Abstract
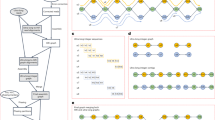
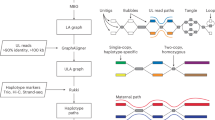
Draft genomes generated from Oxford Nanopore Technologies (ONT) long reads are known to have a higher error rate. Although existing genome polishers can enhance their quality, the error rate (including mismatches, indels and switching errors between paternal and maternal haplotypes) can be significant. Here, we develop two polishers, hypo-short and hypo-hybrid to address this issue. Hypo-short utilizes Illumina short reads to polish an ONT-based draft assembly, resulting in a high-quality assembly with low error rates and switching errors. Expanding on this, hypo-hybrid incorporates ONT long reads to further refine the assembly into a diploid representation. Leveraging on hypo-hybrid, we have created a diploid genome assembly pipeline called hypo-assembler. Hypo-assembler automates the generation of highly accurate, contiguous and nearly complete diploid assemblies using ONT long reads, Illumina short reads and optionally Hi-C reads. Notably, our solution even allows for the production of telomere-to-telomere diploid genomes with additional manual steps. As a proof of concept, we successfully assembled a fully phased telomere-to-telomere diploid genome of HG00733, achieving a quality value exceeding 50.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All the assemblies and annotations are available in https://zenodo.org/doi/10.5281/zenodo.10494612. CHM13 are mostly taken from the CHM13 GitHub page https://github.com/marbl/CHM13: ONT reads, https://s3-us-west-2.amazonaws.com/human-pangenomics/T2T/CHM13/nanopore/rel3/rel3.fastq.gz; Illumina reads, https://www.ncbi.nlm.nih.gov/sra/SRX1009644 [accn]; Reference: https://s3-us-west-2.amazonaws.com/human-pangenomics/T2T/CHM13/assemblies/chm13.draft_v1.1.fasta.gz; Annotation, https://s3-us-west-2.amazonaws.com/human-pangenomics/T2T/CHM13/assemblies/annotation/chm13.draft_v2.0.gene_annotation.gff3. HG002 are mostly taken from HG002 data freeze https://github.com/human-pangenomics/HG002_Data_Freeze_v1.0 with the only difference on ONT reads, where we take a newer set of Guppy pangenomics: ONT reads, https://s3-us-west-2.amazonaws.com/human-pangenomics/NHGRI_UCSC_panel/HG002/nanopore/Guppy_4.2.2/HG002_GIAB_MinION_GridION_Guppy_4.2.2.fastq.gz and https://s3-us-west-2.amazonaws.com/human-pangenomics/NHGRI_UCSC_panel/HG002/nanopore/Guppy_4.2.2/HG002_GIAB_PromethION_Guppy_4.2.2_prom.fastq.gz; Illumina reads, https://ftp-trace.ncbi.nlm.nih.gov/ReferenceSamples/giab/data_indexes/AshkenazimTrio/sequence.index.AJtrio_Illumina300X_wgs_07292015_updated.HG002; HiFi reads, https://s3-us-west-2.amazonaws.com/human-pangenomics/NHGRI_UCSC_panel/HG002/hpp_HG002_NA24385_son_v1/PacBio_HiFi/15kb/m64012_190920_173625.Q20.fastq and https://s3-us-west-2.amazonaws.com/human-pangenomics/NHGRI_UCSC_panel/HG002/hpp_HG002_NA24385_son_v1/PacBio_HiFi/15kb/m64012_190921_234837.Q20.fastq; Hi-C reads, https://s3-us-west-2.amazonaws.com/human-pangenomics/index.html?prefix=NHGRI_UCSC_panel/HG002/hpp_HG002_NA24385_son_v1/hic/downsampled/; Reference, https://www.ncbi.nlm.nih.gov/assembly/GCA_021951015.1/ and https://www.ncbi.nlm.nih.gov/assembly/GCA_021950905.1. HG00733 are taken from human pangenomics: ONT reads, https://s3-us-west-2.amazonaws.com/human-pangenomics/index.html?prefix=NHGRI_UCSC_panel/HG00733/nanopore/Guppy_4.2.2/; Illumina reads, https://www.ncbi.nlm.nih.gov/sra/ERX4439205 [accn] and https://www.ncbi.nlm.nih.gov/sra/ERX4439180 [accn]; HiFi reads, https://www.ncbi.nlm.nih.gov/sra/ERX3831682; Hi-C reads, https://trace.ncbi.nlm.nih.gov/Traces/sra/?run=SRR11347815. HG003 data for HG002 (trio) are taken from https://ftp-trace.ncbi.nlm.nih.gov/ReferenceSamples/giab/data_indexes/AshkenazimTrio/sequence.index.AJtrio_Illumina300X_wgs_07292015_updated.HG003. HG004 data for HG002 (trio) are taken from https://ftp-trace.ncbi.nlm.nih.gov/ReferenceSamples/giab/data_indexes/AshkenazimTrio/sequence.index.AJtrio_Illumina300X_wgs_07292015_updated.HG004. HG00733 trio data (HG00731 and HG00732) are taken from 1000 genomes database https://www.internationalgenome.org/data-portal/sample/HG00732 and https://www.internationalgenome.org/data-portal/sample/HG00731.
