Abstract
Inhibitors of cancer cell migration and invasion should be useful to inhibit metastasis. Then, we have screened microbial culture filtrates for the inhibitors of cancer cell migration. As a result, we isolated an antibiotic ketomycin from a culture filtrate of Actinomycetes SF2912 as an inhibitor of cancer cell migration. It is a known antibiotic, but its biological activity on mammalian cells has not been reported. Ketomycin inhibited cellular migration and invasion in human breast carcinoma MDA-MB-231 and MCF-7 cells at the non-toxic concentrations. Ketomycin decreased the expressions of MMP-9 and MMP-11 in MDA-MB-231 cells. Knockdown of each gene by siRNA inhibited the cellular migration and invasion. Ketomycin was then found to inhibit the cellular NF-κB activity that may be involved in the upstream signaling. For the mechanism of NF-κB inhibition, ketomycin inhibited autophosphorylation of IKK-α/IKK-β. Ketomycin also inhibited the 3D-invasion of MDA-MB-231 cells at the non-toxic concentrations. Thus, ketomycin having a comparatively simple structure may become a seed of anti-metastasis agent.
Similar content being viewed by others
Introduction
Compared with conventional anticancer agents that inhibit cancer cell growth causing apoptosis, metastasis inhibitors can be less toxic because they may be free of cytotoxicity. Metastasis is a complex process that involves the spread of a tumor to distant parts of the body from its original site. Before a tumor cell successfully colonize to a new area, it must complete those steps of metastasis consisting of detachment from the primary tumor, migration, invasion, transport in the blood or lymphatic vessels, and attachment and growth at the secondary site. Especially, migration and invasion are often involved in the mechanism of metastasis. Therefore, we looked for cellular migration inhibitors of low molecular weight with less toxicity from microbial culture filtrates. Previously, we reported novel compounds, migracin A and B, isolated from the culture filtrate of Streptomyces sp [1]. Without showing any cytotoxicity, migracin A and B inhibited cellular migration in human breast carcinoma MDA-MB-231, fibrosarcoma HT1080, and lung carcinoma A549 cells. Later, migracin A was found to inhibit IGF-1 expression to inhibit migration and invasion of human ovarian clear cell carcinoma cells [2]. However, large-scale preparation of migracin A appears difficult, since yield that form the producing agent is low and it is difficult to synthesize because of the complicated structure.
Until recently, monolayer 2D cultured cells have been mainly used for the study of cell-based cancer growth and metastasis [2]. Since cancer cells grow and metastasize with the 3D organization in the body, 3D cell culture assay should be closer to the in vivo analysis. Currently, 3D cultured cells have become available for the study of cancer cell invasion [3]. For the 3D invasion assay, multicellular spheroids are formed in spheroid formation extracellular matrix. Then, they are embedded in the invasion matrix forming a hydrogel network on which invasive cells can travel, such as the basement membrane extract (BME) or collagen I [4]. Meanwhile we designed and synthesized a novel NF-κB inhibitor dehydroxymethylepoxyquinomicin (DHMEQ) [5, 6]. This inhibitor covalently binds to the specific cysteine residues of NF-κB component such as p65, p50, RelB, and cRel [7, 8]. Recently, we found that DHMEQ inhibited 2D and 3D invasion of MDA-MB-231 cells without toxicity [9]. We have first introduced the 3D invasion as a model of early phase metastasis consisting of cell detachment and invasion [9, 10].
Late phase of metastasis includes tumor cell attachment to the secondary site and growth. Among in vivo metastasis experiments, analysis of late stage metastasis is popular as lung [11] and liver [12] metastases by intravenous and portal injection of cells, respectively. Therefore, 3D invasion assay appears useful for the screening of early phase metastasis inhibitors.
In the present research, we have continued to screen new cancer cell migration inhibitors from microbial secondary metabolites. After the screening of several hundreds of samples, we found a hit sample, and isolated ketomycin from the culture filtrate of Actinomycetes as an inhibitor of breast cancer cell migration. It inhibited 2D and 3D invasion of MDA-MB-231 cells without toxicity. We also studied the mechanism of inhibition.
