Abstract
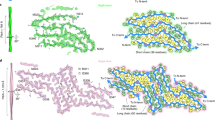
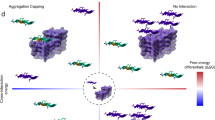
Proteins in the fibrous amyloid state are a major hallmark of neurodegenerative disease. Understanding the multiple conformations, or polymorphs, of amyloid proteins at the molecular level is a challenge of amyloid research. Here, we detail the wide range of polymorphs formed by a segment of human TAR DNA-binding protein 43 (TDP-43) as a model for the polymorphic capabilities of pathological amyloid aggregation. Using X-ray diffraction, microelectron diffraction (MicroED) and single-particle cryo-EM, we show that the 247DLIIKGISVHI257 segment from the second RNA-recognition motif (RRM2) forms an array of amyloid polymorphs. These associations include seven distinct interfaces displaying five different symmetry classes of steric zippers. Additionally, we find that this segment can adopt three different backbone conformations that contribute to its polymorphic capabilities. The polymorphic nature of this segment illustrates at the molecular level how amyloid proteins can form diverse fibril structures.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Chiti, F. & Dobson, C. M. Protein misfolding, functional amyloid, and human disease. Annu. Rev. Biochem. 75, 333–366 (2006).
Hardy, J. & Selkoe, D. J. The amyloid hypothesis of Alzheimer’s disease: progress and problems on the road to therapeutics. Science 297, 353–356 (2002).
Knowles, T. P. J., Vendruscolo, M. & Dobson, C. M. The amyloid state and its association with protein misfolding diseases. Nat. Rev. Mol. Cell. Biol. 15, 384–396 (2014).
Greenwald, J. & Riek, R. Biology of amyloid: structure, function, and regulation. Structure. 18, 1244–1260 (2010).
Goldsbury, C., Frey, P., Olivieri, V., Aebi, U. & Müller, S. A. Multiple assembly pathways underlie amyloid-β fibril polymorphisms. J. Mol. Biol. 352, 282–298 (2005).
Paravastu, A. K., Leapman, R. D., Yau, W.-M. & Tycko, R. Molecular structural basis for polymorphism in Alzheimer’s β-amyloid fibrils. Proc. Natl Acad. Sci. USA 105, 18349–18354 (2008).
Wiltzius, J. J. W. et al. Molecular mechanisms for protein-encoded inheritance. Nat. Struct. Mol. Biol. 16, 973–978 (2009).
Nelson, R. et al. Structure of the cross-β spine of amyloid-like fibrils. Nature 435, 773–778 (2005).
Sawaya, M. R. et al. Atomic structures of amyloid cross-beta spines reveal varied steric zippers. Nature 447, 453–457 (2007).
Stroud, J. C. The zipper groups of the amyloid state of proteins. Acta. Crystallogr. D. Biol. Crystallogr. 69, 540–545 (2013).
Tuttle, M. D. et al. Solid-state NMR structure of a pathogenic fibril of full-length human α-synuclein. Nat. Struct. Mol. Biol. 23, 409–415 (2016).
Wälti, M. A. et al. Atomic-resolution structure of a disease-relevant Aβ(1-42) amyloid fibril. Proc. Natl Acad. Sci. USA 113, E4976–E4984 (2016).
Colvin, M. T. et al. Atomic resolution structure of monomorphic Aβ42 amyloid fibrils. J. Am. Chem. Soc. 138, 9663–9674 (2016).
Petkova, A. T. et al. A structural model for Alzheimer’s β-amyloid fibrils based on experimental constraints from solid state NMR. Proc. Natl Acad. Sci. USA 99, 16742–16747 (2002).
Xiao, Y. et al. Aβ(1-42) fibril structure illuminates self-recognition and replication of amyloid in Alzheimer’s disease. Nat. Struct. Mol. Biol. 22, 499–505 (2015).
