Abstract
As animals navigate, they must identify features within context. In the mammalian brain, the hippocampus has the ability to separately encode different environmental contexts, even when they share some prominent features. To do so, neurons respond to sensory features in a context-dependent manner; however, it is not known how this encoding emerges. To examine this, we performed electrical recordings in the hippocampus as mice navigated in two distinct virtual environments. In CA1, both synaptic input to single neurons and population activity strongly tracked visual cues in one environment, whereas responses were almost completely absent when the same cue was presented in a second environment. A very similar, highly context-dependent pattern of cue-driven spiking was also observed in CA3. These results indicate that CA1 inherits a complex spatial code from upstream regions, including CA3, that have already computed a context-dependent representation of environmental features.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Source data for statistics in all figures are accessible through the following figShare link: https://figshare.com/articles/Zhao_et_al_2020_SourceData_zip/12071226.
Code availability
All analyses were conducted with customized MATLAB (R2014) codes. All raw data and analysis codes are archived on the Janelia Research Campus server and are available upon reasonable request.
References
Hartley, T., Lever, C., Burgess, N. & O’Keefe, J. Space in the brain: how the hippocampal formation supports spatial cognition. Philos. Trans. R. Soc. Lond. B Biol. Sci. 369, 20120510 (2014).
O’Keefe, J. & Dostrovsky, J. The hippocampus as a spatial map. Preliminary evidence from unit activity in the freely-moving rat. Brain Res. 34, 171–175 (1971).
Muller, R. U. & Kubie, J. L. The effects of changes in the environment on the spatial firing of hippocampal complex-spike cells. J. Neurosci. 7, 1951–1968 (1987).
Yoganarasimha, D. & Knierim, J. J. Coupling between place cells and head direction cells during relative translations and rotations of distal landmarks. Exp. Brain Res. 160, 344–359 (2005).
Rolls, E. T. & O’Mara, S. M. View-responsive neurons in the primate hippocampal complex. Hippocampus 5, 409–424 (1995).
Phillips, R. G. & LeDoux, J. E. Differential contribution of amygdala and hippocampus to cued and contextual fear conditioning. Behav. Neurosci. 106, 274–285 (1992).
Deshmukh, S. S. & Knierim, J. J. Influence of local objects on hippocampal representations: landmark vectors and memory. Hippocampus 23, 253–267 (2013).
Shapiro, M. L., Tanila, H. & Eichenbaum, H. Cues that hippocampal place cells encode: dynamic and hierarchical representation of local and distal stimuli. Hippocampus 7, 624–642 (1997).
Fyhn, M., Hafting, T., Treves, A., Moser, M. B. & Moser, E. I. Hippocampal remapping and grid realignment in entorhinal cortex. Nature 446, 190–194 (2007).
Lisman, J. E. Relating hippocampal circuitry to function: recall of memory sequences by reciprocal dentate–CA3 interactions. Neuron 22, 233–242 (1999).
Kaifosh, P. & Losonczy, A. Mnemonic functions for nonlinear dendritic integration in hippocampal pyramidal circuits. Neuron 90, 622–634 (2016).
Ito, H. T., Zhang, S. J., Witter, M. P., Moser, E. I. & Moser, M. B. A prefrontal-thalamo-hippocampal circuit for goal-directed spatial navigation. Nature 522, 50–55 (2015).
Geiller, T., Fattahi, M., Choi, J. S. & Royer, S. Place cells are more strongly tied to landmarks in deep than in superficial CA1. Nat. Commun. 8, 14531 (2017).
Nakashiba, T., Young, J. Z., McHugh, T. J., Buhl, D. L. & Tonegawa, S. Transgenic inhibition of synaptic transmission reveals role of CA3 output in hippocampal learning. Science 319, 1260–1264 (2008).
Harvey, C. D., Collman, F., Dombeck, D. A. & Tank, D. W. Intracellular dynamics of hippocampal place cells during virtual navigation. Nature 461, 941–946 (2009).
Cohen, J. D., Bolstad, M. & Lee, A. K. Experience-dependent shaping of hippocampal CA1 intracellular activity in novel and familiar environments. eLife 6, e233040 (2017).
