Abstract
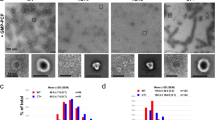
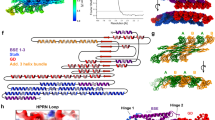
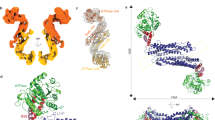
Dynamin 1-like proteins (DNM1-L) are mechanochemical GTPases that induce membrane fission in mitochondria and peroxisomes. Their mechanism depends on conformational changes driven by nucleotide and lipid cycling. Here we show the crystal structure of a mitochondrial fission dynamin (CmDnm1) from the algae Cyanidioschyzon merolae. Unlike other eukaryotic dynamin structures, CmDnm1 is in a hinge 1 closed conformation, with the GTPase domain compacted against the stalk. Within the crystal, CmDnm1 packs as a diamond-shaped tetramer that is consistent with an inactive off-membrane state. Crosslinking, photoinduced electron transfer assays, and electron microscopy verify these structures. In vitro, CmDnm1 forms concentration-dependent rings and protein–lipid tubes reminiscent of DNM1-L and classical dynamin with hinge 1 open. Our data provides a mechanism for filament collapse and membrane release that may extend to other dynamin family members. Additionally, hinge 1 closing may represent a key conformational change that contributes to membrane fission.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
08 August 2018
In the version of this article initially published, the file with supplementary figures had formatting problems. This has now been corrected.
References
Smirnova, E., Shurland, D. L., Ryazantsev, S. N. & van der Bliek, A. M. A human dynamin-related protein controls the distribution of mitochondria. J. Cell Biol. 143, 351–358 (1998).
Smirnova, E., Griparic, L., Shurland, D. L. & van der Bliek, A. M. Dynamin-related protein Drp1 is required for mitochondrial division in mammalian cells. Mol. Biol. Cell 12, 2245–2256 (2001).
Low, H. H., Sachse, C., Amos, L. A. & Löwe, J. Structure of a bacterial dynamin-like protein lipid tube provides a mechanism for assembly and membrane curving. Cell 139, 1342–1352 (2009).
Kishida, H. & Sugio, S. Crystal structure of GTPase domain fused with minimal stalks from human dynamin-1-like protein (Dlp1) in complex with several nucleotide analogues. Curr. Top. Pept. Protein Res 14, 67–77 (2013).
Wenger, J. et al. Functional mapping of human dynamin-1-like GTPase domain based on x-ray structure analyses. PLoS One 8, e71835 (2013).
Chappie, J. S. et al. A pseudoatomic model of the dynamin polymer identifies a hydrolysis-dependent powerstroke. Cell 147, 209–222 (2011).
Yan, L. et al. Structural basis for mechanochemical role of Arabidopsis thaliana dynamin-related protein in membrane fission. J. Mol. Cell Biol. 3, 378–381 (2011).
Rennie, M. L., McKelvie, S. A., Bulloch, E. M. M. & Kingston, R. L. Transient dimerization of human MxA promotes GTP hydrolysis, resulting in a mechanical power stroke. Structure 22, 1433–1445 (2014).
Chen, Y. et al. Conformational dynamics of dynamin-like MxA revealed by single-molecule FRET. Nat. Commun. 8, 15744 (2017).
Fröhlich, C. et al. Structural insights into oligomerization and mitochondrial remodelling of dynamin 1-like protein. EMBO J. 32, 1280–1292 (2013).
DeVay, R. M. et al. Coassembly of Mgm1 isoforms requires cardiolipin and mediates mitochondrial inner membrane fusion. J. Cell Biol. 186, 793–803 (2009).
Cao, Y. L. et al. MFN1 structures reveal nucleotide-triggered dimerization critical for mitochondrial fusion. Nature 542, 372–376 (2017).
Gao, S. et al. Structure of myxovirus resistance protein a reveals intra- and intermolecular domain interactions required for the antiviral function. Immunity 35, 514–525 (2011).
Gao, S. et al. Structural basis of oligomerization in the stalk region of dynamin-like MxA. Nature 465, 502–506 (2010).