Materials and Methods
Screening and isolation
After screening of about 500 Actinomycetes culture filtrates, we found that the SF2912 broth inhibited migration of MDA-MB-231 cells without toxicity. Then, we purified the active compound using methanol extraction, silica gel chromatography, HW-40, and LH-20. The butanol extract of 11.4 g gave 40 mg of active principle as a light yellow crystal. The structure was determined to be ketomycin (Fig. 1a) using several spectrum methods including mass spectrum, 1H, 13C, and 2D-NMR and infrared spectroscopy.
Inhibition of cellular migration and invasion by ketomycin in human breast carcinoma cells. a Structure of ketomycin. b Left: effect on viability in MDA-MB-231 cells. The cells were cultured for 24 h with ketomycin. The cell viability was measured by MTT. Middle: inhibition of cellular migration in MDA-MB-231 cells. The cells were incubated for 24 h with ketomycin in wound healing assay. Right: inhibition of cellular invasion in MDA-MB-231 cells. The cells were incubated for 24 h in Matrigel invasion assay chambers. c Left: effect on viability in MCF-7 cells. The cells were cultured for 24 h with ketomycin. The cell viability was measured by MTT. Middle: inhibition of cellular migration in MCF-7 cells. The cells were incubated for 24 h with ketomycin in wound healing assay. Right: inhibition of cellular invasion in MCF-7 cells. The cells were incubated for 24 h in Matrigel invasion assay chambers. *P < 0.05, **P < 0.01, ***P < 0.001, n = 3
Actinomycetes strain SF2912 was isolated from a soil collected in Hokkaido Prefecture, Japan. A slant culture of SF2912 was inoculated in 100-ml Erlenmeyer flasks containing 15 ml of a seed medium (starch 2.5%, glucose 2.0%, polypepton 0.7%, wheat germ 0.6%, yeast extract 0.45%, soy bean meal 0.3%, Lab-Lemco-Powder 0.3%, CaCO3 0.2%, pH 7.2). The flasks were incubated at 28 °C for 2 days while shaking at 220 rpm. A portion of the seed culture was transferred into each of 500-ml Erlenmeyer flasks containing 80 ml of production medium (glucose 2.0%, starch 1.0%, soy bean meal 1.5%, wheat germ 0.8%, polypepton 0.1%, NaCl 0.1%, CaCO3 0.2%, ZnSO4 0.001%, Mg3(PO)4・8H2O 1.0%, pH 7.2) and incubated at 28 °C for 5 days on a rotary shaker.
Materials and cell culture
DHMEQ was synthesized in our laboratory as described before [13]. Human breast adenocarcinoma MDA-MB-231 and MCF-7 cells were purchased from RIKEN Cell Bank (Tsukuba, Japan). The cells were grown in Dulbecco’s modified Eagle’s medium (DMEM) (Sigma, St. Louis, MO), supplemented with 10% FBS (HyClone, Logan, UT) and 100 U ml−1 penicillin and 100 μg ml−1 streptomycin (Gibco, Grand Island, NY) under a humidified atmosphere with 5% CO2 at 37 °C.
MTT assay
Cell suspensions (100 μl) at a density of 1 × 105 cells per ml were plated in 96-well microtiter plates and incubated for 24 h. Then, the ketomycin solutions at different concentrations were added to each well and further incubated for 24 h. Ten microliter of MTT solution was added to each well and incubated for 2 h. Then, the culture supernatant was replaced by 100 μl DMSO and pipetted to dissolve formazan crystals which formed. Absorbance at 570 nm was then determined on a Microplate Reader (Bio-Rad Laboratories, Inc., Hercules, CA).
Wound healing assay
The cells (1 × 105) were seeded in 24-well until confluency. The confluent cells in 24-well was uniformly scratched across the center of each well with a 200 μl pipette tip. The wells were then rinsed twice with serum-free media to remove floating cells and growth media, and then the cells were cultured in serum-free media for 24 h. The wound areas at the initial and end time endpoints were recorded as the movement of the cells into the scratched area. Each determination was performed in triplicate in three independent experiments.