Gauthier, J. L. & Tank, D. W. A dedicated population for reward coding in the hippocampus. Neuron 99, 179–193 (2018).
Fuhrmann, F. et al. Locomotion, theta oscillations, and the speed-correlated firing of hippocampal neurons are controlled by a medial septal glutamatergic circuit. Neuron 86, 1253–1264 (2015).
Kropff, E., Carmichael, J. E., Moser, M. B. & Moser, E. I. Speed cells in the medial entorhinal cortex. Nature 523, 419–424 (2015).
Danielson, N. B. et al. Sublayer-specific coding dynamics during spatial navigation and learning in hippocampal area CA1. Neuron 91, 652–665 (2016).
Aghajan, Z. M. et al. Impaired spatial selectivity and intact phase precession in two-dimensional virtual reality. Nat. Neurosci. 18, 121–128 (2015).
Bittner, K. C. et al. Conjunctive input processing drives feature selectivity in hippocampal CA1 neurons. Nat. Neurosci. 18, 1133–1142 (2015).
Bittner, K. C., Milstein, A. D., Grienberger, C., Romani, S. & Magee, J. C. Behavioral time scale synaptic plasticity underlies CA1 place fields. Science 357, 1033–1036 (2017).
Wang, Y., Romani, S., Lustig, B., Leonardo, A. & Pastalkova, E. Theta sequences are essential for internally generated hippocampal firing fields. Nat. Neurosci. 18, 282–288 (2015).
Bloss, E. B. et al. Structured dendritic inhibition supports branch-selective integration in CA1 pyramidal cells. Neuron 89, 1016–1030 (2016).
Golding, N. L., Mickus, T. J., Katz, Y., Kath, W. L. & Spruston, N. Factors mediating powerful voltage attenuation along CA1 pyramidal neuron dendrites. J. Physiol. 568, 69–82 (2005).
Nicholson, D. A. et al. Distance-dependent differences in synapse number and AMPA receptor expression in hippocampal CA1 pyramidal neurons. Neuron 50, 431–442 (2006).
Bittner, K. C., Andrasfalvy, B. K. & Magee, J. C. Ion channel gradients in the apical tuft region of CA1 pyramidal neurons. PLoS ONE 7, e46652 (2012).
Brun, V. H. et al. Place cells and place recognition maintained by direct entorhinal–hippocampal circuitry. Science 296, 2243–2246 (2002).
Alme, C. B. et al. Place cells in the hippocampus: eleven maps for eleven rooms. Proc. Natl Acad. Sci. USA 111, 18428–18435 (2014).
Wilson, M. A. & McNaughton, B. L. Dynamics of the hippocampal ensemble code for space. Science 261, 1055–1058 (1993).
Grienberger, C., Milstein, A. D., Bittner, K. C., Romani, S. & Magee, J. C. Inhibitory suppression of heterogeneously tuned excitation enhances spatial coding in CA1 place cells. Nat. Neurosci. 20, 417–426 (2017).
Morris, R. Spatial localization does not require the presence of local cues. Learn. Motiv. 12, 239–260 (1981).
Jayakumar, R. P. et al. Recalibration of path integration in hippocampal place cells. Nature 566, 533–537 (2019).
Anderson, M. I. & Jeffery, K. J. Heterogeneous modulation of place cell firing by changes in context. J. Neurosci. 23, 8827–8835 (2003).
Gulli, R. A. et al. Context-dependent representations of objects and space in the primate hippocampus during virtual navigation. Nat. Neurosci. 23, 103–112 (2020).
McHugh, T. J. & Tonegawa, S. CA3 NMDA receptors are required for the rapid formation of a salient contextual representation. Hippocampus 19, 1153–1158 (2009).
Middleton, S. J. & McHugh, T. J. Silencing CA3 disrupts temporal coding in the CA1 ensemble. Nat. Neurosci. 19, 945–951 (2016).
Deshmukh, S. S. & Knierim, J. J. Representation of non-spatial and spatial information in the lateral entorhinal cortex. Front. Behav. Neurosci. 5, 69 (2011).
Losonczy, A. & Magee, J. C. Integrative properties of radial oblique dendrites in hippocampal CA1 pyramidal neurons. Neuron 50, 291–307 (2006).