Schmidt, A., Annamalai, K., Schmidt, M., Grigorieff, N. & Fändrich, M. Cryo-EM reveals the steric zipper structure of a light chain-derived amyloid fibril. Proc. Natl Acad. Sci. USA 113, 6200–6205 (2016).
Schmidt, M. et al. Peptide dimer structure in an Aβ(1-42) fibril visualized with cryo-EM. Proc. Natl Acad. Sci. USA 112, 11858–11863 (2015).
Bai, X. C. et al. An atomic structure of human γ-secretase. Nature 525, 212–217 (2015).
Tanaka, M., Chien, P., Naber, N., Cooke, R. & Weissman, J. S. Conformational variations in an infectious protein determine prion strain differences. Nature 428, 323–328 (2004).
Kaufman, S. K. et al. Tau prion strains dictate patterns of cell pathology, progression rate, and regional vulnerability in vivo. Neuron 92, 796–812 (2016).
Riek, R. & Eisenberg, D. S. The activities of amyloids from a structural perspective. Nature 539, 227–235 (2016).
Eisenberg, D. & Jucker, M. The amyloid state of proteins in human diseases. Cell 148, 1188–1203 (2012).
Krotee, P. et al. Atomic structures of fibrillar segments of hIAPP suggest tightly mated β-sheets are important for cytotoxicity. eLife 6, 1–26 (2017).
Soriaga, A. B., Sangwan, S., Macdonald, R., Sawaya, M. R. & Eisenberg, D. Crystal structures of IAPP amyloidogenic segments reveal a novel packing motif of out-of-register beta sheets. J. Phys. Chem. B. 120, 5810–5816 (2016).
Colletier, J.-P. et al. Molecular basis for amyloid-beta polymorphism. Proc. Natl Acad. Sci. USA 108, 16938–16943 (2011).
Neumann, M. et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314, 130–133 (2006).
Hasegawa, M. et al. Phosphorylated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Ann. Neurol. 64, 60–70 (2008).
Schwab, C., Arai, T., Hasegawa, M., Yu, S. & McGeer, P. L. Colocalization of transactivation-responsive DNA-binding protein 43 and huntingtin in inclusions of Huntington disease. J. Neuropathol. Exp. Neurol. 67, 1159–1165 (2008).
Amador-Ortiz, C. et al. TDP-43 immunoreactivity in hippocampal sclerosis and Alzheimer’s disease. Ann. Neurol. 61, 435–445 (2007).
Nakashima-Yasuda, H. et al. Co-morbidity of TDP-43 proteinopathy in Lewy body related diseases. Acta. Neuropathol. 114, 221–229 (2007).
Higashi, S. et al. Concurrence of TDP-43, tau and α-synuclein pathology in brains of Alzheimer’s disease and dementia with Lewy bodies. Brain. Res. 1184, 284–294 (2007).
Kraemer, B. C. et al. Loss of murine TDP-43 disrupts motor function and plays an essential role in embryogenesis. Acta. Neuropathol. 119, 409–419 (2010).
Sephton, C. F. et al. TDP-43 is a developmentally regulated protein essential for early embryonic development. J. Biol. Chem. 285, 6826–6834 (2010).
Wu, L.-S. S. et al. TDP-43, a neuro-pathosignature factor, is essential for early mouse embryogenesis. Genesis 48, 56–62 (2010).
Chiang, P. M. et al. Deletion of TDP-43 down-regulates Tbc1d1, a gene linked to obesity, and alters body fat metabolism. Proc. Natl Acad. Sci. USA 107, 16320–16324 (2010).
Zhu, L. et al. An ALS-mutant TDP-43 neurotoxic peptide adopts an anti-parallel β-structure and induces TDP-43 redistribution. Hum. Mol. Genet. 23, 6863–6877 (2014).
Igaz, L. M. et al. Expression of TDP-43 C-terminal fragments in vitro recapitulates pathological features of TDP-43 proteinopathies. J. Biol. Chem. 284, 8516–8524 (2009).
Iguchi, Y. et al. TDP-43 depletion induces neuronal cell damage through dysregulation of Rho family GTPases. J. Biol. Chem. 284, 22059–22066 (2009).
