Article
|
Open Access
Featured
-
-
Article
| Open Access Single-cell analysis of chromatin accessibility in the adult mouse brain
Single-cell analysis of chromatin accessibility in the adult mouse brain
An atlas of candidate cis-regulatory DNA elements (cCREs) in the adult mouse brain unravels the transcriptional regulatory programs that drive the heterogeneity and complexity of brain structure and function.
- Songpeng Zu
- , Yang Eric Li
- & Bing Ren
-
Article
| Open Access Brain-wide correspondence of neuronal epigenomics and distant projections
Brain-wide correspondence of neuronal epigenomics and distant projections
This study uses epi-retro-seq to link single-cell epigenomes and cell types to long-distance projections for neurons dissected from different regions projecting to different targets across the whole mouse brain.
- Jingtian Zhou
- , Zhuzhu Zhang
- & Edward M. Callaway
-
Article
| Open Access MSL2 ensures biallelic gene expression in mammals
MSL2 ensures biallelic gene expression in mammals
After loss of MSL2, a class of dosage-sensitive genes transitions from biallelic to monoallelic expression, whereby one allele remains active, retaining active histone modifications and transcription factor binding, and the other allele is silenced, exhibiting loss of promoter–enhancer contacts and the acquisition of DNA methylation.
- Yidan Sun
- , Meike Wiese
- & Asifa Akhtar
-
Article
| Open Access Transient naive reprogramming corrects hiPS cells functionally and epigenetically
Transient naive reprogramming corrects hiPS cells functionally and epigenetically
A new reprogramming strategy used to produce human induced pluripotent stem cells from somatic cells results in epigenetic and functional profiles that are highly similar to those of human embryonic stem cells.
- Sam Buckberry
- , Xiaodong Liu
- & Ryan Lister
-
Article |
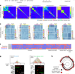 Cycles of satellite and transposon evolution in Arabidopsis centromeres
Cycles of satellite and transposon evolution in Arabidopsis centromeres
Inter- and intra-species comparison of Arabidopsis centromere variation identifies rapid cycles of transposon invasion and purging through satellite homogenization that drive centromere evolution.
- Piotr Wlodzimierz
- , Fernando A. Rabanal
- & Ian R. Henderson
-
Article
| Open Access Genomic–transcriptomic evolution in lung cancer and metastasis
Genomic–transcriptomic evolution in lung cancer and metastasis
Computational and machine-learning approaches that integrate genomic and transcriptomic variation from paired primary and metastatic non-small cell lung cancer samples from the TRACERx cohort reveal the role of transcriptional events in tumour evolution.
- Carlos Martínez-Ruiz
- , James R. M. Black
- & Nicholas McGranahan
-
Article
| Open Access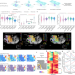 Spatial epigenome–transcriptome co-profiling of mammalian tissues
Spatial epigenome–transcriptome co-profiling of mammalian tissues
The authors present two technologies for spatially resolved, genome-wide, joint profiling of the epigenome and transcriptome by cosequencing chromatin accessibility and gene expression, or histone modifications and gene expression on the same tissue section at near-single-cell resolution.
- Di Zhang
- , Yanxiang Deng
- & Rong Fan
-
Article |
 BRD8 maintains glioblastoma by epigenetic reprogramming of the p53 network
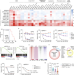
BRD8 maintains glioblastoma by epigenetic reprogramming of the p53 network
BRD8 is identified as a specific epigenetic vulnerability for glioblastomas that harbour wild-type p53.
- Xueqin Sun
- , Olaf Klingbeil
- & Alea A. Mills
-
Article
| Open Access The co-evolution of the genome and epigenome in colorectal cancer
The co-evolution of the genome and epigenome in colorectal cancer
A study maps genetic and epigenetic heterogeneity of primary colorectal adenomas and cancers at single-clone resolution through spatial multi-omic profiling of individual glands and adjacent normal tissue.
- Timon Heide
- , Jacob Househam
- & Andrea Sottoriva
-
Article
| Open Access Spatial profiling of chromatin accessibility in mouse and human tissues
Spatial profiling of chromatin accessibility in mouse and human tissues
Spatial-ATAC-seq—spatially resolved chromatin accessibility profiling of tissue sections using next-generation sequencing—delineated tissue-region-specific epigenetic landscapes in mouse embryos and identified gene regulators involved in the development of the central nervous system and the lymphoid tissue.
