Article
|
Open Access
Featured
-
-
Article
| Open Access Functional and evolutionary significance of unknown genes from uncultivated taxa
Functional and evolutionary significance of unknown genes from uncultivated taxa
We analysed 149,842 environmental genomes from multiple habitats and compiled a curated catalogue of 404,085 functionally and evolutionarily significant novel gene families exclusive to uncultivated prokaryotic taxa spanning multiple species.
- Álvaro Rodríguez del Río
- , Joaquín Giner-Lamia
- & Jaime Huerta-Cepas
-
Perspective |
 Hold out the genome: a roadmap to solving the cis-regulatory code
Hold out the genome: a roadmap to solving the cis-regulatory code
A roadmap towards solving the cis-regulatory code using a combination of machine learning and massively parallel assays of exogenous DNA is proposed.
- Carl G. de Boer
- & Jussi Taipale
-
Article |
 The complete sequence of a human Y chromosome
The complete sequence of a human Y chromosome
We present the complete 62,460,029-base-pair sequence of a human Y chromosome from the HG002 genome (T2T-Y) that corrects multiple errors in GRCh38-Y and adds over 30 million base pairs of sequence to the reference.
- Arang Rhie
- , Sergey Nurk
- & Adam M. Phillippy
-
Article |
 Assembly of 43 human Y chromosomes reveals extensive complexity and variation
Assembly of 43 human Y chromosomes reveals extensive complexity and variation
De novo assemblies of 43 Y chromosomes spanning 182,900 years of human evolution reveal considerable diversity in the size and structure of the human Y chromosome.
- Pille Hallast
- , Peter Ebert
- & Charles Lee
-
Article |
 Cancer aneuploidies are shaped primarily by effects on tumour fitness
Cancer aneuploidies are shaped primarily by effects on tumour fitness
A study reports the development of an algorithm, BISCUT, that detects genomic loci under selective pressure by relying on the distribution of breakpoints across chromosome arms, and uses it to explore how aneuploidies affect tumorigenesis.
- Juliann Shih
- , Shahab Sarmashghi
- & Rameen Beroukhim
-
Article
| Open Access A pangenome reference of 36 Chinese populations
A pangenome reference of 36 Chinese populations
A study reports data from the first phase of the Chinese Pangenome Consortium including 116 de novo assemblies from 58 core samples representing 36 minority Chinese ethnic groups.
- Yang Gao
- , Xiaofei Yang
- & Shuhua Xu
-
Article
| Open Access A draft human pangenome reference
A draft human pangenome reference
An initial draft of the human pangenome is presented and made publicly available by the Human Pangenome Reference Consortium; the draft contains 94 de novo haplotype assemblies from 47 ancestrally diverse individuals.
- Wen-Wei Liao
- , Mobin Asri
- & Benedict Paten
-
Article
| Open Access The person-to-person transmission landscape of the gut and oral microbiomes
The person-to-person transmission landscape of the gut and oral microbiomes
Data from more than 9,700 human stool and oral metagenomes has been used to decipher the strain transmission patterns of the human microbiome from mother to infant, within households and within populations.
- Mireia Valles-Colomer
- , Aitor Blanco-Míguez
- & Nicola Segata
-
Article
| Open Access Wastewater sequencing reveals early cryptic SARS-CoV-2 variant transmission
Wastewater sequencing reveals early cryptic SARS-CoV-2 variant transmission
Emerging SARS-CoV-2 variants of concern were detected early and multiple cases of virus spread not captured by clinical genomic surveillance were identified using high-resolution wastewater and clinical sequencing.
- Smruthi Karthikeyan
- , Joshua I. Levy
- & Rob Knight
-
Article
| Open Access Signatures of copy number alterations in human cancer
Signatures of copy number alterations in human cancer
A new framework enables a pan-cancer reference set of copy number signatures derived from allele-specific profiles from different experimental assays.
- Christopher D. Steele
- , Ammal Abbasi
- & Nischalan Pillay
-
Perspective |
 The Human Pangenome Project: a global resource to map genomic diversity
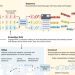
The Human Pangenome Project: a global resource to map genomic diversity
The Human Pangenome Reference Consortium aims to offer the highest quality and most complete human pangenome reference that provides diverse genomic representation across human populations.
