confocal microscopy (Olympus FV1000 IX81-SIM) with excitation wavelengths between 550‒630 nm.51
Pollen germination and pollen tube growth observation
Panicles containing stems were harvested in the morning, about 2 h before anthesis, and were kept with the stems submerged in water. Some of the florets at the middle of the inflorescence were emasculated without damaging the stigma, while other florets were left alone. When the top florets reached anthesis, florets above the emasculated florets were shaken to release pollen and the emasculated florets were quickly examined by a stereoscope to identify ones with pollen on the stigma. The pollinated pistils were sampled at 5 min, 30 min, and 1 h and fixed in ethanol and acetic acid (mixed 3:1) for 30 min. The pistils were then immersed in 1 M KOH overnight for softening. The samples were washed with distilled water three times and stained with 0.1% aniline blue dissolved in K3PO4 buffer (pH 8.5) for 2 h. The samples were washed with distilled water three times and immersed in 50% glycerol diluted in K3PO4 buffer (pH 8.5). A single pistil was placed on a slide and sealed by a cover slip, which was gently pressed to mount the pistil in glycerol. The pistil was then observed under a fluorescence microscope (Nikon E80i) according to previous studies.51,52,53
Floral primordia development observation by SEM
Young panicles of 1 mm to 2 mm in length were harvested and fixed in 2.5% glutaraldehyde in PBS (pH 7.2) for 24 h at 4 °C, then washed with PBS three times. The samples were dehydrated in an ethanol series and dried in tetramethylsilane at 28 °C for 1 h. The samples were then sputter-coated with platinum and observed by a scanning electron microscope (Hitachi S-4000, Tokyo) at an accelerating voltage of 15 kV.
Map-based cloning
An F2 population from the cross between tfs1 and Nip was used for map-based cloning. Bulk segregant analysis was first performed in two F2 pools, F (fertile) and S (sterile), with 40 individuals in each pool for primary mapping. 20,000 more individuals were used for fine mapping. 81 Simple Sequence Repeats (SSR) markers distributed on the 12 chromosomes were used for the primary mapping and more SSR markers around the candidate mutation position were developed based on the reference DNA sequence of Nipponbare (japonica, MSU7.0) and 9311 (indica, ASM465v1) for fine mapping.
MutMap analysis
The F2 population used for MutMap analysis was generated from the cross between tfs1 and 4266. DNA from individuals showing sterility and abnormal hull in the F2 population was separately isolated. DNA from 30 individuals was equally mixed to generate a pool, which was subjected to library construction and sequencing on the Illumina High-seq 4000 platform (Annoroad, Beijing). Three such pools of 30 individuals were separately sequenced together with 4266 and tfs1 plants as controls.54 Sequencing of each pool produced about 104 million clean reads, which represented approximately 30× coverage of the genome. Sequencing of tfs1 and 4266 resulted in 193 and 160 million clean reads, respectively. Reads were aligned to the Nipponbare reference genome (MSU7.0) using the software BWA (version 0.7.0) with default settings. SNP sites between the pool and the wild-type parent were detected using SAMtools (version 0.0.19). After filtering for sequencing quality and depth of high confident reads, 527 SNPs between tfs1 and 4266 were retained for MutMap analysis. The SNP index was calculated for each SNP to show the extent the SNP was linked to the target gene.28 SNPs unlinked to the target gene are expected to segregate randomly such that the ratio of the tfs1 type to the 4266 type in the 3 pools of F2 would be 1:1, resulting in an SNP index of 0.5. An SNP closely linked to the target gene should be shared by all the F2 individuals, thus the SNP index is expected to be around 1. All SNPs with a SNP index over 0.9 were annotated by snpEff (version 3.6).55
Generation of transgenic plants
The TFS1 gene was amplified using primers TFS1-1F, TFS1-2F, TFS1-2Freverse, and TFS1-1R from genomic DNA isolated from 4266. The resulting 9.7 kb fragment included 4.7 kb upstream of the start codon, the coding region, and 0.8 kb downstream of the stop codon (Supplementary information, Fig. S2c). The fragment was cloned into pCAMBIA1300 (Waryong VECT0070) between BamHI (5ʹ) and SalI (3ʹ) sites by the In-Fusion method (ClonExpress one-step cloning kit, Vazyme) and verified by sequencing. Both the complementation plasmid and the empty vector were introduced into tfs1 via Agrobacterium-mediated genetic transformation. Primer sequences are listed in Supplementary information, Table S10.
To generate HA-TFS1 or HA-mTFS1 lines, the TFS1:HA-TFS1 and TFS1:HA-mTFS1 plasmids were first constructed and then used to transform 4266 and tfs1, respectively. The plasmids each contained 4.7 kb upstream of the start codon, 4.1 kb coding region, and 0.8 kb downstream of the stop codon. Primers TFS1-HA-1F and TFS1-HA-1R were used to amplify the 4.7 kb fragment; primers TFS1-HA-2F and TFS1-HA-2R were used for 3× HA amplification; primers TFS1-HA-3F and TFS1-HA-3R were used to amplify the 4.1 kb coding region together with the 0.8 kb 3ʹ region from either 4266 or tfs1. The fragments were cloned into pCAMBIA1300 between the KpnI (5ʹ) and HindIII (3ʹ) sites by the In-Fusion method and verified by sequencing. Primer sequences are listed in Supplementary information, Table S10.