Hsu, C. L., Zhao, X., Milstein, A. D. & Spruston, N. Persistent sodium current mediates the steep voltage dependence of spatial coding in hippocampal pyramidal neurons. Neuron 99, 147–162 (2018).
Golding, N. L., Staff, N. P. & Spruston, N. Dendritic spikes as a mechanism for cooperative long-term potentiation. Nature 418, 326–331 (2002).
Takahashi, H. & Magee, J. C. Pathway interactions and synaptic plasticity in the dendritic tuft regions of CA1 pyramidal neurons. Neuron 62, 102–111 (2009).
Megías, M., Emri, Z., Freund, T. F. & Gulyás, A. I. Total number and distribution of inhibitory and excitatory synapses on hippocampal CA1 pyramidal cells. Neuroscience 102, 527–540 (2001).
Davoudi, H. & Foster, D. J. Acute silencing of hippocampal CA3 reveals a dominant role in place field responses. Nat. Neurosci. 22, 337–342 (2019).
Ravassard, P. et al. Multisensory control of hippocampal spatiotemporal selectivity. Science 340, 1342–1346 (2013).
Bellmund, J. L. S., Gärdenfors, P., Moser, E. I. & Doeller, C. F. Navigating cognition: spatial codes for human thinking. Science 362, eaat6766 (2018).
Wood, E. R., Dudchenko, P. A., Robitsek, R. J. & Eichenbaum, H. Hippocampal neurons encode information about different types of memory episodes occurring in the same location. Neuron 27, 623–633 (2000).
Frank, L. M., Brown, E. N. & Wilson, M. Trajectory encoding in the hippocampus and entorhinal cortex. Neuron 27, 169–178 (2000).
Acknowledgements
We thank M. Bolstad, S. Sawtelle, A. Lee and J. Cohen for their help with the design and manufacture of the virtual reality system. We thank K. Bittner for her development of intracellular recording techniques. We thank all Magee and Spruston lab members for insightful discussions. This study was supported by the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
X.Z., N.S. and J.C.M. designed experiments. X.Z. performed in vivo whole-cell recordings. X.Z. and Y.W. performed extracellular recordings. X.Z. and Y.W. analyzed the data with input from N.S. and J.C.M. X.Z., N.S. and J.C.M. wrote the manuscript with input from Y.W.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Neuroscience thanks Laura Colgin, Debora Ledergerber, Aman Saleem and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Virtual tracks with distinguishable visual cues.
Oval track a, oval track with barriers b, and triangular track c, used throughout this study. Besides the common cue (vertical gratings) shared on both tracks, the triangular track contained another 2 cues (45° gratings and rings) that were not present on the oval track.
Extended Data Fig. 2 Detailed characterization of place fields during manipulations of isolated cues.
a, Position-variant running speeds on oval tracks with (blue, n = 12 recording session from 7 animals) or without (black, n = 19 recording session from 14 animals) virtual curtains in the fixed reward paradigm (reward locations marked as red lines). Although the overall averaged speeds slightly differ under the two conditions due to variations among individual animals, their position dependencies are very similar, likely determined by the turning and reward locations. b, Position-variant running speed on the oval track with random reward delivery (n = 4 sessions from 3 animals). Dashed lines depict location 1 and 4, as in Fig. 1. c, Position-variant running speed on the triangular track (red, n = 8 sessions from 6 animals). Speed on the oval track with virtual curtains was plotted as comparison (blue). Shadings depict the location where the same cue was available on both tracks. In all panels, population data are presented as mean±s.e.m.
