Abstract
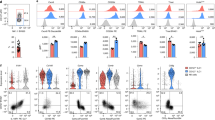
Group 2 innate lymphoid cells (ILC2s) are distributed systemically and produce type 2 cytokines in response to a variety of stimuli, including the epithelial cytokines interleukin (IL)-25, IL-33, and thymic stromal lymphopoietin (TSLP). Transcriptional profiling of ILC2s from different tissues, however, grouped ILC2s according to their tissue of origin, even in the setting of combined IL-25-, IL-33-receptor-, and TSLP-receptor-deficiency. Single-cell profiling confirmed a tissue-organizing transcriptome and identified ILC2 subsets expressing distinct activating receptors, including the major subset of skin ILC2s, which were activated preferentially by IL-18. Tissue ILC2 subsets were unaltered in number and expression in germ-free mice, suggesting that endogenous, tissue-derived signals drive the maturation of ILC2 subsets by controlling expression of distinct patterns of activating receptors, thus anticipating tissue-specific perturbations occurring later in life.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Moltke, von,J. & Locksley, R. M. I-L-C-2 it: type 2 immunity and group 2 innate lymphoid cells in homeostasis. Curr. Opin. Immunol. 31, 58–65 (2014).
Klose, C. S. N. & Artis, D. Innate lymphoid cells as regulators of immunity, inflammation and tissue homeostasis. Nat. Immunol. 17, 765–774 (2016).
Neill, D. R. et al. Nuocytes represent a new innate effector leukocyte that mediates type-2 immunity. Nature 464, 1367–1370 (2010).
Klein Wolterink, R. G. J. et al. Pulmonary innate lymphoid cells are major producers of IL-5 and IL-13 in murine models of allergic asthma. Eur. J. Immunol. 42, 1106–1116 (2012).
Salimi, M. et al. A role for IL-25 and IL-33-driven type-2 innate lymphoid cells in atopic dermatitis. J. Exp. Med. 210, 2939–2950 (2013).
Licona-Limón, P., Kim, L. K., Palm, N. W. & Flavell, R. A. TH2, allergy and group 2 innate lymphoid cells. Nat. Immunol. 14, 536–542 (2013).
Van Dyken, S. J. et al. A tissue checkpoint regulates type 2 immunity. Nat. Immunol. 17, 1381–1387 (2016).
Robinette, M. L. et al. Transcriptional programs define molecular characteristics of innate lymphoid cell classes and subsets. Nat. Immunol. 16, 306–317 (2015).
Simoni, Y. et al. Human Innate Lymphoid Cell Subsets Possess Tissue-Type Based Heterogeneity in Phenotype and Frequency. Immunity 46, 148–161 (2017).
Gury-BenAri, M. et al. The Spectrum and Regulatory Landscape of Intestinal Innate Lymphoid Cells Are Shaped by the Microbiome. Cell 166, 1231–1246.e13 (2016).
Nussbaum, J. C. et al. Type 2 innate lymphoid cells control eosinophil homeostasis. Nature 502, 245–248 (2013).
Huang, Y. et al. S1P-dependent interorgan trafficking of group 2 innate lymphoid cells supports host defense. Science 359, 114–119 (2018).
Bando, J. K., Nussbaum, J. C., Liang, H.-E. & Locksley, R. M. Type 2 innate lymphoid cells constitutively express arginase-I in the naive and inflamed lung. J. Leukoc. Biol. 94, 877–884 (2013).
Liang, H.-E. et al. Divergent expression patterns of IL-4 and IL-13 define unique functions in allergic immunity. Nat. Immunol. 13, 58–66 (2011).
Moltke, von,J., Ji, M., Liang, H.-E. & Locksley, R. M. Tuft-cell-derived IL-25 regulates an intestinal ILC2-epithelial response circuit. Nature 529, 221–225 (2016).
Bando, J. K., Liang, H.-E. & Locksley, R. M. Identification and distribution of developing innate lymphoid cells in the fetal mouse intestine. Nat. Immunol. 16, 153–160 (2015).
Shih, H.-Y. et al. Developmental Acquisition of Regulomes Underlies Innate Lymphoid Cell Functionality. Cell 165, 1120–1133 (2016).
Van Dyken, S. J. et al. Chitin activates parallel immune modules that direct distinct inflammatory responses via innate lymphoid type 2 and γδ T cells. Immunity 40, 414–424 (2014).
Molofsky, A. B. et al. Interleukin-33 and Interferon-γ Counter-Regulate Group 2 Innate Lymphoid Cell Activation during Immune Perturbation. Immunity 43, 161–174 (2015).
Moltke, von,J. et al. Leukotrienes provide an NFAT-dependent signal that synergizes with IL-33 to activate ILC2s. J. Exp. Med. 214, 27–37 (2017).
Cardoso, V. et al. Neuronal regulation of type 2 innate lymphoid cells via neuromedin U. Nature 549, 277–281 (2017).