For CRISPR/Cas9-based editing of the linker 2 region, a guide RNA corresponding to the 80th–99th nt (5ʹ- AAATTAAAGCTTGGCAGCAG -3ʹ) of the third exon of TFS1 was used. The guide RNA sequence was cloned into BGK03056 downstream of the OsU6 promoter as follows. Two pairs of primers (CRISPR-F/CRISPR-SGR and CRISPR-SGF/CRISPR-R) were designed to amplify one region of BGK030 (between the HindIII (5ʹ) and AarI (3ʹ) sites) into two small fragments, both containing the guide RNA sequence. Afterwards, the two fragments were cloned to BGK030 (between the HindIII (5ʹ) and AarI (3ʹ) sites) by the In-Fusion method to replace the 20 bp sequence of the empty vector with that of the guide RNA sequence. The clone was verified by sequencing. Primer sequences are listed in Supplementary information, Table S10.
For CRISPR/Cas9-based editing of MIR390 to create small deletions, three 20 bp guide RNAs (5ʹ- GCTATCCCTCCTGAGCTTCAAGG -3ʹ; 5ʹ-ACAATCCTTGAAGCTCAGGAGGG-3ʹ; 5ʹ-TGGAACAATCCTTGAAGCTCAGG-3ʹ) targeting the miR390-5P region were cloned in one transformation vector as follows. Two pairs of primers (OsU3-F/SG1R and SG1F/OsU3-mR) were used to amplify two fragments, then the two fragments were used as templates to obtain sequence 1 (OsU3-guide RNA1) by overlapping PCR. Then primers OsU3-mF/SG2R and SG2F/OsU3-mR, and primers OsU3-mF/SG3R and SG3F/OsU3-R were used to obtain sequences 2 (OsU3-guide RNA2) and 3 (OsU3-guide RNA3), respectively, as for sequence 1. Sequence 1 and sequence 2 were then used as templates to obtain sequence 1 + 2 by overlapping PCR. Sequence 1 + 2 and sequence 3 were used as templates to produce sequence 1 + 2 + 3. The OsU3 guide RNA fragment was further cloned into pCAMBIA1390-OsNLSCas9 between PacI and HindIII with T4 DNA ligase (M0202L, NEB). The resulting clone was verified by sequencing. Primer sequences are listed in Supplementary information, Table S10.
For CRISPR/Cas9-based editing of MIR390, a 20 bp guide RNA (5ʹ- GCTATCCCTCCTGAGCTTCA -3ʹ) targeting the miR390-5P region was used. Two pairs of primers (OsU3-F/MIR390-SGR and MIR390-SGF/OsU3-R) were designed to amplify two fragments, one containing OsU3 and the 20 bp miR390-5P guide RNA sequence, and the other containing the miR390-5P guide RNA sequence and the stem-loop sequence, and then the two fragments were assembled to an OsU3-miR390-sgRNA fragment by overlapping PCR. The OsU3-miR390-sgRNA fragment was further cloned into pCAMBIA1390-OsNLSCas957 as described in the previous paragraph. Primer sequences are listed in Supplementary information, Table S10.
The plasmid used for the one-step engineering of female sterility contained two units: (1) the CAS9 and a guide RNA (targeting the 341st‒360th nt (5ʹ- GGGTGGCTTGCCGTAGAACG -3ʹ) of the first exon of TFS1), and (2) a 9.7 kb genomic fragment (pTFS1:mTFS1gDNA) with the tfs1 mutation as well as a modification at the PAM (338th‒340th nt, (CCA) to (CAA)) in the first exon without changing the amino acid. The PAM modification was meant to protect the mTFS1 transgene from the guide RNA (Supplementary information, Fig. S9E). This constructed plasmid, which can knock out the endogenous TFS1 and simultaneously introduce the mTFS1 sequence, was used for both japonica and indica rice transformation.
RNA isolation and quantification
RNA used for RT-qPCR and cDNA amplification was isolated from young panicles using Plant RNA Extraction Kit (Qiagen), and DNA was removed with the RNase-free DNase Set (Qiagen). Reverse transcription was performed with RevertAid First Strand cDNA Synthesis Kit (Thermo). RT-qPCR analysis was performed using iQ™ SYBR® Green Supermix (Bio-Rad) with the CFX Real-Time PCR Detection system (Bio-Rad). Three technical repeats were included. U6 or UBQ5 served as the internal standard. Primers used are listed in Supplementary information, Table S11.