Extended Data Fig. 3 Cue duplication induced a second place field with smaller amplitude without virtual curtains.
a–d, Population averages and statistics of theta amplitudes and spiking rates following the cue duplication (n = 10 cells from 7 animals in a and b, 9 cells from 7 animals in e and f). All data are presented as mean±s.e.m. The same color conventions as in Fig. 3. In (c): P = 1.936e-5 for control loc. 1 vs. control loc. 4, 0.641 for duplication loc. 1 vs. duplication loc. 4, 0.0564 for control loc. 1 vs. duplication loc. 1, and 3.451e-4 for control loc. 4 vs. duplication loc. 4. In (d): P = 3.97e-5 for control loc. 1 vs. control loc. 4, 0.5135 for duplication loc. 1 vs. duplication loc. 4, 0.1553 for control loc. 1 vs. duplication loc. 1, and 2.04e-4 for control loc. 4 vs. duplication loc. 4. e–h, Population averages and statistics of theta amplitudes and spiking rates following the cue shift. Paired student t-test was conducted in all analyses. In (g): P = 2.744e-5 for control loc. 1 vs. control loc. 4, 1.85e-4 for shift loc. 1 vs. shift loc. 4, 2.454e-4 for control loc. 1 vs. shift loc. 1, 0.0632 for control loc. 1 vs. shift loc. 4, 9.723e-4 for control loc. 4 vs. shift loc. 4, and 0.2953 for control loc. 4 vs. shift loc. 1. In (h): P = 1.959e-4 for control loc. 1 vs. control loc. 4, 0.0035 for shift loc. 1 vs. shift loc. 4, 2.667e-4 for control loc. 1 vs. shift loc. 1, 0.2406 for control loc. 1 vs. shift loc. 4, 0.0034 for control loc. 4 vs. shift loc. 4, and 0.4954 for control loc. 4 vs. shift loc. 1. Solid curves and shaded areas depict mean±s.e.m. Bar graphs and error bars represent mean and s.e.m., respectively. Paired student t-test (two-sided) was conducted in all statistical analyses. *: P < 0.05, **: P < 0.01, ***: P < 0.001, N.S.: not significant (P > = 0.05).
Extended Data Fig. 4 Effects of cue duplication on CA1 place fields with random reward delivery.
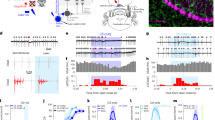
a, An example cell with the duplication of cue A during recording. b, Ramp depolarization, theta amplitude and spiking rate before (black) and after (red) the cue duplication in the example cell shown in (a). c–e, Position-variant ramp depolarization (c), theta amplitude (d) and spiking rate (e) before (black) and after (red) the cue duplication (n = 12 cells from 7 animals). Solid curves and shaded areas depict mean±s.e.m. Through (b) to (e), locations of cue A and D were marked with light blue and yellow shadings, respectively, as in Fig. 1. f–h, Statistical comparisons of peak ramp depolarization (f), theta amplitude (g) and spiking rate (h) following the cue duplication. Peak values of each metric were calculated within location 1 (between the two light blue lines in (c)-(e); grey for control, pink for cue duplication) and location 4 (between the two yellow lines in (c)-(e); black for control, red for cue duplication). Ramp depolarization (f): P = 1.485e-7 for control, loc. 1 vs. loc. 4; P = 0.0027 for duplication, loc. 1 vs. loc. 4; P = 0.9783 for control loc. 1 vs. duplication loc. 1; P = 0.001 for control loc. 4 vs. duplication loc. 4. Theta amplitude (g): P = 1.029e-5 for control, loc. 1 vs. loc. 4; P = 0.0061 for duplication, loc. 1 vs. loc. 4; P = 0.7968 for control loc. 1 vs. duplication loc. 1; P = 0.0116 for control loc. 4 vs. duplication loc. 4. Spiking rate (h): P = 8.163e-7 for control, loc. 1 vs. loc. 4; P = 0.0024 for duplication, loc. 1 vs. loc. 4; P = 0.1988 for control loc. 1 vs. duplication loc. 1; P = 0.0083 for control loc. 4 vs. duplication loc. 4 (n = 12 cells from 7 animals). Solid curves and shaded areas depict mean±s.e.m. Bar graphs and error bars represent mean and s.e.m., respectively. Paired student t-test (2-sided) was conducted in all analyses. *: P < 0.05, **: P < 0.01, ***: P < 0.001, N.S.: not significant (P > = 0.05).