Matrigel invasion assay
MDA-MB-231 cells (4 × 104) were suspended in 500 μl of serum-free medium containing ketomycin or DMSO and seeded into the upper chambers coated with BD Matrigel Basement Membrane Matrix (Corning Inc., Corning, NY). The lower chambers were filled with 750 μl of medium containing 10% FBS and incubated for 24 h at 37 °C in incubator. After incubation, non-invading cells were removed by wiping with a cotton swab from the upper surface of the membrane. Invading cells on the lower surface of the membrane were stained with Diff-Quick (Sysmex, Kobe, Japan), and counted.
PCR array
Total RNA was extracted from MDA-MB-231 cells using RNeasy Mini Kit (Qiagen, Hilden, Germany). Reverse transcription was carried out with RT2 First Strand Kit (Qiagen, Maryland, USA). The cDNA was added to the qPCR Master Mix and the aliquot mixture across the Human Tumor Metastasis PCR Array (Qiagen, Maryland, USA). Data analysis was carried out by the comparative Ct method.
RNA isolation and semi-quantitative RT-PCR analysis
Total RNA was extracted from cultured cells using TRIzol reagent (Life Technologies, Carlsbad, CA). Reverse transcription was carried out at 37 °C for 120 min with High-Capacity cDNA Reverse Transcription Kit (Life Technologies, Carlsbad, CA). The prepared cDNA was used for PCR amplification with Quick Taq® HS DyeMix (Toyobo, Tokyo, Japan). The primer, number of PCR cycles, and the annealing temperature, as follows: MMP-9, 5′-ATTTCTGCCAGGACCGCTTCTACT-3′ (forward) and 5′-CAGTTTGTATCCGGCAAACTGGCT-3′ (reverse), 35 cycles, 58 °C; MMP-11, 5′-GAGCAGGTGCGGCAGACGA-3′ (forward) and 5′-CGAAAGGTGTAGAAGGCGGACA-3′ (reverse), 29 cycles, 58 °C; β-actin, 5′-CTTCTACAATGAGCTGCGTG-3′ (forward) and 5′-TCATGAGGTAGTCAGTCAGG-3′ (reverse), 21 cycles, 58 °C. PCR products were electrophoresed on 2% agarose gels, stained with ethidium bromide, and visualized with a UV illuminator.
Knockdown by siRNA
siMMP-9 (sc-29400), siMMP-11 (sc-35947), and control siRNA-A (sc-37007) were purchased from Santa Cruz Biotechnology Inc. (Dallas, TX). Transfection of cells with siRNAs was carried out using the Lipofectamine RNAiMax transfection reagent (Life Technologies, Carlsbad, CA) according to the manufacturer’s instruction. The efficiency of transfection was determined by mRNA expression.
Cellular and in vitro NF-κB activity
After the incubation with ketomycin, the nuclear extract was prepared with Nuclear Extract Kit (Active Motif, Carlsbad, CA) according to the instructions of the manufacturer. NF-κB binding activity with 5 μg of nuclear extract was measured with the TransAM NF-κB p65 Transcription Factor Assay Kit (Active Motif, Carlsbad, CA). For the in vitro assay, recombinant p65 (Active Motif, Carlsbad, CA) was treated with or without ketomycin for 15 min on ice. The DNA binding activity of p65 was measured as above.
IKK-α/IKK-β phosphorylation assay
Detection of phosphorylated IKK-α/IKK-β was carried out according to the protocol of the IKK-α (Phospho-Ser176)/IKK-β (phosphor-Ser177) Colorimetric Cell-based ELISA Kit (AssaybioTech, Fremont, CA). Briefly, 2 × 104 MDA-MB-231 cells were seeded in 200 μl of medium in 96-well plate. Incubate the cells overnight in incubator. The cells were treated with ketomycin for 4 h. The cells were fixed by 4% formaldehyde (Sigma, St. Louis, MO) for 20 min at room temperature. Then, the quenching buffer (100 μl) was added for 20 min. Blocking buffer 200 μl was added for 1 h. Then, the cells were treated with 50 μl of 1x primary antibodies (1/100) and incubated for 2 h at room temperature. After washing, the cells were added by 50 μl of 1x secondary antibodies (1/100). Then, the cells were added by 50 μl substrate for 30 min. After adding 50 μl of stop solution, OD at 450 nm was measured. Following the colorimetric measurement of HRP activity via substrate addition, the crystal violet whole-cell staining was employed to determine the cell density according to the protocol. The final results were normalized to the internal GAPDH and crystal violet cell staining.
