Featured
-
-
Article |
 Bioaccumulation of therapeutic drugs by human gut bacteria
Bioaccumulation of therapeutic drugs by human gut bacteria
An analysis of the interactions between 15 drugs and 25 gut bacterial strains shows that bioaccumulation of drugs within bacterial cells is another mechanism through which gut microorganisms can alter drug availability and efficacy.
- Martina Klünemann
- , Sergej Andrejev
- & Kiran R. Patil
-
Article |
 Extensive signal integration by the phytohormone protein network
Extensive signal integration by the phytohormone protein network
A systems-level map of the Arabidopsis hormone signalling network, comprising more than 2,000 binary protein–protein interactions, reveals hundreds of interpathway contact points, many of which mediate crosstalk between different hormone pathways.
- Melina Altmann
- , Stefan Altmann
- & Pascal Falter-Braun
-
Article |
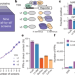 A reference map of the human binary protein interactome
A reference map of the human binary protein interactome
A human binary protein interactome map that includes around 53,000 protein–protein interactions involving more than 8,000 proteins provides a reference for the study of human cellular function in health and disease.
- Katja Luck
- , Dae-Kyum Kim
- & Michael A. Calderwood
-
Letter |
 An orthogonal proteomic survey uncovers novel Zika virus host factors
An orthogonal proteomic survey uncovers novel Zika virus host factors
Integrative analyses identify host proteins that are modulated by Zika virus at multiple levels and provide a comprehensive framework for the understanding of Zika virus-induced changes to cellular pathways.
- Pietro Scaturro
- , Alexey Stukalov
- & Andreas Pichlmair
-
Letter |
 Interactome map uncovers phosphatidylserine transport by oxysterol-binding proteins
Interactome map uncovers phosphatidylserine transport by oxysterol-binding proteins
The lipid-binding profiles of all lipid-transfer proteins in Saccharomyces cerevisiae are determined and a new subfamily of oxysterol-binding proteins that function in phosphatidylserine homeostasis and transport is identified.
- Kenji Maeda
- , Kanchan Anand
- & Anne-Claude Gavin
-
Letter |
 A latent capacity for evolutionary innovation through exaptation in metabolic systems
A latent capacity for evolutionary innovation through exaptation in metabolic systems
A computational analysis of the ability of a metabolic reaction network to synthesize all biomass from a single source of carbon and energy shows that when such networks are required to be viable on one particular carbon source, they are typically also viable on multiple other carbon sources that were not targets of selection.
- Aditya Barve
- & Andreas Wagner
-

 Defining mitochondrial protein functions through deep multiomic profiling
Defining mitochondrial protein functions through deep multiomic profiling Cell mechanics and the cytoskeleton
Cell mechanics and the cytoskeleton