Extended Data Fig. 5 Spatial coding in the hippocampus following cue manipulations.
a, Behavior of an example animal. Two rewards (red) were delivered in each lap at random locations. Note that the animal licked (blue) along the entire track. b–d, Population averages and statistics of ramp depolarizations, theta amplitudes and spiking rates following the cue duplication (n = 4 cells from 3 animals). All data are presented as mean±s.e.m. The same color conventions as in Fig. 2. Paired student t-test (2-sided) was conducted in all analyses. In e,: P = 0.0204 for control, loc. 1 vs. loc. 4; P = 0.2255 for duplication, loc. 1 vs. loc. 4; P = 0.6152 for control loc. 1 vs. duplication loc. 1; P = 0.2318 for control loc. 1 vs. duplication loc. 4; P = 0.0019 for control loc. 4 vs. duplication loc. 4. In f,: P = 0.0011 for control, loc. 1 vs. loc. 4; P = 0.0794 for duplication, loc. 1 vs. loc. 4; P = 0.9538 for control loc. 1 vs. duplication loc. 1; P = 0.3426 for control loc. 1 vs. duplication loc. 4; P = 0.0169 for control loc. 4 vs. duplication loc. 4. In g,: P = 0.0091 for control, loc. 1 vs. loc. 4; P = 0.2591 for duplication, loc. 1 vs. loc. 4; P = 0.6522 for control loc. 1 vs. duplication loc. 1; P = 0.5469 for control loc. 1 vs. duplication loc. 4; P = 0.0216 for control loc. 4 vs. duplication loc. 4. Bar graphs and error bars represent mean and s.e.m., respectively.
Extended Data Fig. 6 Spatial coding in the hippocampus following cue manipulations.
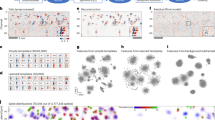
a, Rate map of simultaneously recorded CA1 place cells (n = 35 cells) on the oval track in one session during cue duplication and shift. Cells were sorted with the center of mass of their place fields. Note that red boxes labeled cells that decreased their firing rates at location 4 but increased firing at location 1 after cue A was shifted from 1 to 4. b, Population vector-based decoding using the control rate map as the base. Decoding error under the cue duplication condition was plotted against the actual position on the track. Location 1 and 4 were marked by blue and yellow shadings, respectively. c, Decoding errors under the cue shift condition. d–f, The same plots as (a–c) with another CA1 recording session (n = 29 cells). g–l, The same as in (a–c) with two CA3 recording sessions (n = 18 and 23 cells, respectively).
Extended Data Fig. 7 Isolation of single units from extracellular recordings in CA3.
a, Representative histology confirmation of extracellular recordings in CA3 using silicon probes. b, Recording paradigm. Most of recording sessions (n = 7/9) in this report included 5 epochs. Two cue manipulation epochs (duplication and shift) were sandwiched between control epochs. The rest 2 sessions only included the cue shift epoch and two control epochs. c, Spike clustering based on principle component analysis (PCA) of spike waveforms. Red and blue dots are two isolated spike clusters, while grey dots represent all other spikes (364694 spikes were detected in this session. A randomly sampled 10% of them were plotted). d, Color coded rate maps of the two sorted units shown in c (color scale: 0–45 Hz). Unit 1 is a putative interneuron (spatial selectivity: 0.06). Unit 2 is a place cell (spatial selectivity: 0.72). Boundaries between epochs are marked by cyan lines. Vertical white lines mark the locations of cue A and B, as in Extended Fig. 6. In this recording session, the second and fourth sessions were cue A’s duplication and shift, respectively, as labeled on the right. e, and f, Example sessions of simultaneously recorded place cells in CA3 during shift of cue A (e) or B (f). Spiking rate of each cell was normalized to its peak rate under the control condition. Cells were sorted based on center of mass of their firing field under the control condition.
Extended Data Fig. 8 Identification of place fields in CA3 units.
a, Position-variant spiking rates (black) were fitted with skewed von Mises equations (red). 20% peak amplitude in the fitted curve was used to define the beginning and end of place fields (red circles). Two examples were shown with narrow and wide fields, respectively. b, Rate map of all 129 cells (from 3 animals) included in Fig. 4 (under the control condition). Spiking rate of each cell was normalized by its peak. Cells were sorted by the center of mass of their place fields. For each cell, white and magenta ticks marked the beginning and end of its place field, determined by the method described in a. c, Distribution of place field width. The median field width is 36.4 cm.