TFS1 protein localization in rice protoplasts
Protoplasts were prepared according to reported methods58 with modifications. 8-day-old rice seedlings were used for protoplast isolation. Stem and leaf sheath tissues were cut into approximately 0.5 mm strips, which were immediately transferred into enzymatic digestion buffer (20 mM MES, pH 5.7, 10 mM CaCl2, 0.4 M Mannitol, 1.5% Cellulase RS, 0.4% Macerozyme R-10, 0.1% BSA, 20 mM KCl) and incubated in the dark with gentle shaking for 4 h. The protoplasts were collected by filtration through 40 μm nylon meshes and centrifugation for 10 min at 100 × g. The W5 solution (2 mM MES, pH 5.7, 5 mM KCl, 5 mM NaCl, 125 mM CaCl2) was added to resuspend the protoplasts. The protoplasts were centrifuged at 100 × g and resuspended in MMG solution (4 mM MES, pH 5.7, 0.4 M Mannitol, 15 mM MgCl2).
The open reading frames of TFS1 and mTFS1 were amplified and then cloned into pENTR11 (Thermo fisher A10467) using primers P11-TFS1F and P11-TFS1R (Supplementary information, Table S10). LR recombination reaction using the Gateway system was performed with the binary vector of pHW-EGFP-avi to result in the fusion with GFP at the N-terminus of TFS1 which was driven by a CaMV 35 S promoter. Plasmids were introduced into protoplasts by PEG (40% PEG 4000, 0.2 M Mannitol, 0.1 M CaCl2)-mediated transformation. Briefly, 5 μg plasmid DNA was mixed with 100 μL protoplasts (about 2 × 105 cells). After incubation for 10 min, 440 μL W5 solution was slowly added to the protoplasts, followed by gently inverting the tube. The protoplasts were pelleted by centrifugation at 100 × g for 3 min and gently resuspended in 1 mL WI solution (0.5 M mannitol, 20 mM KCl, 4 mM MES, pH 5.7). Finally, the protoplasts were cultured in the dark at room temperature for 16 h. The protoplasts were observed by confocal microscopy for GFP fluorescence.
Small RNA sequencing and analysis
Young panicles sampled with biological replicates were used for total RNA extraction. Small RNAs of 18 to 30 nt were isolated from 1 μg total RNA subjected to size fractionation by gel electrophoresis. Library preparation and sequencing (BGISEQ-500RS Illumina platform) were performed at BGI, Hong Kong. More than 50 million clean reads were obtained for each sample. The raw reads were trimmed to remove adaptor sequences and filtered to obtain clean tags between 18 nt and 30 nt long.59 The reads were mapped to the rice reference genome (MSU7).60,61 The small RNA levels were calculated as RPM (Reads Per Million).62 Annotated miRNAs were extracted from miRBase version 21 (http://www.mirbase.org). Levels of phasiRNAs were quantified by mapping reads from our small RNA sequencing data to PHAS loci identified previously.33 Similarly, levels of RDR6-dependent phasiRNAs were quantified by mapping reads from our small RNA sequencing data to PHAS loci showing reduced phasiRNA levels in an osrdr6-2 weak allele as previously reported.33
In situ hybridization
Young panicles, pistils and anthers were fixed in 4% PFA (4% paraformaldehyde, 0.1% Tween-20, 0.1% Triton X-100) for 24 h at 4 °C. The samples were dehydrated sequentially in 30%–100% ethanol at 10% increments in ethanol, each with 1 h incubation. The tissue was then immersed in 25%, 50%, 75%, and 100% xylene with 1 h incubation in each. Samples were then immersed in molten paraffin for 3 days with changes of fresh molten paraffin twice a day. After that, the samples were embedded in paraffin at 60 °C. The paraffin blocks were cut to 8 µm sections with a rotary microtome. The sections were floated on 42 °C water and mounted onto slides. The slides were dried overnight in a 42 °C oven. Samples were then dewaxed by immersing the slides in 100% xylene twice, each with 10 min incubation. After sample dewaxing, slides were rehydrated by an ethanol series (95%, 85%, 50%, and 30%, each for 1 min). After washes with PBS, proteinase K was added, followed by incubation for 20 min at 37 °C. Then, slides were incubated in 0.1 M glycine for 2 min and washed twice with PBS. The samples were fixed by 4% PFA for 10 min, and then washed with PBS twice. The slides were put back to the box and treated with acetic anhydride followed by a PBS wash. Slides were dehydrated by an ethanol series (30%, 50%, 85%, 95%, and 100%), wrapped in paper towel and left on the bench to dry.
Fluorescence in situ hybridization
Dry slides were put in a dark box, and 0.1 μM FAM-labeled probe (Supplementary information, Table S12) was added onto the slides, which were incubated at 55 °C for 48 h, then washed with SSC buffer (1× SSC, pH 4.5, 50% Formamide) three times at 55 °C for 20 min each time. After five washes with PBS, the samples were stained with DAPI and washed three times with PBS. Finally, the slides were observed under a fluorescence microscope (Leica DM2000-MC170).
