Abstract
During X-chromosome inactivation (XCI), one of the two X-inactivation centers (Xics) upregulates the noncoding RNA Xist to initiate chromosomal silencing in cis. How one Xic is chosen to upregulate Xist remains unclear. Models proposed include localization of one Xic at the nuclear envelope or transient homologous Xic pairing followed by asymmetric transcription factor distribution at Xist’s antisense Xite/Tsix locus. Here, we use a TetO/TetR system that can inducibly relocate one or both Xics to the nuclear lamina in differentiating mouse embryonic stem cells. We find that neither nuclear lamina localization nor reduction of Xic homologous pairing influences monoallelic Xist upregulation or choice-making. We also show that transient pairing is associated with biallelic expression, not only at Xist/Tsix but also at other X-linked loci that can escape XCI. Finally, we show that Xic pairing occurs in wavelike patterns, coinciding with genome dynamics and the onset of global regulatory programs during early differentiation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in the published article (and its supplementary information). The raw datasets generated during and or analyzed during the current study are available from the corresponding author on reasonable request.
References
Rastan, S. & Robertson, E. J. X-chromosome deletions in embryo-derived (EK) cell lines associated with lack of X-chromosome inactivation. J. Embryol. Exp. Morphol. 90, 379–388 (1985).
Augui, S., Nora, E. P. & Heard, E. Regulation of X-chromosome inactivation by the X-inactivation centre. Nat. Rev. Genet. 12, 429–442 (2011).
Galupa, R. & Heard, E. X-chromosome inactivation: new insights into cis and trans regulation. Curr. Opin. Genet. Dev. 31, 57–66 (2015).
Pollex, T. & Heard, E. Recent advances in X-chromosome inactivation research. Curr. Opin. Cell Biol. 24, 825–832 (2012).
Borsani, G. et al. Characterization of a murine gene expressed from the inactive X chromosome. Nature 351, 325–329 (1991).
Brockdorff, N. et al. Conservation of position and exclusive expression of mouse Xist from the inactive X chromosome. Nature 351, 329–331 (1991).
Brockdorff, N. et al. The product of the mouse Xist gene is a 15 kb inactive X-specific transcript containing no conserved ORF and located in the nucleus. Cell 71, 515–526 (1992).
Brown, C. J. et al. A gene from the region of the human X inactivation centre is expressed exclusively from the inactive X chromosome. Nature 349, 38–44 (1991).
Brown, C. J. et al. The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell 71, 527–542 (1992).
Okamoto, I. et al. Eutherian mammals use diverse strategies to initiate X-chromosome inactivation during development. Nature 472, 370–374 (2011).
Marahrens, Y., Panning, B., Dausman, J., Strauss, W. & Jaenisch, R. Xist-deficient mice are defective in dosage compensation but not spermatogenesis. Genes Dev. 11, 156–166 (1997).
Borensztein, M. et al. Xist-dependent imprinted X inactivation and the early developmental consequences of its failure. Nat. Struct. Mol. Biol. 24, 226–233 (2017).
Takagi, N. & Abe, K. Detrimental effects of two active X chromosomes on early mouse development. Development 109, 189–201 (1990).
Lee, J. T. Regulation of X-chromosome counting by Tsix and Xite sequences. Science 309, 768–771 (2005).
Luikenhuis, S., Wutz, A. & Jaenisch, R. Antisense transcription through the Xist locus mediates Tsix function in embryonic stem cells. Mol. Cell. Biol. 21, 8512–8520 (2001).
Stavropoulos, N., Lu, N. & Lee, J. T. A functional role for Tsix transcription in blocking Xist RNA accumulation but not in X-chromosome choice. Proc. Natl Acad. Sci. USA 98, 10232–10237 (2001).
Navarro, P., Pichard, S., Ciaudo, C., Avner, P. & Rougeulle, C. Tsix transcription across the Xist gene alters chromatin conformation without affecting Xist transcription: implications for X-chromosome inactivation. Genes Dev. 19, 1474–1484 (2005).
Sado, T., Hoki, Y. & Sasaki, H. Tsix silences Xist through modification of chromatin structure. Dev. Cell. 9, 159–165 (2005).