- Yanxiang Deng
- , Marek Bartosovic
- & Rong Fan
-
Article
| Open Access Cohesin-mediated loop anchors confine the locations of human replication origins
Cohesin-mediated loop anchors confine the locations of human replication origins
A study shows that the three-dimensional conformation of the human genome influences the positioning of DNA replication initiation zones, highlighting cohesin-mediated loop anchors as essential determinants of their precise location.
- Daniel J. Emerson
- , Peiyao A. Zhao
- & Jennifer E. Phillips-Cremins
-
Article |
 Brahma safeguards canalization of cardiac mesoderm differentiation
Brahma safeguards canalization of cardiac mesoderm differentiation
The BAF chromatin-remodelling complex ATPase gene Brm safeguards cell identity during directed cardiogenesis of mouse embryonic stem cells.
- Swetansu K. Hota
- , Kavitha S. Rao
- & Benoit G. Bruneau
-
Article
| Open Access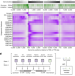 Mutation bias reflects natural selection in Arabidopsis thaliana
Mutation bias reflects natural selection in Arabidopsis thaliana
Data on de novo mutations in Arabidopsis thaliana reveal that mutations do not occur randomly; instead, epigenome-associated mutation bias reduces the occurrence of deleterious mutations.
- J. Grey Monroe
- , Thanvi Srikant
- & Detlef Weigel
-
Article
| Open Access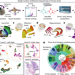 DNA methylation atlas of the mouse brain at single-cell resolution
DNA methylation atlas of the mouse brain at single-cell resolution
A comprehensive survey of the epigenome from 45 regions of the mouse cortex, hippocampus, striatum, pallidum and olfactory areas using single-nucleus DNA methylation sequencing enables identification of 161 cell clusters with distinct locations and projection targets and provides insights into the regulatory landscape underlying neuronal diversity and spatial regulation.
- Hanqing Liu
- , Jingtian Zhou
- & Joseph R. Ecker
-
Article
| Open Access A transcriptomic and epigenomic cell atlas of the mouse primary motor cortex
A transcriptomic and epigenomic cell atlas of the mouse primary motor cortex
The authors describe an integrated atlas of the diverse cell types in the mouse primary motor cortex.
- Zizhen Yao
- , Hanqing Liu
- & Eran A. Mukamel
-
Article
| Open Access Epigenomic diversity of cortical projection neurons in the mouse brain
Epigenomic diversity of cortical projection neurons in the mouse brain
Quantitative analysis of the methylation of mouse cortical neurons that project to different cortical and subcortical target regions provides insight into genetic mechanisms that contribute to differences in cell function.
- Zhuzhu Zhang
- , Jingtian Zhou
- & Edward M. Callaway
-
Article
| Open Access An atlas of gene regulatory elements in adult mouse cerebrum
An atlas of gene regulatory elements in adult mouse cerebrum
A comprehensive analysis of gene regulatory elements in 160 distinct cell types from the mouse cerebrum.
- Yang Eric Li
- , Sebastian Preissl
- & Bing Ren
-
Article |
 Interpreting type 1 diabetes risk with genetics and single-cell epigenomics
Interpreting type 1 diabetes risk with genetics and single-cell epigenomics
A genome-wide association study combined with single-cell epigenomics identifies risk loci for type 1 diabetes.
- Joshua Chiou
- , Ryan J. Geusz
- & Kyle J. Gaulton
-
Article |
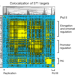 A high-resolution protein architecture of the budding yeast genome
A high-resolution protein architecture of the budding yeast genome
A ChIP–exo method is used to define the genome-wide positional organization of proteins associated with gene transcription, DNA replication, centromeres, subtelomeres and transposons, revealing distinct protein assemblies for constitutive and inducible gene expression.
- Matthew J. Rossi
- , Prashant K. Kuntala
- & B. Franklin Pugh
-
Article
| Open Access Regulatory genomic circuitry of human disease loci by integrative epigenomics
Regulatory genomic circuitry of human disease loci by integrative epigenomics
The authors present EpiMap, a compendium that comprises 10,000 epigenomic maps across more than 800 biosamples for the annotation of genome-wide association study circuitry.