Faelber, K. et al. Crystal structure of nucleotide-free dynamin. Nature 477, 556–560 (2011).
Reubold, T. F. et al. Crystal structure of the dynamin tetramer. Nature 525, 404–408 (2015).
Mears, J. A. et al. Conformational changes in Dnm1 support a contractile mechanism for mitochondrial fission. Nat. Struct. Mol. Biol. 18, 20–26 (2011).
Chappie, J. S., Acharya, S., Leonard, M., Schmid, S. L. & Dyda, F. G domain dimerization controls dynamin’s assembly-stimulated GTPase activity. Nature 465, 435–440 (2010).
Alvarez, F. J. D. et al. CryoEM structure of MxB reveals a novel oligomerization interface critical for HIV restriction. Sci. Adv. 3, e1701264 (2017).
Antonny, B. et al. Membrane fission by dynamin: what we know and what we need to know. EMBO J. 35, 2270–2284 (2016).
Nishida, K. et al. Dynamic recruitment of dynamin for final mitochondrial severance in a primitive red alga. Proc. Natl. Acad. Sci. USA 100, 2146–2151 (2003).
Imoto, Y. et al. Single-membrane-bounded peroxisome division revealed by isolation of dynamin-based machinery. Proc. Natl. Acad. Sci. USA 110, 9583–9588 (2013).
Koch, A. et al. Dynamin-like protein 1 is involved in peroxisomal fission. J. Biol. Chem. 278, 8597–8605 (2003).
Narayanan, R., Leonard, M., Song, B. D., Schmid, S. L. & Ramaswami, M. An internal GAP domain negatively regulates presynaptic dynamin in vivo: a two-step model for dynamin function. J. Cell Biol. 169, 117–126 (2005).
Ingerman, E. et al. Dnm1 forms spirals that are structurally tailored to fit mitochondria. J. Cell Biol. 170, 1021–1027 (2005).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Ford, M. G. J., Jenni, S. & Nunnari, J. The crystal structure of dynamin. Nature 477, 561–566 (2011).
Low, H. H. & Löwe, J. A bacterial dynamin-like protein. Nature 444, 766–769 (2006).
Mattila, J. P. et al. A hemi-fission intermediate links two mechanistically distinct stages of membrane fission. Nature 524, 109–113 (2015).
Doose, S., Neuweiler, H. & Sauer, M. Fluorescence quenching by photoinduced electron transfer: a reporter for conformational dynamics of macromolecules. ChemPhysChem 10, 1389–1398 (2009).
Mansoor, S. E., Dewitt, M. A. & Farrens, D. L. Distance mapping in proteins using fluorescence spectroscopy: the tryptophan-induced quenching (TrIQ) method. Biochemistry 49, 9722–9731 (2010).
De Vecchis, D. et al. A membrane-inserted structural model of the yeast mitofusin Fzo1. Sci. Rep. 7, 10217 (2017).
Binns, D. D. et al. Correlation between self-association modes and GTPase activation of dynamin. J. Protein Chem. 18, 277–290 (1999).
Muhlberg, A. B., Warnock, D. E. & Schmid, S. L. Domain structure and intramolecular regulation of dynamin GTPase. EMBO J. 16, 6676–6683 (1997).
Okamoto, P. M., Tripet, B., Litowski, J., Hodges, R. S. & Vallee, R. B. Multiple distinct coiled-coils are involved in dynamin self-assembly. J. Biol. Chem. 274, 10277–10286 (1999).
Tuma, P. L. & Collins, C. A. Activation of dynamin GTPase is a result of positive cooperativity. J. Biol. Chem. 269, 30842–30847 (1994).
Chen, Y. J., Zhang, P., Egelman, E. H. & Hinshaw, J. E. The stalk region of dynamin drives the constriction of dynamin tubes. Nat. Struct. Mol. Biol. 11, 574–575 (2004).
Zhang, P. & Hinshaw, J. E. Three-dimensional reconstruction of dynamin in the constricted state. Nat. Cell Biol. 3, 922–926 (2001).