3D spheroid proliferation assay
The 3D spheroid proliferation assay kit was purchased from Trevigen Inc. (Gaithersburg, MD) and used according to the manufacturer’s protocol as before [9]. Briefly, 3000 cells in 1x Spheroid Formation ECM were seeded for spheroid formation. Third day, 50 μl of warm (37 °C) cell culture medium containing ketomycin was added. After 4 days incubation, 10 μl of MTT was added for 24 h. Then 100 μl of detergents was added to solubilize cells and MTT formazan crystals. 24 h later, absorbance was read at 570 nm and data graphed.
3D spheroid invasion assay
3D spheroid invasion assay kit was purchased from Trevigen Inc. (Gaithersburg, MD) and used according to the manufacturer’s protocol as before [9]. In all, 3000 cells in 1x Spheroid Formation ECM were seeded for spheroid formation in 96 round-bottom well plate. After 3 days, put the well with spheroid on ice for 15 min. Then, they were added by the invasion matrix and medium containing ketomycin, and incubated for 4 days. The spheroids were photographed every 24 h at 4x objective. Spheroid areas were measured using Image J (National Institute of Health, Bethesda, MD).
IL-6 secretion in on-top 3D culture
We purchased 3D culture Matrix Reduced Growth Factor Basement Membrane Extract (BME) from Trevigen Inc. On-top 3D culture was performed according to the manufacturer’s protocol. After thawing BME at 4 ˚C overnight, 80 μl of BME was added into the pre-chilled 48-well plate, working on ice, and the well plate was incubated for 30 min at 37 ˚C. Then, 1.6 × 104 cells were plated onto the BME-coated surface and the well plate was incubated for 30 min at 37 ˚C. After 30 min, 200 μl of pre-chilled medium containing 10% BME was added. Cells were cultured for 4 days and the medium was changed every 2 days. After 4 days, the medium was replaced to the new one including ketomycin and the cells were incubated for 24 h. The supernatants were used for the human IL-6 ELISA assay (R&D Systems, Inc. Minneapolis, MN).
Statistical analysis
All experiments were performed independently at least three times. Data are expressed as mean ± SEM. Differences between groups were examined for significance with ANOVA and/or the Student’s t-test where appropriate.
Results
Inhibition of migration and invasion by ketomycin in breast carcinoma cells
Ketomycin (Fig. 1a) did not decrease the cellular viability below 3 μg ml−1 in MDA-MB-231 cells after incubation for 24 h (Fig. 1b left). It inhibited the migration of cells at 0.1–3 μg ml−1 monitored by Wound healing assay (Fig. 1b middle). It also inhibited cellular invasion at 0.3–3 μg ml−1 (Fig. 1b right). We also studied the effect on migration and invasion in human breast carcinoma MCF-7 cells. Ketomycin did not decrease the viability again at 3 μg ml−1 (Fig. 1c left). Ketomycin inhibited cellular migration (Fig. 1c middle) and invasion (Fig. 1c right) in MCF-7 cells at the nontoxic concentrations.