Navarro, P., Page, D. R., Avner, P. & Rougeulle, C. Tsix-mediated epigenetic switch of a CTCF-flanked region of the Xist promoter determines the Xist transcription program. Genes Dev. 20, 2787–2792 (2006).
Ohhata, T., Hoki, Y., Sasaki, H. & Sado, T. Crucial role of antisense transcription across the Xist promoter in Tsix-mediated Xist chromatin modification. Development 135, 227–235 (2008).
Debrand, E., Chureau, C., Arnaud, D., Avner, P. & Heard, E. Functional analysis of the DXPas34 locus, a 3’ regulator of Xist expression. Mol. Cell. Biol. 19, 8513–8525 (1999).
Lee, J. T., Davidow, L. S. & Warshawsky, D. Tsix, a gene antisense to Xist at the X-inactivation centre. Nat. Genet. 21, 400–404 (1999).
Lee, J. T. & Lu, N. Targeted mutagenesis of Tsix leads to nonrandom X inactivation. Cell 99, 47–57 (1999).
Navarro, P. et al. Molecular coupling of Xist regulation and pluripotency. Science 321, 1693–1695 (2008).
Navarro, P. et al. Molecular coupling of Tsix regulation and pluripotency. Nature 468, 457–460 (2010).
Jonkers, I. et al. RNF12 is an X-Encoded dose-dependent activator of X chromosome inactivation. Cell 139, 999–1011 (2009).
Barakat, T. S. et al. RNF12 activates Xist and is essential for X chromosome inactivation. PLoS Genet. 7, e1002001 (2011).
Gontan, C. et al. RNF12 initiates X-chromosome inactivation by targeting REX1 for degradation. Nature 485, 386–390 (2012).
Barakat, T. S. et al. The trans-activator RNF12 and cis-acting elements effectuate X chromosome inactivation independent of X-pairing. Mol. Cell 53, 965–978 (2014).
Schulz, E. G. & Heard, E. Role and control of X chromosome dosage in mammalian development. Curr. Opin. Genet. Dev. 23, 109–115 (2013).
Xu, N., Tsai, C.-L. & Lee, J. T. Transient homologous chromosome pairing marks the onset of X inactivation. Science 311, 1149–1152 (2006).
Bacher, C. P. et al. Transient colocalization of X-inactivation centres accompanies the initiation of X inactivation. Nat. Cell Biol. 8, 293–299 (2006).
Xu, N., Donohoe, M. E., Silva, S. S. & Lee, J. T. Evidence that homologous X-chromosome pairing requires transcription and Ctcf protein. Nat. Genet. 39, 1390–1396 (2007).
Masui, O. et al. Live-cell chromosome dynamics and outcome of X chromosome pairing events during ES cell differentiation. Cell 145, 447–458 (2011).
Augui, S. et al. Sensing X chromosome pairs before X inactivation via a novel X-pairing region of the Xic. Science 318, 1632–1636 (2007).
Sun, S., Fukue, Y., Nolen, L., Sadreyev, R. & Lee, J. T. Characterization of Xpr (Xpct) reveals instability but no effects on X-chromosome pairing or Xist expression. Transcription 1, 46–56 (2010).
Donohoe, M. E., Silva, S. S., Pinter, S. F., Xu, N. & Lee, J. T. The pluripotency factor Oct4 interacts with Ctcf and also controls X-chromosome pairing and counting. Nature 460, 128–132 (2009).
Comings, D. E. The rationale for an ordered arrangement of chromatin in the interphase nucleus. Am. J. Hum. Genet. 20, 440–460 (1968).
Heard, E., Chaumeil, J., Masui, O. & Okamoto, I. Mammalian X-chromosome inactivation: an epigenetics paradigm. Cold Spring Harb. Symp. Quant. Biol. 69, 89–102 (2004).
Chen, C.-K. et al. Xist recruits the X chromosome to the nuclear lamina to enable chromosome-wide silencing. Science 354, 468–472 (2016).
Kosak, S. T. et al. Subnuclear compartmentalization of immunoglobulin loci during lymphocyte development. Science 296, 158–162 (2002).