DIG in situ hybridization
Dry slides were put in a dark box, and 0.1 μM DIG-labeled probe (Supplementary information, Table S12) was added onto the slides, which were incubated at 55 °C for 48 h and then washed with 0.1× SSC twice at 55 °C. After washed with Buffer 1 (0.1 M Tris-HCl pH 7.5, 0.15 M NaCl), 2% Boheringer blocking reagent and 1% BSA dissolved in Buffer1 were used for blocking for 30 min respectively. Anti-digoxigenin-alkaline phosphatase (Roche, 11093274910) diluted 3000 times in 1% BSA dissolved in Buffer 1 was used to soak slides for 1.5 h with gentle shaking. Slides were washed with 1% BSA (4 times of 20 min each), Bufer 1 (5 min) and Buffer 2 (0.1 M Tris-HCl, pH 9.5, 0.1 M NaCl, 50 mM MgCl2). The slides were placed in a clean box, and NBT/BCIP made with fast tablets (Sigma, 085H-8909) were applied on the slides, which were kept in the dark for 48 h. To stop the enzymatic reaction and remove background signals, slides were washed with distilled water, 70%, 95%, 100%, 95%, 70% ethanol, and DEPC water in this sequence. Slides were then observed under a microscope (Leica DM2000-MC170).
AGO7 IP and small RNA sequencing
TFS1 and mTFS1 IPs were performed with pTFS1:HA-TFS1/4266 and pTFS1:HA-mTFS1/tfs1 plants, respectively, and the corresponding control IPs were conducted from nontransgenic plants 4266 and tfs1. 0.5 g young panicles sampled with two biological replicates were fully ground and mixed well with 1.5 mL IP buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 0.1% NP40, 4 mM MgCl2, 5 mM DTT, Protease inhibitor, RNase inhibitor), followed by incubation of the suspension on ice for 20 min. The supernatant was collected after centrifugation at 5000× g transferred to a 1.5 mL Eppendorf tube and centrifuged at 12,000× g to further remove any debris. 5 μg HA antibody (sigma, H6908) and protein A Dynabeads (Invitrogen, 10002D) were sequentially added to the supernatant to immunoprecipitate the AGO7 protein. RNA was extracted from the above IP samples for small RNA sequencing. The 4266 control IP and tfs1 control IP served as controls for the TFS1 IP and mTFS1 IP, respectively. RNA extraction, library construction, sequencing, and data analysis were performed in two independent experiments as described above for total small RNAs.
RNA gel blotting analysis
The RNA gel blotting analysis was performed as follows. 10 μg total RNA (60 μg total RNA for miR390* detection) was resolved in a 15% urea-PAGE gel, and RNA was then transferred to a Hybond NX membrane. The EDC cross-linking buffer (0.16M EDC, 0.13M 1-methylimidazole, pH 8.0) was used to crosslink RNA to the membrane at 65 °C for 90 min. Membrane was washed with DEPC water three times and dried at 65 °C for 30 min. Then, the membrane was incubated in hybridization buffer (sigma, H7033) containing a biotin-labeled probe (see Supplementary information, Table S12 for the sequences of probes) at 55 °C overnight (about 16 h). After two washes (2× SSC, 0.1% SDS, 55 °C, 20 min each time), the membrane was processed using the Chemiluminescent Nucleic Acid Detection Module Kit (ThermoFisher, 89880) according to the instruction manual. The relative gray values of the RNA signals were calculated by Image J.
TFS1 and mTFS1 RISC assembly and slicer activity assays
An evacuolated protoplast extract was prepared from suspension-cultured tobacco BY-2 cells,63 and was centrifuged to remove membranes. The resulting membrane-depleted extract, called “membrane-depleted BYL” in the previous publication,64 was named “BYL” in this study. BYL was used for in vitro translation and RISC assembly. The method of using BYL for RISC assembly has been demonstrated with reproducibility.65
TFS1 and mTFS1 coding regions were cloned into the pJET1.2 blunt vector (ThermoFisher, K1231), such that they were fused to 3× FLAG at the N-termini. XbaI-linearized plasmids served as templates for in vitro transcription using the AmpliCap SP6 High Yield Message Maker Kit (CELLSCRIPT) to produce FLAG-TFS1 and FLAG-mTFS1 mRNAs. In vitro translation and miR390/miR390* loading were performed using BYL as previously described66(miR390 and miR390* sequences are listed in Supplementary information, Table S12). The translation reaction was terminated by adding one fiftieth volume of 10 mM puromycin. Primer sequences are listed in Supplementary information, Table S10.
Following RISC assembly, the FLAG-TFS1 and FLAG-mTFS1 proteins and their associated miRNAs were analyzed as follows. The RISC assembly reaction (20 μL) was mixed with 0.6 μg anti-FLAG M2 antibody (Sigma-Aldrich F1804) and 9.4 μL of the translation reaction (TR) buffer (30 mM HEPES, pH 7.4, 80 mM KOAc, 1.8 mM MgCl2, 2 mM DTT, protease inhibitor) and incubated for 40 min on ice, followed by further incubation with 0.18 mg of protein G-conjugated Dynabeads (Invitrogen 10003D) in 36 μL TR buffer for 20 min on ice. The magnetic beads were washed three times with 100 μL TR buffer. One-tenth of the magnetic beads was used for WB analysis. The remaining magnetic beads were mixed with 10 μL of annealing buffer (50 mM Tris-HCl, pH 7.5, 100 mM KCl and 5 mM MgCl2) and 10 μL TE-saturated phenol. After centrifugation, the aqueous phase was mixed with one-fourth of native dye solution (2.5× TBE, 0.5 mg/mL bromophenol blue, 0.5 mg/mL xylene cyanol, 25% glycerol) and analyzed with native 15% PAGE.