Di Carlo, V. et al. TDP-43 regulates the microprocessor complex activity during in vitro neuronal differentiation. Mol. Neurobiol. 48, 952–963 (2013).
Soragni, A. et al. A designed inhibitor of p53 aggregation rescues p53 tumor suppression in ovarian carcinomas. Cancer Cell 29, 90–103 (2016).
Guo, W. et al. An ALS-associated mutation affecting TDP-43 enhances protein aggregation, fibril formation and neurotoxicity. Nat. Struct. Mol. Biol. 18, 822–830 (2011).
Jiang, L. L. et al. Structural transformation of the amyloidogenic core region of TDP-43 protein initiates its aggregation and cytoplasmic inclusion. J. Biol. Chem. 288, 19614–19624 (2013).
Jiang, L.-L. et al. Two mutations G335D and Q343R within the amyloidogenic core region of TDP-43 influence its aggregation and inclusion formation. Sci. Rep. 6, 23928 (2016).
Budini, M. et al. Cellular model of TAR DNA-binding protein 43 (TDP-43) aggregation based on its C-terminal Gln/Asn-rich region. J. Biol. Chem. 287, 7512–7525 (2012).
Mompeán, M. et al. “Structural characterization of the minimal segment of TDP-43 competent for aggregation”. Arch. Biochem. Biophys. 545, 53–62 (2014).
Saini, A. & Chauhan, V. S. Self-assembling properties of peptides derived from TDP-43 C-terminal fragment. Langmuir 30, 3845–3856 (2014).
Saini, A. & Chauhan, V. S. Delineation of the core aggregation sequences of TDP-43 C-terminal fragment. Chembiochem. 12, 2495–2501 (2011).
Goldschmidt, L., Teng, P. K., Riek, R. & Eisenberg, D. Identifying the amylome, proteins capable of forming amyloid-like fibrils. Proc. Natl Acad. Sci. USA 107, 3487–3492 (2010).
Kyte, J. & Doolittle, R. F. A simple method for displaying the hydropathic character of a protein. J. Mol. Biol. 157, 105–132 (1982).
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Sunde, M. et al. Common core structure of amyloid fibrils by synchrotron X-ray diffraction. J. Mol. Biol. 273, 729–739 (1997).
Shi, D., Nannenga, B. L., Iadanza, M. G. & Gonen, T. Three-dimensional electron crystallography of protein microcrystals. eLife 2, e01345 (2013).
Nannenga, B. L., Shi, D., Leslie, A. G. W. & Gonen, T. High-resolution structure determination by continuous-rotation data collection in MicroED. Nat. Methods 11, 927–930 (2014).
Liu, S. et al. Atomic resolution structure determination by the cryo-EM method MicroED. Protein. Sci. 26, 8–15 (2017).
de la Cruz, M. J. et al. Atomic-resolution structures from fragmented protein crystals with the cryoEM method MicroED. Nat. Methods 14, 399–402 (2017).
Rodriguez, J. A. et al. Structure of the toxic core of α-synuclein from invisible crystals. Nature 525, 486–490 (2015).
Sept, D., Baker, N. A. & McCammon, J. A. The physical basis of microtubule structure and stability. Protein. Sci. 12, 2257–2261 (2003).
Eisenberg, D. S. & Sawaya, M. R. Structural studies of amyloid proteins at the molecular level. Annu. Rev. Biochem. 86, 69–95 (2017).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta. Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Jiang, L. L. et al. The N-terminal dimerization is required for TDP-43 splicing activity. Sci. Rep. 7, 6196 (2017).
Mompeán, M. et al. Point mutations in the N-terminal domain of transactive response DNA-binding protein 43 kDa (TDP-43) compromise its stability, dimerization, and functions. J. Biol. Chem. 292, 11992–12006 (2017).
Afroz, T. et al. Functional and dynamic polymerization of the ALS-linked protein TDP-43 antagonizes its pathologic aggregation. Nat. Commun. 8, 45 (2017).