- Carles A. Boix
- , Benjamin T. James
- & Manolis Kellis
-
Article |
 A map of cis-regulatory elements and 3D genome structures in zebrafish
A map of cis-regulatory elements and 3D genome structures in zebrafish
A comprehensive map of transcriptomes, cis-regulatory elements, heterochromatin structure, the methylome and 3D genome organization in the zebrafish (Danio rerio) enables identification of species-specific and evolutionarily conserved regulatory features, and provides a foundation for modelling studies on human disease and development.
- Hongbo Yang
- , Yu Luan
- & Feng Yue
-
Article |
 Epigenetic therapy induces transcription of inverted SINEs and ADAR1 dependency
Epigenetic therapy induces transcription of inverted SINEs and ADAR1 dependency
Inverted-repeat Alu elements are the main source of drug-induced immunogenic double-stranded RNAs, which are destabilized by the RNA deaminase ADAR1, thereby limiting activation of the immune response.
- Parinaz Mehdipour
- , Sajid A. Marhon
- & Daniel D. De Carvalho
-
Article |
 Cell-type-specific 3D epigenomes in the developing human cortex
Cell-type-specific 3D epigenomes in the developing human cortex
Analysis of cis-regulatory chromatin interactions, open chromatin and transcriptomes for different cell types isolated from mid-gestational human cortex samples provides insights into gene regulation during development.
- Michael Song
- , Mark-Phillip Pebworth
- & Yin Shen
-
Article
| Open Access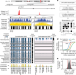 Global reference mapping of human transcription factor footprints
Global reference mapping of human transcription factor footprints
A high-density DNase I cleavage map from 243 human cell and tissue types provides a genome-wide, nucleotide-resolution map of human transcription factor footprints.
- Jeff Vierstra
- , John Lazar
- & John A. Stamatoyannopoulos
-
Article
| Open Access Index and biological spectrum of human DNase I hypersensitive sites
Index and biological spectrum of human DNase I hypersensitive sites
High-resolution maps of DNase I hypersensitive sites from 733 human biosamples are used to identify and index regulatory DNA within the human genome.
- Wouter Meuleman
- , Alexander Muratov
- & John Stamatoyannopoulos
-
Perspective |
 Perspectives on ENCODE
Perspectives on ENCODE
The authors summarize the history of the ENCODE Project, the achievements of ENCODE 1 and ENCODE 2, and how the new data generated and analysed in ENCODE 3 complement the previous phases.
- Federico Abascal
- , Reyes Acosta
- & Richard M. Myers
-
Article
| Open Access An atlas of dynamic chromatin landscapes in mouse fetal development
An atlas of dynamic chromatin landscapes in mouse fetal development
Analysis of chromatin state and accessibility in mouse tissues from twelve sites and eight developmental stages provides a comprehensive view of chromatin dynamics.
- David U. Gorkin
- , Iros Barozzi
- & Bing Ren
-
Article
| Open Access Expanded encyclopaedias of DNA elements in the human and mouse genomes
Expanded encyclopaedias of DNA elements in the human and mouse genomes
The authors summarize the data produced by phase III of the Encyclopedia of DNA Elements (ENCODE) project, a resource for better understanding of the human and mouse genomes.
- Federico Abascal
- , Reyes Acosta
- & Zhiping Weng
-
Article
| Open Access Spatiotemporal DNA methylome dynamics of the developing mouse fetus
Spatiotemporal DNA methylome dynamics of the developing mouse fetus
Analysis of 168 methylomes from 12 mouse tissues at 9 developmental stages sheds light on the epigenetic and regulatory landscape during mammalian fetal development.
- Yupeng He
- , Manoj Hariharan
- & Joseph R. Ecker
-
Article
| Open Access Occupancy maps of 208 chromatin-associated proteins in one human cell type
Occupancy maps of 208 chromatin-associated proteins in one human cell type
ChIP–seq and CETCh–seq data are used to analyse binding maps for 208 transcription factors and other chromatin-associated proteins in a single human cell type, providing a comprehensive catalogue of the transcription factor landscape and gene regulatory networks in these cells.