Skok, J. A. et al. Reversible contraction by looping of the Tcra and Tcrb loci in rearranging thymocytes. Nat. Immunol. 8, 378–387 (2007).
Brown, K. E. et al. Association of transcriptionally silent genes with Ikaros complexes at centromeric heterochromatin. Cell 91, 845–854 (1997).
Skok, J. A. et al. Nonequivalent nuclear location of immunoglobulin alleles in B lymphocytes. Nat. Immunol. 2, 848–854 (2001).
Hewitt, S. L. et al. RAG-1 and ATM coordinate monoallelic recombination and nuclear positioning of immunoglobulin loci. Nat. Immunol. 10, 655–664 (2009).
Duncan, I. W. Transvection effects in Drosophila. Annu. Rev. Genet. 36, 521–556 (2002).
Fukaya, T. & Levine, M. Transvection. Curr. Biol. 27, R1047–R1049 (2017).
Lim, B., Heist, T., Levine, M. & Fukaya, T. Visualization of transvection in living Drosophila embryos. Mol. Cell 70, 287–296.e6 (2018).
Kumaran, R. I. & Spector, D. L. A genetic locus targeted to the nuclear periphery in living cells maintains its transcriptional competence. J. Cell. Biol. 180, 51–65 (2008).
Finlan, L. E. et al. Recruitment to the nuclear periphery can alter expression of genes in human cells. PLoS Genet. 4, e1000039 (2008).
Reddy, K. L., Zullo, J. M., Bertolino, E. & Singh, H. Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature 452, 243–247 (2008).
Dialynas, G., Speese, S., Budnik, V., Geyer, P. K. & Wallrath, L. L. The role of Drosophila Lamin C in muscle function and gene expression. Development 137, 3067–3077 (2010).
Pollex, T., Piolot, T. & Heard, E. Live-cell imaging combined with immunofluorescence, RNA, or DNA FISH to study the nuclear dynamics and expression of the X-inactivation center. Methods Mol. Biol. 1042, 13–31 (2013).
Krueger, C. et al. Pairing of homologous regions in the mouse genome is associated with transcription but not imprinting status. PLoS ONE 7, e38983 (2012).
Chow, J. C. et al. LINE-1 activity in facultative heterochromatin formation during X chromosome inactivation. Cell 141, 956–969 (2010).
Paulose, J. K., Rucker, E. B. & Cassone, V. M. Toward the beginning of time: circadian rhythms in metabolism precede rhythms in clock gene expression in mouse embryonic stem cells. PLoS ONE 7, e49555 (2012).
Yagita, K. et al. Development of the circadian oscillator during differentiation of mouse embryonic stem cells in vitro. Proc. Natl Acad. Sci. USA 107, 3846–3851 (2010).
Clowney, E. J. et al. Nuclear aggregation of olfactory receptor genes governs their monogenic expression. Cell 151, 724–737 (2012).
Armelin-Correa, L. M., Gutiyama, L. M., Brandt, D. Y. C. & Malnic, B. Nuclear compartmentalization of odorant receptor genes. Proc. Natl Acad. Sci. USA 111, 2782–2787 (2014).
Luo, L. et al. The nuclear periphery of embryonic stem cells is a transcriptionally permissive and repressive compartment. J. Cell. Sci. 122, 3729–3737 (2009).
Boyle, S. et al. The spatial organization of human chromosomes within the nuclei of normal and emerin-mutant cells. Hum. Mol. Genet. 10, 211–219 (2001).
Joyce, E. F., Erceg, J. & Wu, C.-T. Pairing and anti-pairing: a balancing act in the diploid genome. Curr. Opin. Genet. Dev. 37, 119–128 (2016).
Osborne, C. S. et al. Active genes dynamically colocalize to shared sites of ongoing transcription. Nat. Genet. 36, 1065–1071 (2004).
Brown, J. M. et al. Coregulated human globin genes are frequently in spatial proximity when active. J. Cell. Biol. 172, 177–187 (2006).
Brown, J. M. et al. Association between active genes occurs at nuclear speckles and is modulated by chromatin environment. J. Cell. Biol. 182, 1083–1097 (2008).
Chu, H.-P. et al. PAR-TERRA directs homologous sex chromosome pairing. Nat. Struct. Mol. Biol. 24, 620–631 (2017).