Inhibition of MMP-9 and MMP-11 expressions by ketomycin in MDA-MB-231 cells
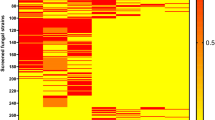
To study the mechanism of inhibition, we employed human tumor metastasis PCR array using MDA-MB-231 cells. As a result, several gene expressions were increased and a few gene expressions were decreased. We selected and further studied the effect on MMP-9 and MMP-11. We confirmed that these gene expressions were inhibited by ketomycin time- and dose-dependently (Fig. 2a, upper and lower, respectively). Next, we knocked down MMP-9 (Fig. 2b, upper) and MMP-11 (Fig. 2b, lower) by siRNA. Knockdown of MMP-9 inhibited the cellular migration, and ketomyicn did not further lower the migration (Fig. 2c). In the same way knockdown of MMP-9 inhibited the cellular invasion, and ketomyicn did not further inhibit the invasion (Fig. 2d). Knockdown of MMP-11 also inhibited the cellular migration, and ketomyicn did not further inhibit the migration (Fig. 2e). In the same way knockdown of MMP-11 inhibited the cellular invasion, and ketomyicn did not further inhibit the invasion (Fig. 2f).
Inhibition of cellular migration and invasion by knockdown of MMP-9 or MMP-11 in MDA-MB-231 cells. a Time and dose-dependent inhibition of MMP-9 and MMP-11. b Knockdown of MMP-9 and MMP-11 by siRNA. The decrease of expressions was confirmed by RT-PCR. c Inhibition of cellular migration by knockdown of MMP-9. d Inhibition of cellular invasion by knockdown of MMP-9. e Inhibition of cellular migration by knockdown of MMP-11. f Inhibition of cellular invasion by knockdown of MMP-11. *P < 0.05, **P < 0.01, n = 3
Inhibition of NF-κB by ketomycin in MDA-MB-231 cells
MMP-9 expression is well known to be dependent on NF-κB [14]. NF-κB-dependent MMP-11 expression has also been reported [15]. Then, we studied the effect on NF-κB. NF-κB is constitutively activated in MDA-MB-231 cells [16]. Larger number of cells are used for the NF-κB assay than for the migration and invasion assay. Ketomycin did not decrease the viability at 10 μg ml−1 in 4 h (Fig. 3a). Ketomycin was found to inhibit cellular NF-κB activity below 10 μg ml−1 (Fig. 3b). We have previously reported that DHMEQ directly inhibits NF-κB components by the covalent binding [7]. However, ketomycin did not prominently inhibit the NF-κB activity of p65 in vitro (Supplementary Fig. 1). On the other hand, ketomycin was found to inhibit the IKK auto-phosphorylation (Fig. 3c). Therefore, it is likely that ketomycin would inhibit I-κB kinase directly or indirectly. Ketomycin also inhibited the secretion of IL-6 which is dependent on NF-κB, as shown in Fig. 3d.
Inhibition of cellular NF-κB activity. a Effect of ketomycin on viability after the 4 h incubation. b Inhibition of cellular NF-κB activity by ketomycin. The cells were incubated for 4 h with ketomycin, and the nuclear extract was prepared for the assay of NF-κB activity. c Inhibition of IKK auto-phosphorylation by ketomycin. The cells were incubated for 4 h with ketomycin, and IKK activity was measured by Colorimetric Cell-based ELISA. d Inhibition of IL-6 secretion by ketomycin. The cells were incubated for 24 h and the secretion of IL-6 was measured by ELISA. *P < 0.05, **P < 0.01, n = 3
Inhibition of 3D invasion of MDA-MB-231 cells by ketomycin
To study the effect on 3D invasion of MDA-MB-231 cells, firstly, we measured the toxicity of ketomycin in 3D-cultured cells. Ketomycin was added after the spheroid formation, and it did not lower the viability even at 100 μg ml−1 (Fig. 4a). Next, ketomycin was shown to inhibit 3D-invasion at 10–30 μg ml−1, as shown in Fig. 4b. To show the decrease of NF-κB activity by ketomycin in 3D culture, we measured the secretion of IL-6. Ketomycin significantly lowered the secretion of IL-6 in the on-top 3D culture assay (Fig. 4c) as in the 2D culture assay (Fig. 3d).