For slicer activity assays, TAS3b RNA was prepared as described.66 The TAS3b template for in vitro transcription was prepared by PCR using the primers TAS3b-1F and TAS3b-1R. In vitro transcription for producing the TAS3b sense strand probe was carried out with SP6 RNA polymerase in the presence of [α-32P]CTP. Poly-adenylated 32P-labeled TAS3b RNAs were purified using Dynabeads Oligo (dT)25 (Invitrogen 61005). After miR390-AGO7 RISC formation, the 32P-labeled TAS3b RNA was added into the reaction mixture and incubated for 20 min. 5 μL of the reaction mixture was mixed with 15 μL of TE buffer and 20 μL of TE-saturated phenol. After centrifugation, the aqueous phase was mixed with 1.6 volume of urea dye mix (12.5 M urea, 0.2 mg/mL bromophenol blue, and 0.2 mg/mL xylene cyanol) and analyzed with 7 M urea/5% PAGE. Radiolabeled products were detected using the Typhoon FLA 7000 imaging system (GE Healthcare).
For the experiments to examine RISC formation at different temperatures in Fig. 4C, FLAG-TFS1 and FLAG-mTFS1 proteins were translated at 25 °C and RISC formation was carried out at the indicated temperatures (20 °C, 25 °C, and 30 °C). FLAG-TFS1 and FLAG-mTFS1 proteins were detected by WB analysis using anti-FLAG antibody and signals were visualized with the LAS-3000 (FUJIFILM) instrument.
Protein half-life measurements
Rice protoplasts isolation and transformation were performed as described in the protein localization section. Protoplasts were counted with a hemocytometer, and the cell concentration was brought to 107 cells/mL with the addition of MMG solution (4 mM MES, pH 5.7, 0.4 M Mannitol, 15 mM MgCl2). A total of 20 ml protoplasts was transformed with plasmids of 35 S:GFP-TFS1 and 35 S:GFP-mTFS1, respectively. At 16 h after transfection, cycloheximide (50 μg/mL) and chloramphenicol (50 μg/mL) were added to the transformed protoplasts to inhibit protein synthesis. 4 mL of protoplasts of each sample was then collected at 0 h, 12 h, 24 h, 36 h, and 48 h. The 4 mL of collected protoplasts was split into two parts. 2 mL was used for TFS1 and mTFS1 protein detection by WB analysis using anti-GFP antibody (Takara, 632381). The other 2 mL was for IP using anti-GFP antibody followed by RNA extraction to determine the abundance of miR390 by RNA gel blot analysis. RDR6 protein half-life measurement was performed with the anti-RDR6 antibody without protoplast transformation.
Slicer activity assays with immunoprecipitated TFS1 and mTFS1
The TFS1 and mTFS1 proteins were immunoprecipitated from transgenic lines TFS1: HA-mTFS1/tfs1 and TFS1: HA-TFS1/4266 as described above. The RNA substrates were TAS3b (wild-type RNA) or TAS3b-5ʹ3ʹm (negative control), which contained mutations in both miR390 binding sites. In vitro transcription for producing the TAS3b sense strand was carried out with SP6 RNA polymerase (Invitrogen MAXIscript kit) and the band corresponding to the expected TAS3b full-length transcript was excised from 7 M urea/5% PAGE. The beads containing HA-TFS1 and HA-mTFS1 were mixed with the TAS3b or TAS3b-5ʹ3ʹm transcript in reaction buffer (1 mM ATP, 0.2 mM GTP, 1.2 mM MgCl2, 25 mM creatine phosphate, 20 mg/mL creatine kinase and 0.4 U/mL RNase Inhibitor). The reaction mix was incubated at room temperature for 2 h, RNA was then extracted and resolved in 7 M urea/5% PAGE followed by RNA gel blot analysis as described above. A biotin-labeled probe for the hybridization was produced as follows. The template DNA for the probe was prepared by PCR using the primers TAS3b-3F and TAS3b-3R. In vitro transcription was carried out with T7 RNA polymerase (Invitrogen MAXIscript kit) in the presence of biotin-UTP (biotin-UTP: UTP = 1:1) to produce a biotin-labeled TAS3b antisense RNA. Primers used are listed in Supplementary information, Table S13.