Saluzzo, S. et al. First-Breath-Induced Type 2 Pathways Shape the Lung Immune Environment. Cell Rep. 18, 1893–1905 (2017).
Schneider, C. et al. A Metabolite-Triggered Tuft Cell-ILC2 Circuit Drives Small Intestinal Remodeling. Cell 174, 271–284.e14 (2018).
Guo, L., Junttila, I. S. & Paul, W. E. Cytokine-induced cytokine production by conventional and innate lymphoid cells. Trends Immunol. 33, 598–606 (2012).
Mohapatra, A. et al. Group 2 innate lymphoid cells utilize the IRF4-IL-9 module to coordinate epithelial cell maintenance of lung homeostasis. Mucosal Immunol. 9, 275–286 (2016).
Kim, B. S. et al. TSLP elicits IL-33-independent innate lymphoid cell responses to promote skin inflammation. Sci. Transl. Med. 5, 170ra16–170ra16 (2013).
Konishi, H. et al. IL-18 contributes to the spontaneous development of atopic dermatitis-like inflammatory skin lesion independently of IgE/stat6 under specific pathogen-free conditions. Proc. Natl Acad. Sci. USA 99, 11340–11345 (2002).
Thijs, J. et al. Biomarkers for atopic dermatitis: a systematic review and meta-analysis. Curr. Opin. Allergy Clin. Immunol. 15, 453–460 (2015).
Zedan, K. et al. Immunoglobulin e, interleukin-18 and interleukin-12 in patients with atopic dermatitis: correlation with disease activity. J. Clin. Diagn. Res. 9, WC01–WC05 (2015).
Ferreira, M. A. et al. Shared genetic origin of asthma, hay fever and eczema elucidates allergic disease biology. Nature Genet. 49, 1752–1757 (2017).
Demenais, F. et al. Multiancestry association study identifies new asthma risk loci that colocalize with immune-cell enhancer marks. Nature Genet. 50, 42–53 (2018).
Steer, C. A. et al. Group 2 innate lymphoid cell activation in the neonatal lung drives type 2 immunity and allergen sensitization. J. Allergy Clin. Immunol. 140, 593–595.e3 (2017).
Mass, E. et al. Specification of tissue-resident macrophages during organogenesis. Science 353, aaf4238–aaf4238 (2016).
Kotas, M. E. & Locksley, R. M. Why Innate Lymphoid Cells? Immunity 48, 1081–1090 (2018).
Li, M. et al. Induction of thymic stromal lymphopoietin expression in keratinocytes is necessary for generating an atopic dermatitis upon application of the active vitamin D3 analogue MC903 on mouse skin. J. Invest. Dermatol. 129, 498–502 (2009).
Molofsky, A. B. et al. Innate lymphoid type 2 cells sustain visceral adipose tissue eosinophils and alternatively activated macrophages. J. Exp. Med. 210, 535–549 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank M. Consengco and M. Ji for technical expertise; Z. Wang for cell sorting; A. Barczak, R. Barbeau, and J. Pollack for assistance with RNA-seq; E. Wan for assistance with scRNA-seq; J. Turnbaugh (University of California San Francisco) for providing germ-free mice; and M. Ansel and A. Marson for comments on the manuscript. This work was supported by the National Institutes of Health (AI030663 and HL128903 to R.M.L., AI122702 to J.L., DK101604 to A.B.M., and AI113143 to J.C.N.), Dermatology Foundation (R.R.R.-G.), A.P. Giannini Foundation (R.R.R.-G.), Robert Wood Johnson Foundation (R.R.R.-G.), Swiss National Science Foundation (P2EZP3_162266 and P300PA_171591 to C.S.), Howard Hughes Medical Institute (R.M.L.), and the Sandler Asthma Basic Research Center at the University of California San Francisco (R.M.L).
Author information
Authors and Affiliations
Contributions
R.R.R.-G. and S.J.V.D. designed and performed experiments, analyzed and interpreted the data, and wrote the manuscript. C.S., J.L., J.C.N., H.-E.L., and A.B.M. contributed to experiments. D.V. analyzed scRNA-seq data, and W.L.E. and D.J.E. provided RNA-seq data analysis and expertise. R.M.L. directed the studies and wrote the manuscript with R.R.R.-G. and S.J.V.D.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Gating strategy for sorting tissue-resident function-marked ILC2s.
a-e, Gating strategy for sorting lung (a), skin (b), gut (c, small intestine lamina propia), fat (d, perigonadal white adipose tissue) and bone marrow (e, BM) ILC2s as described in Methods.
Supplementary Figure 2 RNA-seq of peripheral program gene sets in ILC2s.
a-e, RNA-seq analysis of select transcripts significantly enriched (FDR < 0.01) in R5 + ILC2s from peripheral tissues (lung, gut, fat, skin) versus Yarg + BM ILC2s. a, enzyme / ion channel. b, extracellular / nucleus. c, plasma membrane, d, cytoplasm. e, transporter. Data represent mean normalized read counts (fragments per million mapped reads) from biological replicates (n = 3, BM; n = 5, lung; n = 6, fat, gut, skin).