- Ting Wang
- , Lucinda Antonacci-Fulton
- & David Haussler
-
Analysis
| Open Access A joint NCBI and EMBL-EBI transcript set for clinical genomics and research
A joint NCBI and EMBL-EBI transcript set for clinical genomics and research
Matched Annotation from NCBI and EMBL-EBI (MANE) delivers joint transcript sets from Ensembl/GENCODE and RefSeq for standardizing variant reporting in clinical genomics and research.
- Joannella Morales
- , Shashikant Pujar
- & Terence D. Murphy
-
Article
| Open Access Isoform cell-type specificity in the mouse primary motor cortex
Isoform cell-type specificity in the mouse primary motor cortex
- A. Sina Booeshaghi
- , Zizhen Yao
- & Lior Pachter
-
Article |
 Clonal dynamics in early human embryogenesis inferred from somatic mutation
Clonal dynamics in early human embryogenesis inferred from somatic mutation
Adult human tissues from diverse sites around the body are used to reconstruct cellular phylogenies from early development, using somatic mutations as an internal barcode.
- Seongyeol Park
- , Nanda Maya Mali
- & Young Seok Ju
-
Article
| Open Access A high-quality bonobo genome refines the analysis of hominid evolution
A high-quality bonobo genome refines the analysis of hominid evolution
A high-quality bonobo genome assembly provides insights into incomplete lineage sorting in hominids and its relevance to gene evolution and the genetic relationship among living hominids.
- Yafei Mao
- , Claudia R. Catacchio
- & Evan E. Eichler
-
Article
| Open Access Evolutionary and biomedical insights from a marmoset diploid genome assembly
Evolutionary and biomedical insights from a marmoset diploid genome assembly
A trio-binning approach is used to produce a fully haplotype-resolved diploid genome assembly for the common marmoset, providing insight into the heterozygosity spectrum and the evolution of the sex-differentiation region.
- Chentao Yang
- , Yang Zhou
- & Guojie Zhang
-
Article
| Open Access Progressive Cactus is a multiple-genome aligner for the thousand-genome era
Progressive Cactus is a multiple-genome aligner for the thousand-genome era
The Progressive Cactus program can create reference-free alignments of hundreds of large vertebrate genomes efficiently, and is used for the alignment of more than 600 amniote genomes.
- Joel Armstrong
- , Glenn Hickey
- & Benedict Paten
-
Analysis
| Open Access A comparative genomics multitool for scientific discovery and conservation
A comparative genomics multitool for scientific discovery and conservation
A whole-genome alignment of 240 phylogenetically diverse species of eutherian mammal—including 131 previously uncharacterized species—from the Zoonomia Project provides data that support biological discovery, medical research and conservation.
- Diane P. Genereux
- , Aitor Serres
- & Elinor K. Karlsson
-
Article
| Open Access Telomere-to-telomere assembly of a complete human X chromosome
Telomere-to-telomere assembly of a complete human X chromosome
High-coverage, ultra-long-read nanopore sequencing is used to create a new human genome assembly that improves on the coverage and accuracy of the current reference (GRCh38) and includes the gap-free, telomere-to-telomere sequence of the X chromosome.
- Karen H. Miga
- , Sergey Koren
- & Adam M. Phillippy
-
Article
| Open Access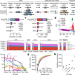 A structural variation reference for medical and population genetics
A structural variation reference for medical and population genetics
A large empirical assessment of sequence-resolved structural variants from 14,891 genomes across diverse global populations in the Genome Aggregation Database (gnomAD) provides a reference map for disease-association studies, population genetics, and diagnostic screening.
- Ryan L. Collins
- , Harrison Brand
- & Michael E. Talkowski
-
Article |
 Longitudinal molecular trajectories of diffuse glioma in adults
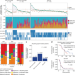
Longitudinal molecular trajectories of diffuse glioma in adults
The GLASS Consortium studies the evolutionary trajectories of 222 patients with a diffuse glioma to aid in our understanding of tumour progression and treatment failure
- Floris P. Barthel
- , Kevin C. Johnson
- & Roel G. W. Verhaak
-
Article |
 Structural variation in the gut microbiome associates with host health
Structural variation in the gut microbiome associates with host health
The authors systematically characterize structural variation in the genomes of gut microbiota and show that they are associated with bacterial fitness and with host risk factors, and that examining genes coded in these regions facilitates investigation of mechanisms that may underlie these associations.