Kung, J. T. et al. Locus-specific targeting to the X chromosome revealed by the RNA interactome of CTCF. Mol. Cell 57, 361–375 (2015).
Aguilar-Arnal, L. et al. Cycles in spatial and temporal chromosomal organization driven by the circadian clock. Nat. Struct. Mol. Biol. 20, 1206–1213 (2013).
Norris, D. P. et al. Evidence that random and imprinted Xist expression is controlled by preemptive methylation. Cell 77, 41–51 (1994).
Schulz, E. G. et al. The two active X chromosomes in female ESCs block exit from the pluripotent state by modulating the ESC signaling network. Cell. Stem. Cell. 14, 203–216 (2014).
Giorgetti, L., Piolot, T. & Heard, E. High-resolution 3D DNA FISH using plasmid probes and computational correction of optical aberrations to study chromatin structure at the sub-megabase scale. Methods Mol. Biol. 1262, 37–53 (2015).
Chaumeil, J., Augui, S., Chow, J. C. & Heard, E. Combined immunofluorescence, RNA fluorescent in situ hybridization, and DNA fluorescent in situ hybridization to study chromatin changes, transcriptional activity, nuclear organization, and X-chromosome inactivation. Methods Mol. Biol. 463, 297–308 (2008).
Giorgetti, L. et al. Predictive polymer modeling reveals coupled fluctuations in chromosome conformation and transcription. Cell 157, 950–963 (2014).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Acknowledgements
We thank the laboratory of E.H. for helpful discussions and input. This work was supported by a Institut Curie PhD student fellowship and a fellowship from Fondation ARC (Aides Individuelles Jeunes Chercheurs) to T.P. E.H. is supported by an ERC Advanced Investigator award (ERC-2014-AdG no. 671027), Labelisation La Ligue, FRM (grant DEI20151234398), ANR DoseX 2017, Labex DEEP (ANR-11-LBX-0044), part of the IDEX Idex PSL (ANR-10-IDEX-0001–02 PSL) and ABS4NGS (ANR-11-BINF-0001). We thank N. Brockdorff (Oxford University), A. Belmont (University of Illinois) and F. Zhang (Broad Institute) for providing materials.
Author information
Authors and Affiliations
Contributions
T.P. and E.H. conceived the project and designed the experiments. T.P. carried out all experiments and analyzed the data. T.P. and E.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Gene expression in the Xic in PGK12.1 XXTetO TetR-EGFP and PGK12.1 XTetOXTetO TetR-EGFP-LaminB1 ESCs.
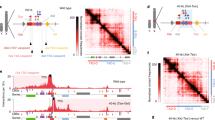
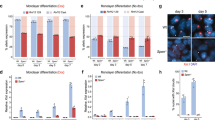
a, Depicted are colonies of PGK12.1 XXTetO or XTetOXTetO cell lines stably expressing TetR-EGFP fusion proteins. The imaged cells were fixed and the EGFP fluorescence signal was acquired. b, DNA FISH for the TetO array locus (green) and the Xist/Tsix region in the Xic (red). Displayed is a cell in which the TetO array locus localized to pericentric heterochromatin (DAPI-dense region, chromocenter) as depicted in the magnified panels next to the projection displaying the entire nucleus. c, Quantification of pericentric heterochromatin association in undifferentiated bound and control PGK12.1 XXTetO TetR-EGFP-LaminB1 cells. Scored was the overlap of the TetO array DNA FISH signal with DAPI-dense regions. d, Mean relative gene expression level (normalized to Arp0 expression levels) assessed by qRT–PCR in control (empty bars) and bound (empty bars, lighter shade) PGK12.1 XXTetO TetR-EGFP cells (*P < 0.05, **P < 0.01, t-test (unpaired, two-tailed); error bars indicate s.d.; n = 3; individual data points are depicted next to the respective bar as filled and empty circles). e, Mean relative gene expression level (normalized to Arp0 expression levels) assessed by qRT–PCR in control (empty bars) and bound (empty bars, lighter shade) PGK12.1 XTetOXTetO TetR-EGFP-LaminB1 cells (*P < 0.05, **P < 0.01, t-test (unpaired, two-tailed); error bars indicate s.d.; n = 3; individual data points are depicted next to the respective bar as filled and empty circles). f, RNA FISH was performed in control and bound PGK12.1 XXTetO TetR-EGFP-LaminB1 ESCs for nascent Tsix