- E. Christopher Partridge
- , Surya B. Chhetri
- & Eric M. Mendenhall
-
Article |
 Epigenetic regulator function through mouse gastrulation
Epigenetic regulator function through mouse gastrulation
An experimental and analytical pipeline is used to assess, at the single-cell level, complex transcriptional and morphological mutant phenotypes that occur in mouse embryos during gastrulation.
- Stefanie Grosswendt
- , Helene Kretzmer
- & Alexander Meissner
-
Article |
 Structural cells are key regulators of organ-specific immune responses
Structural cells are key regulators of organ-specific immune responses
Structural cells implement a broad range of immune-regulatory functions beyond their roles as barriers and connective tissues, and they utilize an epigenetically encoded potential for immune gene activation in their rapid response to viral infection.
- Thomas Krausgruber
- , Nikolaus Fortelny
- & Christoph Bock
-
Article |
 Multi-omics profiling of mouse gastrulation at single-cell resolution
Multi-omics profiling of mouse gastrulation at single-cell resolution
Single-cell mapping of chromatin accessibility, DNA methylation and RNA expression during gastrulation in mouse embryos shows characteristic epigenetic changes that accompany formation of the primary germ layers.
- Ricard Argelaguet
- , Stephen J. Clark
- & Wolf Reik
-
Article |
 Key role for CTCF in establishing chromatin structure in human embryos
Key role for CTCF in establishing chromatin structure in human embryos
The chromatin regulator CTCF has key roles in the gradual development of hierarchical chromatin structure during human embryogenesis.
- Xuepeng Chen
- , Yuwen Ke
- & Zi-Jiang Chen
-
Letter |
 Epigenetic evolution and lineage histories of chronic lymphocytic leukaemia
Epigenetic evolution and lineage histories of chronic lymphocytic leukaemia
A single-cell approach is used to follow the heritable stochastic changes to DNA methylation that occur in primary chronic lymphocytic leukaemia and healthy B cells, allowing the tracing of cell lineage histories and evolution during treatment with ibrutinib.
- Federico Gaiti
- , Ronan Chaligne
- & Dan A. Landau
-
Letter |
 Multiplex chromatin interactions with single-molecule precision
Multiplex chromatin interactions with single-molecule precision
A strategy using droplet-based and barcode-linked sequencing captures multiplex chromatin interactions at single-molecule precision, and here provides topological insight into chromatin structures and transcription in Drosophila.
- Meizhen Zheng
- , Simon Zhongyuan Tian
- & Yijun Ruan
-
Article
| Open Access Amphioxus functional genomics and the origins of vertebrate gene regulation
Amphioxus functional genomics and the origins of vertebrate gene regulation
Genomic, epigenomic and transcriptomic data derived from the Mediterranean amphioxus (Branchiostoma lanceolatum) provide insights into the evolution of the genomic regulatory landscape of chordates.
- Ferdinand Marlétaz
- , Panos N. Firbas
- & Manuel Irimia
-
Article |
 Mechanoresponsive stem cells acquire neural crest fate in jaw regeneration
Mechanoresponsive stem cells acquire neural crest fate in jaw regeneration
Reversion of adult skeletal stem cells to a developmental state underlies the growth of new bone during jaw regeneration, in a process that relies on mechanotransduction via the focal adhesion kinase protein.
- Ryan C. Ransom
- , Ava C. Carter
- & Michael T. Longaker
-
Letter |
 Principles of nucleosome organization revealed by single-cell micrococcal nuclease sequencing
Principles of nucleosome organization revealed by single-cell micrococcal nuclease sequencing
Single-cell micrococcal nuclease sequencing simultaneously measures chromatin accessibility and genome-wide nucleosome positioning in single cells to reveal principles of nucleosome organization.
- Binbin Lai
- , Weiwu Gao
- & Keji Zhao
-
Letter |
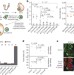 Epigenetic reprogramming enables the transition from primordial germ cell to gonocyte
Epigenetic reprogramming enables the transition from primordial germ cell to gonocyte
Gonadal germline epigenetic reprogramming involves an interplay between DNA methylation, the polycomb complex and Tet1 in both DNA methylation dependent and independent roles, to ensure the activation of a specific subset of genes critical for progression of gametogenesis.