Strack, S. & Cribbs, J. T. Allosteric modulation of Drp1 mechanoenzyme assembly and mitochondrial fission by the variable domain. J. Biol. Chem. 287, 10990–11001 (2012).
Zhang, Y., Gao, X. & Garavito, R. M. Biochemical characterization of human dynamin-like protein 1. J. Biochem 150, 627–633 (2011).
Mozdy, A. D., McCaffery, J. M. & Shaw, J. M. Dnm1p GTPase-mediated mitochondrial fission is a multi-step process requiring the novel integral membrane component Fis1p. J. Cell Biol. 151, 367–380 (2000).
Gandre-Babbe, S. & van der Bliek, A. M. The novel tail-anchored membrane protein Mff controls mitochondrial and peroxisomal fission in mammalian cells. Mol. Biol. Cell 19, 2402–2412 (2008).
Palmer, C. S. et al. MiD49 and MiD51, new components of the mitochondrial fission machinery. EMBO Rep. 12, 565–573 (2011).
Zhao, J. et al. Human MIEF1 recruits Drp1 to mitochondrial outer membranes and promotes mitochondrial fusion rather than fission. EMBO J. 30, 2762–2778 (2011).
Morlot, S. et al. Membrane shape at the edge of the dynamin helix sets location and duration of the fission reaction. Cell 151, 619–629 (2012).
Roux, A., Uyhazi, K., Frost, A. & De Camilli, P. GTP-dependent twisting of dynamin implicates constriction and tension in membrane fission. Nature 441, 528–531 (2006).
Danino, D., Moon, K. H. & Hinshaw, J. E. Rapid constriction of lipid bilayers by the mechanochemical enzyme dynamin. J. Struct. Biol. 147, 259–267 (2004).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Low, H. H., Moncrieffe, M. C. & Löwe, J. The crystal structure of ZapA and its modulation of FtsZ polymerisation. J. Mol. Biol. 341, 839–852 (2004).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Biasini, M. et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 42, W252–8 (2014).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Headd, J. J. et al. Use of knowledge-based restraints in phenix.refine to improve macromolecular refinement at low resolution. Acta Crystallogr. D. Biol. Crystallogr. 68, 381–390 (2012).
Laskowski, R. A., Macarthur, M. W., Moss, D. S., Thornton, J. M. Procheck - a program to check the stereochemical quality of protein structures.J. Appl. Crystallogr. 26, 283–291 (1993).
Loo, T. W. & Clarke, D. M. Determining the dimensions of the drug-binding domain of human P-glycoprotein using thiol cross-linking compounds as molecular rulers. J. Biol. Chem. 276, 36877–36880 (2001).
Acknowledgements
We thank the Beamline staff at the ESRF for data collection support and T. Pape for in-house EM facility support. We acknowledge the gift of C. merolae genomic DNA from T. Kuroiwa. We are grateful to A. Chernyatina and J. Liu for manuscript feedback. This work was supported by a Wellcome Trust Fellowship (097328/Z/11/Z) to H.H.L. O.B. was supported by a BBSRC Doctoral Training Partnership with Imperial College.
Author information
Authors and Affiliations
Contributions
O.B. and H.H.L. designed experiments. H.H.L. initially purified native protein, obtained crystals, and determined structure. O.B. purified proteins and performed all other experiments including GTPase, crosslinking, and PET assays and generated EM data. H.H.L. and O.B. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Bohuszewicz, O., Low, H.H. Structure of a mitochondrial fission dynamin in the closed conformation. Nat Struct Mol Biol 25, 722–731 (2018). https://doi.org/10.1038/s41594-018-0097-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0097-6
This article is cited by
-
Structural basis for regulated assembly of the mitochondrial fission GTPase Drp1
Nature Communications (2024)
-
SynDLP is a dynamin-like protein of Synechocystis sp. PCC 6803 with eukaryotic features
Nature Communications (2023)
-
Measuring Membrane Penetration Depths and Conformational Changes in Membrane Peptides and Proteins
The Journal of Membrane Biology (2022)
-
Function and regulation of the divisome for mitochondrial fission
Nature (2021)
-
The cell biology of mitochondrial membrane dynamics
Nature Reviews Molecular Cell Biology (2020)