RNA gel blotting analysis to detect the RDR6 product
40 ug of total RNA from 4266 and tfs1 was dissolved in RNase-free water. Biotin-labeled RNAs were produced by in vitro transcription to serve as RNA size markers and/or the probe as described below (Fig. 3g). One template was prepared by PCR using the primers TAS3b-2F and TAS3b-5R. In vitro transcription was carried out with SP6 RNA polymerase in the presence of biotin-UTP (biotin-UTP: UTP = 1:1) to generate the 609 nt TAS3b sense transcript, which was used as the hybridization probe as well as the size marker for the TAS3b full-length transcript. The negative control RNA was similarly produced by in vitro transcription without using biotin-UTP. Another template, which covers the region between the two miR390 binding sites, was prepared by PCR using primers TAS3b-4F and TAS3b-4R. Then, in vitro transcription was carried out with SP6 RNA polymerase in the presence of biotin-UTP (biotin-UTP: UTP = 1:1) to produce a 268 nt sense RNA, which was used as an RDR6 product size marker. The RNA gel blot assay was performed as described above. Primers used are listed in Supplementary information, Table S13.
Antibodies
Antibodies | SOURCE | IDENTIFIER |
|---|---|---|
Anti-HA, produced in rabbit, affinity isolated antibody | Sigma | Catalog#H6908 |
Anti-GFP, produced in mouse, affinity isolated antibody | Takara | Catalog#632381 |
Anti-Digoxigenin-AP, Fab fragments | Roche | Catalog#11093274910 |
Anti-FLAG, produced in mouse, affinity isolated antibody | Sigma | Catalog#F1804 |
Anti-FLAG (Anti-DYKDDDDK), monoclonal antibody | Wako | Catalog#012-22384 |
Anti-OsSGS3, produced in rabbit, affinity isolated antibody (Two OsSGS3 genes (Os12g09580 and Os12g09590) homologous to AtSGS3) | Agrisera | Catalog#AS153099 |
Anti-OsRDR6, produced in rabbit, affinity isolated antibody | ABclonal | Catalog#A20319 |
Data availability
All DNA sequencing and sRNA sequencing data from this study have been deposited in the National Center for Biotechnology Information (NCBI) with the accession number PRJNA827282. Source data for all graphs have been provided. All other data are available from the corresponding authors upon reasonable request.
References
Khush, G. S. Green revolution: the way forward. Nat. Rev. Genet. 2, 815 (2001).
Yuan, L. P. Hybrid rice technology for food security in the world. Crop Res. 18, 185–186 (2004).
Fei, Q., Xia, R. & Meyers, B. C. Phased, secondary, small interfering RNAs in posttranscriptional regulatory networks. Plant Cell 25, 2400–2415 (2013).
Chen, X. Small RNAs and their roles in plant development. Annu. Rev. Cell Dev. Biol. 25, 21 (2009).
Allen, E., Xie, Z., Gustafson, A. M. & Carrington, J. C. microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 121, 207–221 (2005).
Axtell, M. J., Jan, C., Rajagopalan, R. & Bartel, D. P. A two-hit trigger for siRNA biogenesis in plants. Cell 127, 565–577 (2006).
Montgomery, T. A. et al. Specificity of ARGONAUTE7-miR390 interaction and dual functionality in TAS3 trans-acting siRNA formation. Cell 133, 128–141 (2008).
Peragine, A., Yoshikawa, M., Wu, G., Albrecht, H. L. & Poethig, R. S. SGS3 and SGS2/SDE1/RDR6 are required for juvenile development and the production of trans-acting siRNAs in Arabidopsis. Genes Dev. 18, 2368–2379 (2004).
Iwakawa, H. et al. Ribosome stalling caused by the Argonaute-microRNA-SGS3 complex regulates the production of secondary siRNAs in plants. Cell Rep. 35, 109300 (2021).
Sakurai, Y. et al. Cell-free reconstitution reveals the molecular mechanisms for the initiation of secondary siRNA biogenesis in plants. Proc. Natl. Acad. Sci. USA 118, 2102889118 (2021).
Adenot, X. et al. DRB4-dependent TAS3 trans-acting siRNAs control leaf morphology through AGO7. Curr. Biol. 16, 927–932 (2006).
Garcia, D., Collier, S. A., Byrne, M. E. & Martienssen, R. A. Specification of leaf polarity in Arabidopsis via the trans-Acting siRNA pathway. Curr. Biol. 16, 933–938 (2006).
Xu, L. et al. Genetic interaction between the AS1-AS2 and RDR6-SGS3-AGO7 pathways for leaf morphogenesis. Plant Cell Physiol. 47, 853–863 (2006).
Fahlgren, N. et al. Regulation of AUXIN RESPONSE FACTOR3 by TAS3 ta-siRNA affects developmental timing and patterning in Arabidopsis. Curr. Biol. 16, 939–944 (2006).
Yoshikawa, M., Peragine, A., Park, M. Y. & Poethig, R. S. A pathway for the biogenesis of trans-acting siRNAs in Arabidopsis. Genes Dev. 19, 2164–2175 (2005).
Marin, E. et al. mir390, Arabidopsis TAS3 tasiRNAs, and their AUXIN RESPONSE FACTOR targets define an autoregulatory network quantitatively regulating lateral root growth. Plant Cell 22, 1104–1117 (2010).
Yoon, E. K. et al. Auxin regulation of the microRNA390-dependent transacting small interfering RNA pathway in Arabidopsis lateral root development. Nucleic Acids Res. 38, 1382–1391 (2009).