Inhibition of 3D invasion by ketomycin in MDA-MB-231 cells. a Effect of ketomycin on the proliferation in 3D-cultured cells. b Inhibition of 3D invasion by ketomycin. Spheroids were incubated with ketomycin for 96 h. c Inhibition of IL-6 secretion in on-top 3D culture assay. Ketomycin was added for 24 h to the 3D-culture of MDA-MB-231 cells. *P < 0.05, **P < 0.01, ***P < 0.001, n = 6; Scale bar, 500 μm
Discussion
Ketomycin was first isolated from Streptomyces in 1969 as an antibiotic [17]. It inhibited the growth of Bacillus subtilis, and the antibiotic activity was found to be canceled by addition of amino acids such as l-serine and threonine, suggesting that it may inhibit amino acid synthesis. Its biosynthetic pathway [18] and the mechanism of resistance in Escherichia coli [19] were studied. However, little is known about the biological activity especially in mammalian cells.
We mainly employed MDA-MB-231 cells throughout this research, because this cell line can be used for 3D invasion research [9]. However, ketomycin also inhibited migration and invasion in human breast carcinoma MCF-7 cells (Fig. 1b).
Ketomycin is likely to inhibit MMP-9 and MMP-11 expressions by inhibition of constitutively activated NF-κB in MDA-MB-231 cells [16]. In fact we found that an NF-κB inhibitor DHMEQ inhibited the expressions of MMP-9 and MMP-11 (Supplementary Fig. 2). DHMEQ was also reported to inhibit migration and invasion in MDA-MB-231 cells [9]. Thus, inhibition of NF-κB should inhibit MMP-9 and MMP-11 expressions. To inhibit NF-κB, ketomycin was found to inhibit IKK-α/IKK-β phosphorylation as shown in Fig. 3c.
Ketomycin was used at 0.3–3 μg ml−1 for the experiments of migration and invasion (Fig. 1b), while 1–10 μg ml−1 for the experiments of cellular NF-κB (Fig. 2b). It is because the cell density is much higher in the NF-κB analysis to prepare the nuclear extracts. We have used ketomycin at the non-toxic concentrations throughout this research.
Cancer cells grow and metastasize in the body with the 3D organization interacting with neighboring cancer cells. Recently, invasion of cancer cells has become available for study in the 3D condition [3]. In the 3D invasion assay, we can observe detachment of tumor cells from the sphere, migration, and invasion into the basement membrane extract (BME) or collagen I. This phenomenon is just like the early phase of metastasis [9, 10]. We previously reported that DHMEQ inhibited 3D invasion of MDA-MB-231 cells [9]. In the present research, ketomycin also inhibited the 3D invasion at the non-toxic concentrations (Fig. 4b). Then, ketomycin is likely to inhibit the early phase of metastasis.
Ketomycin is simple in structure and easy to prepare by synthesis, and it may be a candidate of anti-metastasis agent.
References
Arai Y, et al. Migracins A and B, new inhibitors of cancer cell migration, produced by Streptomyces sp. J Antibiot (Tokyo). 2013;66:225–30.
Ukaji T, Lin Y, Banno K, Okada S, Umezawa K. Inhibition of IGF-1-mediated cellular migration and invasion by migracin A in ovarian clear cell carcinoma cells. PLoS ONE. 2015;10:e0137663.
Sodek KL, Ringuette MJ, Brown TJ. Compact spheroid formation by ovarian cancer cells is associated with contractile behavior and an invasive phenotype. Int J Cancer. 2009;124:2060–70.
Wenzel C, Otto S, Prechtl S, Parczyk K, Steigemann P. A novel 3D high-content assay identifies compounds that prevent fibroblast invasion into tissue surrogates. Exp Cell Res. 2015;339:35–43.
Matsumoto N, et al. Synthesis of NF-κB activation inhibitors derived from epoxyquinomicin C. Bioorg Med Chem Lett. 2000;10:865–69.
Ariga A, Namekawa J, Matsumoto N, Inoue J, Umezawa K. Inhibition of tumor necrosis factor-α-induced nuclear translocation and activation of NF-κB by dehydroxymethylepoxyquinomicin. J Biol Chem. 2002;277:24625–30.