transcripts. Depicted is a single cell with biallelic Tsix expression. g, Quantification of the presence of focal RNA FISH signals indicating the presence of nascent Tsix transcripts. h, Sequential RNA and DNA FISH for Tsix (red) and Linx (white) transcripts (RNA FISH) as well as for the TetO array locus (green) and the Xist/Tsix region in the Xic (red) (DNA FISH). The second focal signal/pinpoint in the green channel is due to cross-hybridization of the TetO plasmid probe (backbone) with the stably inserted pBROAD3-TetR-EGFP-LaminB transgene. i, Logarithmic ratio of the integrated intensity of the RNA FISH signal of Tsix (left) and Linx (right) transcripts on Xic-TetO to the wildtype Xic in control and bound PGK12.1 XXTetO TetR-EGFP-LaminB1 cells based on sequential RNA–DNA FISH as depicted in h (*P < 0.05, KS test). (n is the number of cells; scale bar, 2 µm; representative cells for the respective condition are depicted in a, b, f and h.)
Supplementary Figure 2 Gene expression in bound and control PGK12.1 XXTetO TetR-EGFP-LaminB1cells during differentiation.
Mean relative gene expression assessed by qRT–PCR (normalized to Arp0 expression levels) in control (filled circles) and bound (empty circles) PGK12.1 XXTetO TetR-EGFP-LaminB1 ESCs and their differentiating counterparts (n = 6; error bars indicate s.d.; individual data points are depicted as smaller filled and empty circles in a lighter color shade).
Supplementary Figure 3 Xic–Xic distance distribution in PGK12.1 XXTetO and PGK12.1 XTetOXTetO TetR-EGFP-LaminB1 ESCs and their differentiating counterparts.
a, DNA FISH for different regions of the Xic (green, BAC 7; red, BAC 8; white, BAC 5) in bound PGK12.1 XXTetO TetR-EGFP-LaminB1 ESCs. (Scale bar, 2 µm; one representative cell is depicted.) b, Cumulative frequency plot of Xic–Xic distances (Xist/Tsix region, BAC 8) in control (dotted line, n = 834) and bound (continuous line, n = 799) PGK12.1 XXTetO TetR-EGFP-LaminB1 ESCs assessed by 3D distance measurement after DNA FISH as depicted in a (P = 0.007, Kolmogorov–Smirnov test; pooled results of three replicates). c, Distribution of Xic–Xic distances (Xist/Tsix region, BAC 8) in control (dotted line) and bound (continuous line) PGK12.1 XXTetO TetR-EGFP-LaminB1 ESCs assessed by 3D distance measurement after DNA FISH as depicted in a (500 nm binning). d, DNA FISH for various parts of the Xic (green, BAC 7; red, BAC 8; white, BAC 5) in bound PGK12.1 XTetOXTetO TetR-EGFP-LaminB1 ESCs. (Scale bar, 2 µm; one representative cell is depicted.) e, Cumulative frequency plot of Xic–Xic distances (Xist/Tsix region, BAC 8) in control (dotted line, n = 611) and bound (continuous line, n = 733) PGK12.1 XTetOXTetO TetR-EGFP-LaminB1 ESCs assessed by 3D distance measurement after DNA FISH as depicted in d (P = 1.57 × 10–4, Kolmogorov–Smirnov test; pooled results of three replicates). f, Distribution of Xic–Xic distances (Xist/Tsix region, BAC 8) in control (dotted line) and bound (continuous line) PGK12.1 XTetOXTetO TetR-EGFP-LaminB1 ESCs assessed by 3D distance measurement after DNA FISH as depicted in d (500 nm binning). g, Table depicting pairing frequencies observed in previous studies. h,i, Xic–Xic distance distribution in control (h) and bound (i) PGK12.1 XTetOXTetO TetR-EGFP-LaminB1 ESCs and their differentiating counterparts depicted as a box plot (the box shows the 0.25 quartile to the 0.75 quartile; median Xic–Xic distances are connected by a red line). j, Median Xic–Xic distance distribution in control PGK12.1 XTetOXTetO TetR-EGFP-LaminB1 ESCs and their differentiating counterparts from a single biological replicate. k, Relative gene expression assessed by qRT–PCR (normalized to Arp0 gene expression) in control (filled circles) and bound (empty circles) PGK12.1 XTetOXTetO TetR-EGFP-LaminB1 ESCs and their differentiating counterparts (n = 3; error bars indicate s.d.; individual data points are depicted as smaller filled and empty circles in a lighter color shade).