Hobecker, K. V. et al. The microRNA390/TAS3 pathway mediates symbiotic nodulation and lateral root growth. Plant Physiol. 174, 2469–2486 (2017).
Xia, R., Xu, J. & Meyers, B. C. The emergence, evolution, and diversification of the miR390-TAS3-ARF pathway in land plants. Plant Cell 29, 1232–1247 (2017).
Hunter, C., Sun, H. & Poethig, R. S. The Arabidopsis heterochronic gene ZIPPY is an ARGONAUTE family member. Curr. Biol. 13, 1734–1739 (2003).
Yifhar, T. et al. Failure of the tomato trans-acting short interfering RNA program to regulate AUXIN RESPONSE FACTOR3 and ARF4 underlies the wiry leaf syndrome. Plant Cell 24, 3575–3589 (2012).
Brooks, C., Nekrasov, V., Lipppman, Z. B. & van Eck, J. Efficient gene editing in tomato in the first generation using the clustered regularly interspaced short palindromic repeats/CRISPR-associated9 system. Plant Physiol. 166, 1292–1297 (2014).
Douglas, R. N. et al. ragged seedling2 encodes an ARGONAUTE7-like protein required for mediolateral expansion, but not dorsiventrality, of maize leaves. Plant Cell 22, 1441–1451 (2010).
Itoh, J.-I., Kitano, H., Matsuoka, M. & Nagato, Y. Shoot organization genes regulate shoot apical meristem organization and the pattern of leaf primordium initiation in rice. Plant Cell 12, 2161–2174 (2000).
Nagasaki, H. et al. The small interfering RNA production pathway is required for shoot meristem initiation in rice. Proc. Natl. Acad. Sci. USA 104, 14867–14871 (2007).
Li, S. et al. Natural variation in PTB1 regulates rice seed setting rate by controlling pollen tube growth. Nat. Commun. 4, 2793 (2013).
Zhang, T. et al. LATERAL FLORET 1 induced the three-florets spikelet in rice. Proc. Natl. Acad. Sci. USA 114, 9984–9989 (2017).
Abe, A. et al. Genome sequencing reveals agronomically important loci in rice using MutMap. Nat. Biotechnol. 30, 174 (2012).
Song, X. et al. Rice RNA-dependent RNA polymerase 6 acts in small RNA biogenesis and spikelet development. Plant J. 71, 378–389 (2012).
Liu, B. et al. Oryza sativa Dicer-like4 reveals a key role for small interfering RNA silencing in plant development. Plant Cell 19, 2705–2718 (2007).
Yoshikawa, M. et al. 3′ fragment of miR173-programmed RISC-cleaved RNA is protected from degradation in a complex with RISC and SGS3. Proc. Natl. Acad. Sci. USA 110, 4117–4122 (2013).
Jouannet, V. et al. Cytoplasmic Arabidopsis AGO7 accumulates in membrane-associated siRNA bodies and is required for ta-siRNA biogenesis. EMBO J. 31, 1704–1713 (2012).
Jiang, P. et al. 21-nt phasiRNAs direct target mRNA cleavage in rice male germ cells. Nat. Commun. 11, 5191 (2020).
Han, J. et al. A ubiquitin ligase mediates target-directed microRNA decay independently of tailing and trimming. Science 370, eabc9546 (2020).
Shi, C. Y. et al. The ZSWIM8 ubiquitin ligase mediates target-directed microRNA degradation. Science 370, eabc9359 (2020).
Zhang, X. et al. The Arabidopsis thaliana F-box gene HAWAIIAN SKIRT is a new player in the microRNA pathway. PLoS One 12, e0189788 (2017).
Lang, P. L. M. et al. A role for the F-box protein HAWAIIAN SKIRT in plant microRNA function. Plant Physiol. 176, 730–741 (2018).
Yan, J. et al. Effective small RNA destruction by the expression of a short tandem target mimic in Arabidopsis. Plant Cell 24, 415–427 (2012).
Franco-Zorrilla, J. M. et al. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 39, 1033–1037 (2007).
Kuhn, C.-D. & Joshua-Tor, L. Eukaryotic Argonautes come into focus. Trends Biochem. Sci 38, 263–271 (2013).
Meister, G. Argonaute proteins: functional insights and emerging roles. Nat. Rev. Genet. 14, 447–459 (2013).
Parker, J. S. How to slice: snapshots of Argonaute in action. Silence 1, 3 (2010).
Höck, J. & Meister, G. The Argonaute protein family. Genome Biol 9, 210 (2008).
Hutvagner, G. & Simard, M. J. Argonaute proteins: key players in RNA silencing. Nat. Rev. Mol. Cell Biol. 9, 22–32 (2008).
Boland, A., Huntzinger, E., Schmidt, S., Izaurralde, E. & Weichenrieder, O. Crystal structure of the MID-PIWI lobe of a eukaryotic Argonaute protein. Proc. Natl. Acad. Sci. USA 108, 10466–10471 (2011).