- David Zeevi
- , Tal Korem
- & Eran Segal
-
Article
| Open Access New insights from uncultivated genomes of the global human gut microbiome
New insights from uncultivated genomes of the global human gut microbiome
Draft prokaryotic genomes from faecal metagenomes of diverse human populations enrich our understanding of the human gut microbiome by identifying over two thousand new species-level taxa that have numerous disease associations.
- Stephen Nayfach
- , Zhou Jason Shi
- & Nikos C. Kyrpides
-
Article |
 Predictable and precise template-free CRISPR editing of pathogenic variants
Predictable and precise template-free CRISPR editing of pathogenic variants
The authors use a machine-learning algorithm to predict the spectrum of CRISPR–Cas9-nuclease-mediated DNA repair outcomes at human genomic target sites.
- Max W. Shen
- , Mandana Arbab
- & Richard I. Sherwood
-
Letter |
 RNA velocity of single cells
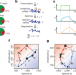
RNA velocity of single cells
RNA velocity, estimated in single cells by comparison of spliced and unspliced mRNA, is a good indicator of transcriptome dynamics and will provide a useful tool for analysis of developmental lineage.
- Gioele La Manno
- , Ruslan Soldatov
- & Peter V. Kharchenko
-
Letter
| Open Access Pan-cancer genome and transcriptome analyses of 1,699 paediatric leukaemias and solid tumours
Pan-cancer genome and transcriptome analyses of 1,699 paediatric leukaemias and solid tumours
Analysis of the genomes, exomes and transcriptomes of 1,699 childhood cancers identifies 142 driver genes.
- Xiaotu Ma
- , Yu Liu
- & Jinghui Zhang
-
Article
| Open Access The genome of Schmidtea mediterranea and the evolution of core cellular mechanisms
The genome of Schmidtea mediterranea and the evolution of core cellular mechanisms
An improved genome assembly for Schmidtea mediterranea shows that the genome is highly polymorphic and repetitive, and lacks multiple genes encoding core components of cell biological mechanisms.
- Markus Alexander Grohme
- , Siegfried Schloissnig
- & Jochen Christian Rink
-
Letter |
 The m1A landscape on cytosolic and mitochondrial mRNA at single-base resolution
The m1A landscape on cytosolic and mitochondrial mRNA at single-base resolution
Transcriptome-wide mapping of N1-methyladenosine (m1A) at single-nucleotide resolution reveals m1A to be scarce in cytoplasmic mRNA, to inhibit translation, and to be highly dynamic at a single site in a mitochondrial mRNA.
- Modi Safra
- , Aldema Sas-Chen
- & Schraga Schwartz
-
Letter
| Open Access Sequencing and de novo assembly of 150 genomes from Denmark as a population reference
Sequencing and de novo assembly of 150 genomes from Denmark as a population reference
A report of high-depth, short-read sequencing and de novo assemblies for 150 individuals from 50 parent–offspring trios as part of establishing a population reference genome for the GenomeDenmark project.
- Lasse Maretty
- , Jacob Malte Jensen
- & Mikkel Heide Schierup
-
Letter
| Open Access Improved maize reference genome with single-molecule technologies
Improved maize reference genome with single-molecule technologies
An improved reference genome for maize, using single-molecule sequencing and high-resolution optical mapping, enables characterization of structural variation and repetitive regions, and identifies lineage expansions of transposable elements that are unique to maize.
- Yinping Jiao
- , Paul Peluso
- & Doreen Ware
-
Letter |
 Capturing pairwise and multi-way chromosomal conformations using chromosomal walks
Capturing pairwise and multi-way chromosomal conformations using chromosomal walks
A conformation capture sequencing method is developed to link multiple genomic loci into three-dimensional proximity chains called chromosomal walks (C-walks), adding to our understanding of how higher-order chromosomal structures participate in genome regulation.
- Pedro Olivares-Chauvet
- , Zohar Mukamel
- & Amos Tanay
-
Article |
 Genomic architecture of heterosis for yield traits in rice
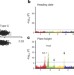
Genomic architecture of heterosis for yield traits in rice
Insights into the genomic architecture of heterosis for grain yield in rice are presented, and further mapping of grain yield loci resolves candidate genes that could be useful for breeding.