Code availability
Hypo-assembler, related pipelines, and evaluation scripts are available on GitHub: https://github.com/kensung-lab/hypo-assembler.
References
Garg, S. et al. Chromosome-scale, haplotype-resolved assembly of human genomes. Nat. Biotechnol. 39, 309–312 (2021).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Nurk, S. et al. HiCanu: accurate assembly of segmental duplications, satellites, and allelic variants from high-fidelity long reads. Genome Res. 30, 1291–1305 (2020).
Luo, X., Kang, X. & Schönhuth, A. phasebook: haplotype-aware de novo assembly of diploid genomes from long reads. Genome Biol. 22, 299 (2021).
Chin, C.-S. & Khalak, A. Human genome assembly in 100 minutes. Preprint at bioRxiv https://doi.org/10.1101/705616 (2019).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 37, 540–546 (2019).
Vaser, R. & Šikić, M. Time- and memory-efficient genome assembly with Raven. Nat. Comput. Sci. 1, 332–336 (2021).
Ruan, J. & Li, H. Fast and accurate long-read assembly with wtdbg2. Nat. Methods 17, 155–158 (2020).
Chen, Y. et al. Efficient assembly of nanopore reads via highly accurate and intact error correction. Nat. Commun. 12, 60 (2021).
Lang, D. et al. Comparison of the two up-to-date sequencing technologies for genome assembly: HiFi reads of Pacific Biosciences Sequel II system and ultralong reads of Oxford Nanopore. Gigascience 9, giaa123 (2020).
Warren, R. L. et al. ntedit: scalable genome sequence polishing. Bioinformatics 35, 4430–4432 (2019).
Zimin, A. V. & Salzberg, S. L. The genome polishing tool POLCA makes fast and accurate corrections in genome assemblies. PLoS Comput. Biol. 16, e1007981 (2020).
Aury, J.-M. & Istace, B. Hapo-G, haplotype-aware polishing of genome assemblies with accurate reads. NAR Genom. Bioinform. 3, lqab034 (2021).
Shafin, K. et al. Haplotype-aware variant calling with PEPPER-Margin-DeepVariant enables high accuracy in nanopore long-reads. Nat. Methods 18, 1322–1332 (2021).
Shafin, K. et al. Nanopore sequencing and the shasta toolkit enable efficient de novo assembly of eleven human genomes. Nat. Biotechnol. 38, 1044–1053 (2020).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Vaser, R., Sović, I., Nagarajan, N. & Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746 (2017).
Hu, J., Fan, J., Sun, Z. & Liu, S. NextPolish: a fast and efficient genome polishing tool for long-read assembly. Bioinformatics 36, 2253–2255 (2020).
Formenti, G. et al. Merfin: improved variant filtering, assembly evaluation and polishing via k-mer validation. Nat. Methods 19, 696–704 (2022).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://doi.org/10.48550/arXiv.1303.3997 (2013).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Rajaby, R. et al. INSurVeyor: improving insertion calling from short read sequencing data. Nat. Commun. 14, 3243 (2023).
Rajaby, R. & Sung, W.-K. SurVIndel: improving CNV calling from high-throughput sequencing data through statistical testing. Bioinformatics 37, 1497–1505 (2021).