Supplementary Figure 3 ILC2 counts and select transcripts in resting SPF versus GF mice.
a-b, Number of ILC2s (Lin–CD45+Thy1.2+CD25+) in fat (a), and bone marrow (b). c-h, Quantitative RT-PCR (qPCR) of Gata3 (c), Arg1 (d), Il1rl1 (e), Il17rb (f), Tnfaip3 (g), and Il18r1 (h) transcript abundance among germ-free (GF) or specific pathogen free (SPF) ILC2s sorted from BM, lung, gut, skin, and fat. Data represent biological replicates (n = 3; mean ± SD) and transcripts are normalized to Rps17. * p < 0.05.
Supplementary Figure 4 Steady-state ILC2 reporter expression.
Representative flow cytometry of CD45+Lin– cells from indicated single knockout (KO) or TKO mice on YRS backgrounds. Numbers within the gate represent percentage of activated (IL-5-producing) ILC2s among total ILC2s (Lin–CD45+Thy1.2+CD25+ in lungs, fat, and skin; Lin–CD45+IL17Rb+KLRG1+ in gut). Data are representative of 2 or more independent experiments representing at least 3 mice in each group.
Supplementary Figure 5 Gene expression in ILC2s from multiple tissues.
a, Heat map of log2 fold change relative global average for top 1000 genes by variance. b-g, Mean normalized read counts (fragments per million mapped reads) for Gata3 (b), Il7r (c), Il5 (d), Il13 (e), Il17rb (f), and Il18r1 (g) transcripts among WT and TKO ILC2s from BM, lung, fat, gut and skin from biological replicates (n = 3, WT BM, TKO BM, TKO gut; n = 4, TKO lung, TKO fat; n = 5, WT lung, TKO skin; n = 6, WT fat, WT gut, WT skin). * p < 0.05; ** p < 0.005; *** p < 0.0005.
Supplementary Figure 6 Single-cell transcriptome analysis of tissue-resident ILC2s.
a, tSNE plot representing 35,396 ILC2s sorted from BM (Lin–CD45+Yarg+), lung, fat, gut, and skin (Lin–CD45+Red5+) analyzed by single-cell RNA sequencing (scRNA-seq). b, tSNE plot of graph-based clustering of tissue-resident scRNA-seq reveals distinct intra-tissue ILC2 subsets. c, Heat map of hierarchical clustering of top 100 up-regulated (log2 fold change) genes from intra-tissue clusters shown in (b). Arrows highlights differences in Il1rl1 and Il18r1 between clusters.
Supplementary Figure 7 Single-cell RNA-seq reveals gradation in peripheral activation program.
Normalized relative expression of Tbx21, Rorc, Il2rg, Arg1, Il13, Thy1, Crlf2, Areg, Klrg1, Il2ra, Il2rb, Itgae, Icos, Cd69, and Cd44 from tissue ILC2s (BM (Lin–CD45+Yarg+), lung, fat, gut, and skin (Lin–CD45+Red5+) by scRNA-seq. Data are representative of 2 independent experiments.
Supplementary Figure 8 Number of R5+ ILC2s in IL-18 KO mice.
a, b, Number of R5 + ILC2s (Lin–CD45+Thy1.2+Red5+) in skin (a) and lung (b) of WT and IL-18KO mice. Data pooled from 2 individual experiments with n ≥ 10 per group and represented as mean±SD.
Supplementary information
Supplementary Figures
Supplementary Figures 1–8
Supplementary Table 1
RNA sequencing analysis of tissue ILC2s. Bulk RNA-seq data table related to Figures 1, 4, and Supplementary Figs. 2 and 5.
Supplementary Table 2
Primers used for qRT-PCR.
Supplementary Table 3
Pipeline code for scRNA-seq Cellranger aggr.
Rights and permissions
About this article
Cite this article
Ricardo-Gonzalez, R.R., Van Dyken, S.J., Schneider, C. et al. Tissue signals imprint ILC2 identity with anticipatory function. Nat Immunol 19, 1093–1099 (2018). https://doi.org/10.1038/s41590-018-0201-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0201-4
This article is cited by
-
A type 1 immunity-restricted promoter of the IL−33 receptor gene directs antiviral T-cell responses
Nature Immunology (2024)
-
Biological and clinical roles of IL-18 in inflammatory diseases
Nature Reviews Rheumatology (2024)
-
A diversity of novel type-2 innate lymphoid cell subpopulations revealed during tumour expansion
Communications Biology (2024)
-
Disease pathogenesis and barrier functions regulated by group 3 innate lymphoid cells
Seminars in Immunopathology (2024)
-
Group 2 innate lymphoid cells and their surrounding environment
Inflammation and Regeneration (2023)