Yamamoto M, Horie R, Takeiri M, Kozawa I, Umezawa K. Inactivation of NF-κB components by covalent binding of (-)-dehydroxymethylepoxyquinomicin to specific cysteine residues. J Med Chem. 2008;51:5780–88.
Takeiri M, et al. Involvement of DNA binding domain in the cellular stability and importin affinity of NF-κB component RelB. Org Biomol Chem. 2012;10:3053–59.
Ukaji T, Lin Y, Okada S, Umezawa K. Inhibition of MMP-2-mediated cellular invasion by NF-κB inhibitor DHMEQ in 3D culture of breast carcinoma MDA-MB-231 cells: a model for early phase of metastasis. Biochem Biophys Res Commun. 2017;485:76–81.
Lin Y, Ukaji T, Koide N, Umezawa K. Inhibition of late and early phases of cancer metastasis by the NF-κB inhibitor DHMEQ derived from microbial bioactive metabolite epoxyquinomicin: a Review. Int J Mol Sci. 2018;19:729 https://doi.org/10.3390/ijms19030729
Saito D, et al. Inhibition of experimental blood-borne lung metastasis by protease inhibitors. Cancer Res. 1980;40:2539–42.
Suzuki K, et al. Combined effect of dehydroxymethylepoxyquinomicin and gemcitabine in a mouse model of liver metastasis of pancreatic cancer. Clin Exp Metastasis. 2013;30:381–92.
Suzuki Y, Sugiyama C, Ohno O, Umezawa K. Preparation and biological activities of optically active dehydroxymethylepoxyquinomicin, a novel NF-κB inhibitor. Tetrahedron. 2004;60:7061–66.
Kang H, et al. N-(4-hydroxyphenyl)retinamide inhibits breast cancer cell invasion through suppressing NF-KB activation and inhibiting matrix metalloproteinase-9 expression. J Cell Biochem. 2012;113:2845–55.
Pires BR, et al. NF-kappaB is involved in the regulation of EMT genes in breast cancer cells. PLoS ONE. 2017;12:e0169622.
Matsumoto G, et al. Targeting of NF-κB pathways by DHMEQ, a novel inhibitor of breast carcinomas: Anti-tumor and anti-angiogenic activity in vivo. Clin Cancer Res. 2005;11:1287–93.
Keller-Schierlein W, Poralla K, Zahner H. Metabolic products of microorganisms 78. isolation, identification and mechanism of action of ketomycin [(R)-3-Cyclohexeneglyoxylic Acid] and of the conversion product 3-cyclohexeneglycine. Arch Mikrobiol. 1969;67:339–56.
Takeda Y, Mak V, Chang CC, Chang CJ, Floss HG. Biosynthesis of ketomycin. J Antibiot. 1984;37:868–75.
Jackson JH, Umbarger HE. Defective transamination, a mechanism for resistance to ketomycin in Escherichia coli. Antimicrob Agents Chemother. 1973;3:510–16.
Acknowledgements
We thank Meiji Seika Pharma Co. Ltd. for the preparation of Actinomycetes culture filtrates. We also thank Drs. Shinichi Kondo and Yoko Ikeda, Bioscience Associates, Tokyo, for the valuable suggestions on isolation and structure determination. This work was financially supported in part by JSPS Kakenhi Grant Number 17K01967, AMED under Grant Number JP18fk0310118, and the Aichi Medical University Research Unit Fund.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
K.U. is supported by Meiji Seika Pharma Co., Ltd, Tokyo, Japan, Shenzhen Wanhe Pharmaceutical Co., Ltd, Shenzhen, China, Fukuyu Hospital, Nisshin, Japan, and Brunese Co., Ltd, Nagoya, Japan. The remaining authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Lin, Y., Chen, Y., Ukaji, T. et al. Isolation of ketomycin from Actinomycetes as an inhibitor of 2D and 3D cancer cell invasion. J Antibiot 72, 148–154 (2019). https://doi.org/10.1038/s41429-018-0129-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-018-0129-9
This article is cited by
-
Induction of apoptosis and cell cycle arrest in colorectal cancer cells by novel anticancer metabolites of Streptomyces sp. 801
Cancer Cell International (2022)