Nurk, S. et al. The complete sequence of a human genome. Science 376, 44–53 (2022).
Vollger, M. R. et al. Long-read sequence and assembly of segmental duplications. Nat. Methods 16, 88–94 (2019).
Logsdon, G. A. et al. The structure, function and evolution of a complete human chromosome 8. Nature 593, 101–107 (2021).
Kronenberg, Z. N. et al. Extended haplotype-phasing of long-read de novo genome assemblies using Hi-C. Nat. Commun. 12, 1935 (2021).
Cheng, H. et al. Haplotype-resolved assembly of diploid genomes without parental data. Nat. Biotechnol. 40, 1332–1335 (2022).
Porubsky, D. et al. Fully phased human genome assembly without parental data using single-cell strand sequencing and long reads. Nat. Biotechnol. 39, 302–308 (2021).
Xie, M. et al. gcaPDA: a haplotype-resolved diploid assembler. BMC Bioinformatics 23, 68 (2022).
Sullivan, L. L. & Sullivan, B. A. Genomic and functional variation of human centromeres. Exp. Cell Res. 389, 111896 (2020).
Mikheenko, A., Bzikadze, A. V., Gurevich, A., Miga, K. H. & Pevzner, P. A. TandemTools: mapping long reads and assessing/improving assembly quality in extra-long tandem repeats. Bioinformatics 36, i75–i83 (2020).
Kim, J.-H. et al. Variation in human chromosome 21 ribosomal RNA genes characterized by tar cloning and long-read sequencing. Nucleic Acids Res. 46, 6712–6725 (2018).
Fiddes, I. T. et al. Comparative annotation toolkit (cat)–simultaneous clade and personal genome annotation. Genome Res. 28, 1029–1038 (2018).
Shumate, A. & Salzberg, S. L. Liftoff: accurate mapping of gene annotations. Bioinformatics 37, 1639–1643 (2021).
Ceballos, F. C., Joshi, P. K., Clark, D. W., Ramsay, M. & Wilson, J. F. Runs of homozygosity: windows into population history and trait architecture. Nat. Rev. Genet. 19, 220–234 (2018).
Ariyaratne, P. N. & Sung, W.-K. Pe-assembler: de novo assembler using short paired-end reads. Bioinformatics 27, 167–174 (2011).
Sudmant, P. H. et al. Diversity of human copy number variation and multicopy genes. Science 330, 641–646 (2010).
Carvalho, A. B., Dupim, E. G. & Goldstein, G. Improved assembly of noisy long reads by k-mer validation. Genome Res. 26, 1710–1720 (2016).
Miga, K. H. et al. Telomere-to-telomere assembly of a complete human X chromosome. Nature 585, 79–84 (2020).
Kundu, R., Casey, J. & Sung, W.-K. Hypo: super fast & accurate polisher for long read genome assemblies. Preprint at bioRXiv https://doi.org/10.1101/2019.12.19.882506 (2019).
Stanke, M. et al. Augustus: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439 (2006).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Acknowledgements
The authors received no specific funding for this work.
Author information
Authors and Affiliations
Contributions
J.C. did the computations and experiments. R.R. developed variant callers that are crucial to the result. R.K. developed the early versions of the hypo polisher that become the starting point of the work. W.-K.S. supervises the work and encouraged the idea of solid k-mers. All authors discussed the results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks Kai Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Lei Tang and Lin Tang, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CHM13 assembly benchmark.
Several measures of qualities of assembly of CHM13. (a) The number of errors per 100kbp and (b) YAK’s estimated quality value of various assemblies of CHM13.
Extended Data Fig. 2 HG002 assembly benchmark.
Several measures of qualities for diploid assemblies of HG002. (a) The number of errors per 100kbp. (b) YAK’s estimated quality value. (c) Kmer Mixture Rate (d) Switch Error Rate.
Supplementary information
Supplementary Information
Supplementary Sections A–O, Figs. 1–29 and Tables 1–30.
Supplementary Dataset
Data used to generate the box plots in Supplementary Figs. 16 and 17.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Darian, J.C., Kundu, R., Rajaby, R. et al. Constructing telomere-to-telomere diploid genome by polishing haploid nanopore-based assembly. Nat Methods 21, 574–583 (2024). https://doi.org/10.1038/s41592-023-02141-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-023-02141-1