Kuo, P. H., Doudeva, L. G., Wang, Y. T., Shen, C. K. J. & Yuan, H. S. Structural insights into TDP-43 in nucleic-acid binding and domain interactions. Nucleic. Acids. Res. 37, 1799–1808 (2009).
Nonaka, T., Kametani, F., Arai, T., Akiyama, H. & Hasegawa, M. Truncation and pathogenic mutations facilitate the formation of intracellular aggregates of TDP-43. Hum. Mol. Genet. 18, 3353–3364 (2009).
Li, Q., Yokoshi, M., Okada, H. & Kawahara, Y. The cleavage pattern of TDP-43 determines its rate of clearance and cytotoxicity. Nat. Commun. 6, 6183 (2015).
Zhang, Y.-J. et al. Aberrant cleavage of TDP-43 enhances aggregation and cellular toxicity. Proc. Natl Acad. Sci. USA 106, 7607–7612 (2009).
Fitzpatrick, A. W. P. et al. Cryo-EM structures of tau filaments from Alzheimer’s disease. Nature 547, 185–190 (2017).
Sievers, S. A. et al. Structure-based design of non-natural amino-acid inhibitors of amyloid fibril formation. Nature 475, 96–100 (2011).
Saelices, L. et al. Uncovering the mechanism of aggregation of human transthyretin. J. Biol. Chem. 290, 28932–28943 (2015).
Caspar, D. L. D. & Cohen, C. Polymorphism of tropomyosin and a view of protein function in Nobel Symposium 11. Symmetry and Function of Biological Systems at the Macromolecular Level (eds Engstrom, A. & Strandberg, B.) 393–414 (Almquist and Wiksell, Stockholm, John Wiley and Sons, Inc., New York, 1969).
Arvai, A. Adxv - A Program to Display X-ray Diffraction Images. (2015).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods. Enzymol. 276, 307–326 (1997).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta. Crystallogr. D. Biol. Crystallogr. 67, 235–242 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Vagin, A. A. et al. REFMAC5 dictionary: organization of prior chemical knowledge and guidelines for its use. Acta. Crystallogr. D. Biol. Crystallogr. 60, 2184–2195 (2004).
Hattne, J. et al. MicroED data collection and processing. Acta Crystallogr. A Found. Adv. 71, 353–360 (2015).
Shi, D. et al. The collection of MicroED data for macromolecular crystallography. Nat. Protoc. 11, 895–904 (2016).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993).
Miyazawa, A., Fujiyoshi, Y. & Unwin, N. Structure and gating mechanism of the acetylcholine receptor pore. Nature 423, 949–955 (2003).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013).
Banerjee, S. et al. 2.3 Å resolution cryo-EM structure of human p97 and mechanism of allosteric inhibition. Science 351, 871–875 (2016).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Ge, P. & Zhou, Z. H. Hydrogen-bonding networks and RNA bases revealed by cryo electron microscopy suggest a triggering mechanism for calcium switches. Proc. Natl Acad. Sci. USA 108, 9637–9642 (2011).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, (213–221) (2010).
Clemens, D. L., Ge, P., Lee, B. Y., Horwitz, M. A. & Zhou, Z. H. Atomic structure of T6SS reveals interlaced array essential to function. Cell 160, 940–951 (2015).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
DeLano, W. L. The PyMOL Molecular Graphics System. (DeLano Scientific, 2002).
Lawrence, M. C. & Colman, P. M. Shape complementarity at protein/protein interfaces. J. Mol. Biol. 234, 946–950 (1993).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta. Crystallogr. D. Biol. Crystallogr. 50, 760–763 (1994).
Boc, A., Diallo, A. B. & Makarenkov, V. T-REX: a web server for inferring, validating and visualizing phylogenetic trees and networks. Nucleic. Acids. Res. 40, W573–W579 (2012).