Supplementary Figure 4 Expression of Xist and Tsix in induced and uninduced TX1072 ESCs and their differentiating counterparts.
a, Expression of nascent Xist transcripts assessed by the presence of focal RNA FISH signals in control and differentiating TX1072 cells with either uninduced or induced Xist expression by addition of doxycycline to the culture medium 1 d prior to initiation of differentiation or simultaneous to the initiation of differentiation. b, Expression of nascent Tsix transcripts assessed by the presence of focal RNA FISH signals in control and differentiating TX1072 cells with either uninduced or induced Xist expression by addition of doxycycline to the culture medium 1 d prior to initiation of differentiation or simultaneous to the initiation of differentiation.
Supplementary Figure 5 Gene expression during differentiation of XXTetO ΔXist DKO ESCs.
a, Schematic representation of a wildtype Xist/Tsix locus and the introduced deletion. qPCR amplicons for Tsix 5ʹ, Tsix 3ʹ and Xist are indicated in the scheme. b, Depicted is the mean relative level of gene expression of selected pluripotency genes as well as Xist and Tsix assessed by qRT–PCR (normalized for Arp0 gene expression) during differentiation by LIF withdrawal in PGK12.1 XXTetO wildtype cells (filled circles) and PGK12.1 XXTetO ΔXist DKO cells (empty circles). (n = 3; error bars represent s.d.; individual data points are depicted as smaller filled and empty circles in a lighter color shade.)
Supplementary Figure 6 Gene expression during differentiation of PGK12.1 XXTetO cells.
Depicted is the mean relative level of gene expression of selected genes assessed by qRT–PCR (normalized for Arp0, Rrm2 and Actb gene expression) during differentiation by LIF withdrawal in PGK12.1 XXTetO cells. Blue, pluripotency factor genes; green, lamin genes; red, clock genes; orange, Glut8. Note that Lmna (laminA/C) is not highly expressed in ESCs and during early differentiation. Lmnb expression increases and decreases in a wave-like pattern during differentiation. Expression of Glut8 and the clock gene Per2 appears to initiate cycling after day 2 of differentiation. (n = 3; error bars represent s.d.; individual data points are depicted as smaller filled circles in a lighter color shade.)
Supplementary Figure 7 Vector maps of the expression plasmids used for the generation of stably expressing cell lines.
a,b, Vector maps of pBROAD3-TetR-EGFP (a) and pBROAD3-TetR-EGFP-LaminB1 (b) were generated using Geneious (Biomatters Limited) and SnapGene Viewer (GSL Biotech).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Note
Supplementary Table 1
Oligonucleotides
Supplementary Table 2
Probe templates
Supplementary Table 3
Antibodies
Rights and permissions
About this article
Cite this article
Pollex, T., Heard, E. Nuclear positioning and pairing of X-chromosome inactivation centers are not primary determinants during initiation of random X-inactivation. Nat Genet 51, 285–295 (2019). https://doi.org/10.1038/s41588-018-0305-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0305-7
This article is cited by
-
Gene regulation in time and space during X-chromosome inactivation
Nature Reviews Molecular Cell Biology (2022)
-
Activation of Xist by an evolutionarily conserved function of KDM5C demethylase
Nature Communications (2022)
-
Engineering 3D genome organization
Nature Reviews Genetics (2021)
-
Decapping enzyme 1A breaks X-chromosome symmetry by controlling Tsix elongation and RNA turnover
Nature Cell Biology (2020)
-
The role of 3D genome organization in development and cell differentiation
Nature Reviews Molecular Cell Biology (2019)