Gan, H. H. & Gunsalus, K. C. Assembly and analysis of eukaryotic Argonaute–RNA complexes in microRNA-target recognition. Nucleic Acids Res. 43, 9613–9625 (2015).
Elkayam, E. et al. The structure of human argonaute-2 in complex with miR-20a. Cell 150, 100–110 (2012).
Schirle, N. T. & MacRae, I. J. The crystal structure of human Argonaute2. Science 336, 1037–1040 (2012).
Miyoshi, T., Ito, K., Murakami, R. & Uchiumi, T. Structural basis for the recognition of guide RNA and target DNA heteroduplex by Argonaute. Nat. Commun. 7, 11846 (2016).
Hu, Y. et al. Study of rice pollen grains by multispectral imaging microscopy. Microsc. Res. Tech. 68, 335–346 (2005).
Zeng, Y. X., Hu, C. Y., Lu, Y. G., Li, J. Q. & Liu, X. D. Diversity of abnormal embryo sacs in indica/japonica hybrids in rice demonstrated by confocal microscopy of ovaries. Plant Breeding 126, 574–580 (2007).
Fujii, S. & Toriyama, K. Molecular mapping of the fertility restorer gene for ms-CW-type cytoplasmic male sterility of rice. Theor. Appl. Genet. 111, 696–701 (2005).
Martin, F. W. Staining and observing pollen tubes in the style by means of fluorescence. Stain Technol. 34, 125–128 (1959).
Patel, R. et al. Mutation scanning using MUT-MAP, a high-throughput, microfluidic chip-based, multi-analyte panel. PLoS One 7, e51153 (2012).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 6, 80–92 (2012).
Wang, J. et al. Tissue-specific ubiquitination by IPA1 INTERACTING PROTEIN1 modulates IPA1 protein levels to regulate plant architecture in rice. Plant Cell 29, 697–707 (2017).
Yang, X., Chen, L., He, J. & Yu, W. Knocking out of carotenoid catabolic genes in rice fails to boost carotenoid accumulation, but reveals a mutation in strigolactone biosynthesis. Plant Cell Rep. 36, 1533–1545 (2017).
Lyznik, L. A., Mitchell, J. C., Hirayama, L. & Hodges, T. K. Activity of yeast FLP recombinase in maize and rice protoplasts. Nucleic Acids Res. 21, 969–975 (1993).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
’t Hoen, P. A. C. et al. Deep sequencing-based expression analysis shows major advances in robustness, resolution and inter-lab portability over five microarray platforms. Nucleic Acids Res. 36, e141–e141 (2008).
Tang, C., Xie, Y. & Yan, W. AASRA: an anchor alignment-based small RNA annotation pipeline. Biol. Reprod. 105, 267–277 (2017).
Li, X. et al. Comparative small RNA analysis of pollen development in autotetraploid and diploid rice. Int. J. Mol. Sci. 17, 499 (2016).
Komoda, K., Naito, S. & Ishikawa, M. Replication of plant RNA virus genomes in a cell-free extract of evacuolated plant protoplasts. Proc. Natl Acad. Sci. USA 101, 1863–1867 (2004).
Komoda, K., Mawatari, N., Hagiwara-Komoda, Y., Naito, S. & Ishikawa, M. Identification of a ribonucleoprotein intermediate of tomato mosaic virus RNA replication complex formation. J. Virol. 81, 2584–2591 (2007).
Iki, T. et al. In vitro assembly of plant RNA-induced silencing complexes facilitated by molecular chaperone HSP90. Mol. Cell 39, 282–291 (2010).
Iki, T., Ishikawa, M. & Yoshikawa, M. In vitro formation of plant RNA-induced silencing complexes using an extract of evacuolated tobacco protoplasts. Methods Mol. Biol. 1640, 39–53 (2017).
Acknowledgements
We thank Biogle GeneTech for providing a gene-editing vector and transgenic plants. This work was funded by the National Natural Science Foundation of China (NSFC31971924), Hong Kong Research Grant Council (GRF14122415), and JSPS KAKENHI (18H02380).
Author information
Authors and Affiliations
Contributions
J.Z. led and managed the project. J.Z., X.C., G.W., and N.Y. conceived the study. H.L. and G.W. performed the research. C.Y. analyzed the sequencing data. M.Y. performed the BYL in vitro experiments. H.G. performed the field planting. C.L. provided the mutant lines. J.C. instructed the RNA gel blotting. H.L., G.W., X.C., and X.Y. wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, H., You, C., Yoshikawa, M. et al. A spontaneous thermo-sensitive female sterility mutation in rice enables fully mechanized hybrid breeding. Cell Res 32, 931–945 (2022). https://doi.org/10.1038/s41422-022-00711-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41422-022-00711-0
This article is cited by
-
Pedigree genome data of an early-matured Geng/japonica glutinous rice mega variety Longgeng 57
Scientific Data (2024)
-
Changes in m6A RNA methylation are associated with male sterility in wolfberry
BMC Plant Biology (2023)
-
Producing hybrid seeds like conventional rice
Cell Research (2022)