- Peter W. S. Hill
- , Harry G. Leitch
- & Petra Hajkova
-
Perspective |
 The 4D nucleome project
The 4D nucleome project
The 4D Nucleome Network aims to map the spatial and dynamic organization of the human and mouse genomes to gain insight into the structure and biological functions of the nucleus.
- Job Dekker
- , Andrew S. Belmont
- & Sheng Zhong
-
Letter |
 Allelic reprogramming of 3D chromatin architecture during early mammalian development
Allelic reprogramming of 3D chromatin architecture during early mammalian development
A low-input Hi-C method is used to show that chromatin organization is markedly relaxed in pre-implantation mouse embryos after fertilization and that the subsequent maturation of 3D chromatin architecture is surprisingly slow.
- Zhenhai Du
- , Hui Zheng
- & Wei Xie
-
Letter |
 KRAB zinc-finger proteins contribute to the evolution of gene regulatory networks
KRAB zinc-finger proteins contribute to the evolution of gene regulatory networks
Genomic analyses of KRAB-containing zinc-finger proteins and the transposable elements to which they bind show that a co-evolutionary arms race was not the only driver of their evolution.
- Michaël Imbeault
- , Pierre-Yves Helleboid
- & Didier Trono
-
Article |
 Complex multi-enhancer contacts captured by genome architecture mapping
Complex multi-enhancer contacts captured by genome architecture mapping
A technique called genome architecture mapping (GAM) involves sequencing DNA from a large number of thin nuclear cryosections to develop a map of genome organization without the limitations of existing 3C-based methods.
- Robert A. Beagrie
- , Antonio Scialdone
- & Ana Pombo
-
Letter |
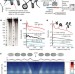 Variable chromatin structure revealed by in situ spatially correlated DNA cleavage mapping
Variable chromatin structure revealed by in situ spatially correlated DNA cleavage mapping
The first genome-wide map of human chromatin conformation at the 1–3 nucleosome (50–500 base pair) scale, obtained using ionizing radiation-induced spatially correlated cleavage of DNA with sequencing (RICC-seq), which identifies spatially proximal DNA–DNA contacts.
- Viviana I. Risca
- , Sarah K. Denny
- & William J. Greenleaf
-
Letter |
 Epigenome-wide association study of body mass index, and the adverse outcomes of adiposity
Epigenome-wide association study of body mass index, and the adverse outcomes of adiposity
A large-scale epigenome-wide association study identifies changes in DNA methylation associated with body mass index in blood and adipose tissue, and correlates DNA methylation sites with high risk of incident type 2 diabetes.
- Simone Wahl
- , Alexander Drong
- & John C. Chambers
-
Letter |
 Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
Allelic reprogramming of the histone modification H3K4me3 in early mammalian development
Three papers in this issue of Nature use highly sensitive ChIP–seq assays to describe the dynamic patterns of histone modifications during early mouse embryogenesis, showing that oocytes have a distinctive epigenome and providing insights into how the maternal gene expression program transitions to the zygotic program.
- Bingjie Zhang
- , Hui Zheng
- & Wei Xie
-
Article |
 The landscape of accessible chromatin in mammalian preimplantation embryos
The landscape of accessible chromatin in mammalian preimplantation embryos
An improved ATAC-seq approach is used to describe a genome-wide view of accessible chromatin and cis-regulatory elements in mouse preimplantation embryos, allowing construction of a regulatory network of early development that helps to identify key modulators of lineage specification.
- Jingyi Wu
- , Bo Huang
- & Wei Xie
-
Article |
 DNA methylation on N6-adenine in mammalian embryonic stem cells
DNA methylation on N6-adenine in mammalian embryonic stem cells
The prevalence of N6-adenine DNA methylation in mammals was previously unknown; this study reveals that N6-methyladenine can be found in mouse embryonic stem cells, especially at subfamilies of young (<1.5 million years old) LINE-1 transposons.
- Tao P. Wu
- , Tao Wang
- & Andrew Z. Xiao

 Conserved and divergent gene regulatory programs of the mammalian neocortex
Conserved and divergent gene regulatory programs of the mammalian neocortex