Acknowledgements
We thank Q. Cao, L. Saelices, S. Sangwan, and P. Seidler for discussion, D. Shi at Janelia for microscope support, and M. Collazo at UCLA-DOE Macromolecular Crystallization Core Technology Center for crystallization support. We thank the staff at the Argonne Photon Source (APS) NECAT Beamline 24-ID-E, which is funded by National Institutes of Health (P41 GM103403). This research was supported in part by grants from the National Science Foundation (NSF MCB 1616265 to D.S.E.) and the National Institutes of Health (GM071940 to Z.H.Z.), Howard Hughes Medical Institute and the Janelia Research Campus visitor program. D.R.B. was supported by the National Science Foundation Graduate Research Fellowship. We acknowledge the use of instruments at the Electron Imaging Center for Nanomachines supported by UCLA and by instrumentation grants from NIH (1S10RR23057 and 1U24GM116792) and NSF (DBI-1338135).
Author information
Authors and Affiliations
Contributions
E.L.G. and D.S.E. designed the project and wrote the manuscript with input from all other authors especially M.R.S. E.L.G. and H.T. conducted fibril growth experiments, stability assays and crystallization and structure determination of 248LIIKGI253. E.L.G. and M.R.S. did the fibril diffraction assays. E.L.G. grew crystals of 247DLIIKGISVHI257, while D.R.B. collected MicroED data of the sample. M.R.S., D.C. and E.L.G. processed data and solved the structure of 247DLIIKGISVHI257 crystal. E.L.G. prepared and optimized the 247DLIIKGISVHI257 fibril sample for cryo-EM while P.G. processed and solved the cryo-EM structure. E.L.G., M.R.S. and D.S.E. analyzed structures, conducted computational analysis such as area buried and designed models of fibril nucleation and elongation. T.G. and Z.H.Z. contributed to analysis of microED and cryoEM structures, respectively.
Corresponding author
Ethics declarations
Competing interests
D.S.E. is an advisor and equity shareholder in ADDRx, Inc.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 RRM2 segments demonstrate high to moderate stability against treatment with SDS and heat.
Fibril samples were first grown by shaking at 37°C at 20mM concentration in PBS, pH 7.5. Following fibril formation, samples were treated with 2% SDS and heated to 70°C for 15 min. The bar graph shows the mean raw absorbance values as well as individual data points as monitored by the Spectramax. Experiments were done in triplicate and error bars represent standard deviation.
Supplementary Figure 2 Fibril diffraction of the RRM2 fibril segments shows classic amyloid diffraction.
Fibril diffraction was collected at the 5um beam at the Advanced Photon Source. All nine samples exhibit the classic 4.6-4.8Å ring indicative of the stacking of β-strands and the 8-12Å ring indicative of mating sheets. We were unable to collect fibril diffraction patterns for 2 samples (#1: 247DLIIKG252 and #11: 254SVHISN259) Fibril diffraction experiments were completed once at APS. Peptide numbering corresponds with the code provided in Fig. 1.
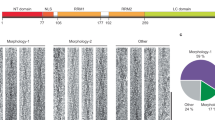
Supplementary Figure 3 Eight unique fibril morphologies are observed in the 247DLIIKGISVHI257 cryo-EM sample.
(a) Negative stain EM image of the 247DLIIKGISVHI257 cryo-EM sample. 247DLIIKGISVHI257 fibrils were grown by shaking for one week at 1mM in water. Sample was stained with 2% uranyl acetate and visualized on the FEI T12 transmission electron microscope. Fibril samples illustrate multiple morphologies, including twisted helical assemblies. (b) Illustration of the eight different types of fibrils observed by cryo-EM. The eight fibrils demonstrate four different conformations of amyloid fibrils: helices, twists, cylinders, and sheets. (c) The relative abundance of the eight unique fibrils are displayed in a table. The 3-start helix represents the most abundant species, comprising approximately 60% of the sample.
Supplementary Figure 4 Classification of images for the 247DLIIKGISVHI257 cryo-EM sample.