Extended Data Fig. 9 Switch between oval and triangle tracks induced global remapping in CA3 and CA1.
a–d, CA1 recordings. Cells that are qualified as place cells (spatial selectivity index>0.3, peak firing rate>1 Hz) in either environment are included in this analysis (n = 283 cells). Cells are pooled from 4 recording sessions, 3 animals. (a) Normalized firing rates of cells sorted by the center of mass (COM) of each cell’s firing activity on the oval track (left). Normalized firing rates of cells with the same ranking (based on oval track activities) on the triangle track (right). (b) Distances of firing activity COM (ΔCOM) between pairs of cells, sorted in the same way as in (a). As expected from the sorting method, ΔCOM has small values along the diagonal of the pairwise matrix on the oval track (left). However, this diagonal pattern disappears on the triangle track (right). (c) ΔCOM on the triangle track was plotted against that on the oval track (blue, mean±s.e.m.). The black line is the averaged ΔCOM on the triangle track. ΔCOM (oval) was binned with 5 cm. ΔCOM on the triangle track shows a nearly flat line around the averaged value. There is a slight trend that pairs with smaller ΔCOM on the oval track tend to have smaller ΔCOM on the triangle track, indicating some maintained topological structures. However, this effect is very weak as the maximal distance between the observed ΔCOM (triangle) is just around 0.1×S.D. from the mean (shown as z-score in d). (d) Z-scores of ΔCOM (triangle) at each bin, compared with the mean value of all ΔCOM (triangle) (the black line in c). e–h, The same as in (a–d) with CA3 recordings (n = 243 cells). Cells are pooled from 3 recording sessions, 3 animals.
Extended Data Fig. 10 CA1 interneurons do not show context-dependent firing.
a, Bimodal distribution of spike width of extracellularly recorded units (n = 558 from 4 animals, see Methods for details). Spike width less than 0.775 ms (dash line) was used as a criterion for fast spiking interneurons. b, Mean spiking rate inversely correlated with spike width. Putative fast spiking interneurons (red) were identified as spike width<0.775 ms and mean spiking rate > 1 Hz. Inset: averaged spike waveform of putative fast spiking interneurons (red) and pyramidal cells (black). Note that spikes from putative fast spiking interneurons decayed faster. SD was plotted as shading areas. Scale bar: 0.5 ms. The vast majority of recorded units (n = 455/558 units) show spike waveforms that exhibit larger negative peaks than the positive one. Only these units were used to calculate the inset figures to avoid distortion of spike waveforms when both polarities were averaged together. c, Spatial rate map of putative fast spiking interneurons. Cells were ranked based on the COM of their firing fields. d, Averaged population activity of putative fast spiking interneurons on oval (black) and triangular (red) tracks. e, Mean spiking rates on oval and triangular tracks (within the shared cue region, between grey lines in e) are not significantly different in fast spiking interneurons (p = 0.471, n = 66 cells from 4 animals). f–i, The same as (b–e) with more strictly defined putative fast spiking interneurons (units distributed at the upper-left corner in f). Mean spiking rates of this population were not dependent on context either (p = 0.8939, n = 17 cells from 4 animals). Solid curves and shaded areas depict mean±s.e.m. Bar graphs and error bars represent mean and s.e.m., respectively. Paired student t-test (two-sided) was conducted in all statistical analyses. N.S.: not significant (P > = 0.05).
Supplementary information
Rights and permissions
About this article
Cite this article
Zhao, X., Wang, Y., Spruston, N. et al. Membrane potential dynamics underlying context-dependent sensory responses in the hippocampus. Nat Neurosci 23, 881–891 (2020). https://doi.org/10.1038/s41593-020-0646-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-0646-2
This article is cited by
-
A consistent map in the medial entorhinal cortex supports spatial memory
Nature Communications (2024)
-
Latent representations in hippocampal network model co-evolve with behavioral exploration of task structure
Nature Communications (2024)
-
Entorhinal cortex directs learning-related changes in CA1 representations
Nature (2022)
-
Prefrontal feature representations drive memory recall
Nature (2022)
-
The functional organization of excitatory synaptic input to place cells
Nature Communications (2021)