- Xuehui Huang
- , Shihua Yang
- & Bin Han
-
Letter |
 Mobile genes in the human microbiome are structured from global to individual scales
Mobile genes in the human microbiome are structured from global to individual scales
Mobile genes, which can be transferred between bacterial species in the microbiome to impart properties such as antibiotic resistance, are reflective of human activity and local diets.
- I. L. Brito
- , S. Yilmaz
- & E. J. Alm
-
Letter |
 Differential DNA repair underlies mutation hotspots at active promoters in cancer genomes
Differential DNA repair underlies mutation hotspots at active promoters in cancer genomes
Analysis of 1,161 cancer genomes across 14 cancer types shows that increased mutation density at gene promoters can be linked to transcription initiation activity and impairment of nucleotide excision repair.
- Dilmi Perera
- , Rebecca C. Poulos
- & Jason W. H. Wong
-
Article |
 Endosymbiotic origin and differential loss of eukaryotic genes
Endosymbiotic origin and differential loss of eukaryotic genes
Eukaryotes acquired their prokaryotic genes in two episodes of evolutionary influx corresponding to the origin of mitochondria and plastids, respectively, followed by extensive differential gene loss, uncovering a massive imprint of endosymbiosis in the nuclear genomes of complex cells.
- Chuan Ku
- , Shijulal Nelson-Sathi
- & William F. Martin
-
Article |
 Lagging-strand replication shapes the mutational landscape of the genome
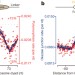
Lagging-strand replication shapes the mutational landscape of the genome
The emRiboSeq sequencing method is used to track polymerase activity genome-wide in vivo; despite Okazaki fragment processing, DNA synthesized by error-prone polymerase-α (Pol-α) is retained in vivo and comprises ∼1.5% of the genome, establishing Pol-α as an important source of genomic variability and providing a mechanism for site-specific variation in nucleotide substitution rates.
- Martin A. M. Reijns
- , Harriet Kemp
- & Martin S. Taylor
-
Article |
 The contribution of de novo coding mutations to autism spectrum disorder
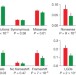
The contribution of de novo coding mutations to autism spectrum disorder
Family-based exome sequencing in a large autism study has identified 27 high-confidence gene targets and accurately estimates the contribution of both de novo gene-disrupting and missense mutations to the incidence of simplex autism, with target genes in affected females overlapping those in males of lower but not higher IQ; targets also overlap known targets for intellectual disability and schizophrenia, and are enriched for chromatin modifiers, FMRP-associated genes and embryonically expressed genes.
- Ivan Iossifov
- , Brian J. O’Roak
- & Michael Wigler
-
Article |
 Alterations of the human gut microbiome in liver cirrhosis
Alterations of the human gut microbiome in liver cirrhosis
Invasion of the gut by oral bacteria in liver cirrhosis.
- Nan Qin
- , Fengling Yang
- & Lanjuan Li
-
Article |
 Homologue engagement controls meiotic DNA break number and distribution
Homologue engagement controls meiotic DNA break number and distribution
DNA double-stranded breaks (DSBs) are shown to form in greater numbers in yeast cells lacking ZMM proteins, which are traditionally regarded as acting strictly downstream of DSB formation; these findings shed light on how cells balance the beneficial and deleterious outcomes of DSB formation.
- Drew Thacker
- , Neeman Mohibullah
- & Scott Keeney
-
Letter
| Open Access The genome of the recently domesticated crop plant sugar beet (Beta vulgaris)
The genome of the recently domesticated crop plant sugar beet (Beta vulgaris)
A full genome sequence is presented of sugar beet Beta vulgaris, the first plant belonging to Caryophyllales to have its genome sequenced; spinach was sequenced to enable inter-clade comparisons, and intraspecific variation was analysed by comparative genomics of a progenitor of all beet crops and additional sugar beet accessions.
- Juliane C. Dohm
- , André E. Minoche
- & Heinz Himmelbauer
-
Letter |
 Programmable single-cell mammalian biocomputers
Programmable single-cell mammalian biocomputers
In synthetic biology, the use of regulatory proteins that bind either DNA or RNA to reprogram mammalian cellular functions allows a variety of computational ‘logic circuits’ to be built in a plug-and-play manner, which may pave the way for precise and robust control of future gene-based and cell-based therapies.
- Simon Ausländer
- , David Ausländer
- & Martin Fussenegger

 The complex polyploid genome architecture of sugarcane
The complex polyploid genome architecture of sugarcane