(a) Round 1 classification illustrates the polymorphic assemblies observed in the fibril sample. (b and c) Round 2 and 3 class averages focus on fibrils exhibiting the 3-start helix morphology. Each panel illustrates a unique face of the fibril as it twists along the EM grid. 5-7Å features of the structure become apparent during these rounds of class averaging.
Supplementary Figure 5 2D projection of the final 3D model of the 247DLIIKGISVHI257 cryo-EM fibril structure.
The panels illustrate a side view of the fibril, perpendicular to the fibril axis. Each panel represents an 8° azimuthal rotation of the fibril. The hydrogen bonding network is visualized by the stacking of β-strands.
Supplementary Figure 6 Fourier shell correlation (FSC) curve for the single-particle cryo-EM structure of RRMcore.
FSC cutoff value of 0.5 results in spatial frequency of 3.83Å for the half-maps and 3.63Å for the map-model.
Supplementary Figure 7 The asymmetric unit of the three-start helix is composed of nine strands.
(a) The electron density of the asymmetric unit is displayed. Here, the N-terminus is displayed in red and the C-terminus is displayed in blue. (b,c) The peptide strands are displayed within the electron density. The β-sheets are classified by their relative conformation; kinked = red, straight = blue, curved = green and partial = yellow. Each strand is represented by a unique color within its designated family.
Supplementary Figure 8 The protofilaments of the three-start helix fill space, exclude water, bury hydrophobic surfaces, and exhibit electrostatic interactions.
Here, glycines and apolar residues are white, polar residues are green, positively charged residues are blue, and negatively charged residues are red. The pH of this sample is ~4 and therefore histidine residues are charged. Red and blue circles indicate the presence of electrostatic interactions between the N and C termini as well as between the aspartic acid and histidine side chains. Yellow ellipses illustrate the presence of hydrophobic cores. The three fold screw axis is indicated by the triangle.
Supplementary Figure 9 Eight different symmetry classes of amyloid steric zippers.
Examples of their occurrence among RRMcore segments of TDP-43 are cataloged below their respective classes.
Supplementary Figure 10 The cryo-EM structure demonstrates key features including a 247DLIIKGISVHI257 kinked backbone that flips orientation and a pseudo-two-fold symmetry is observed in the asymmetric unit.
(a) Below, the kinking of the 247DLIIKGISVHI257 backbone allows for the carbonyl of the glycine and isoleucine to both face down. The glycine and isoleucine residues have been marked by one letter abbreviations. (b-c) Strands 1, 2, 5 and 8 form a complex which exhibits two fold symmetry with strands 3, 4, 6 and 9. The oval shape indicates two fold symmetry.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 1–3
Supplementary Video 1
Segment DLIIKGISVHI forms a three-start helix. Here, the 27 β-strands that form the helical fibril can be viewed down the fibril axis. As the fibril axis become parallel to the screen, the twist of the fibril becomes visible
Supplementary Video 2
The asymmetric unit of DLIIKGISVHI is composed of nine strands. These strands adopt kinked, bent and straight morphologies that form five distinct interfaces
Rights and permissions
About this article
Cite this article
Guenther, E.L., Ge, P., Trinh, H. et al. Atomic-level evidence for packing and positional amyloid polymorphism by segment from TDP-43 RRM2. Nat Struct Mol Biol 25, 311–319 (2018). https://doi.org/10.1038/s41594-018-0045-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0045-5
This article is cited by
-
Structural analysis of cross α-helical nanotubes provides insight into the designability of filamentous peptide nanomaterials
Nature Communications (2021)
-
MicroED for the study of protein–ligand interactions and the potential for drug discovery
Nature Reviews Chemistry (2021)
-
Atomic-level differences between brain parenchymal- and cerebrovascular-seeded Aβ fibrils
Scientific Reports (2021)
-
Tethering-induced destabilization and ATP-binding for tandem RRM domains of ALS-causing TDP-43 and hnRNPA1
Scientific Reports (2021)
-
Triad of TDP43 control in neurodegeneration: autoregulation, localization and aggregation
Nature Reviews Neuroscience (2021)