Article
|
Open Access
Featured
-
-
Article
| Open Access Structural basis of Integrator-dependent RNA polymerase II termination
Structural basis of Integrator-dependent RNA polymerase II termination
Cryo-electron microscopy structures of the human Integrator complex in three different functional states shed light on how Integrator terminates RNA polymerase II (Pol II) transcription by disengaging Pol II from the DNA template.
- Isaac Fianu
- , Moritz Ochmann
- & Patrick Cramer
-
Article |
 Structural basis of exoribonuclease-mediated mRNA transcription termination
Structural basis of exoribonuclease-mediated mRNA transcription termination
A study presents two cryo-EM structures of yeast Pol II pre-termination transcription complexes bound to Rat1–Rai1, and provides the mechanisms for termination of mRNA transcription in yeast and other eukaryotes.
- Yuan Zeng
- , Hong-Wei Zhang
- & Yu Zhang
-
Article
| Open Access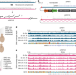 Synthetic reversed sequences reveal default genomic states
Synthetic reversed sequences reveal default genomic states
Introduction of a long synthetic DNA into yeast genomic loci results in high default transcriptional activity in yeast but low activity in mouse, suggesting distinct default levels of genomic activity in these organisms.
- Brendan R. Camellato
- , Ran Brosh
- & Jef D. Boeke
-
Article
| Open Access Incomplete transcripts dominate the Mycobacterium tuberculosis transcriptome
Incomplete transcripts dominate the Mycobacterium tuberculosis transcriptome
A study reveals that most transcripts in Mycobacterium tuberculosis are incomplete, likely because of the tendency of the transcription machinery in this species to pause on genomic DNA.
- Xiangwu Ju
- , Shuqi Li
- & Shixin Liu
-
Article
| Open Access Translation selectively destroys non-functional transcription complexes
Translation selectively destroys non-functional transcription complexes
Translation actively dislodges stalled transcription elongation complexes (ECs) from damaged DNA, which enables lesion repair and restoration of transcription activity, and coupled ribosomes discriminate between active ECs and stalled ECs, ensuring destruction of only the latter.
- Jason Woodgate
- , Hamed Mosaei
- & Nikolay Zenkin
-
Article
| Open Access Affinity-optimizing enhancer variants disrupt development
Affinity-optimizing enhancer variants disrupt development
Low-affinity transcription factor binding sites are prevalent across the genome, and single nucleotide changes that increase binding affinity even slightly can cause gain-of-function gene expression and phenotypes (such as polydactyly).
- Fabian Lim
- , Joe J. Solvason
- & Emma K. Farley
-
Article
| Open Access Single-cell analysis of chromatin accessibility in the adult mouse brain
Single-cell analysis of chromatin accessibility in the adult mouse brain
An atlas of candidate cis-regulatory DNA elements (cCREs) in the adult mouse brain unravels the transcriptional regulatory programs that drive the heterogeneity and complexity of brain structure and function.
- Songpeng Zu
- , Yang Eric Li
- & Bing Ren
-
Article
| Open Access Identification of constrained sequence elements across 239 primate genomes
Identification of constrained sequence elements across 239 primate genomes
Whole-genome alignment of 239 primate species reveals noncoding regulatory elements that are under selective constraint in primates but not in other placental mammals, that are enriched for variants that affect human gene expression and complex traits in diseases.
- Lukas F. K. Kuderna
- , Jacob C. Ulirsch
- & Kyle Kai-How Farh
-
Perspective |
 The status of the human gene catalogue
The status of the human gene catalogue
Although the catalogue of human protein-coding genes is nearing completion, the number of non-coding RNA genes remains highly uncertain, and for all genes much work remains to be done to understand their functions.
- Paulo Amaral
- , Silvia Carbonell-Sala
- & Steven L. Salzberg
-
Article
| Open Access The sex-specific factor SOA controls dosage compensation in Anopheles mosquitoes
The sex-specific factor SOA controls dosage compensation in Anopheles mosquitoes
A newly identified gene, sex chromosome activation (SOA), is a master regulator of dosage compensation in Anopheles gambiae.
- Agata Izabela Kalita
- , Eric Marois
- & Claudia Isabelle Keller Valsecchi
-
Article
| Open Access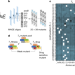 High-resolution landscape of an antibiotic binding site
High-resolution landscape of an antibiotic binding site
A collection of RNA polymerase mutants spanning all possible substitutions of the rifampicin binding site is generated and characterized, increasing our understanding of antibiotic mechanisms and bacterial physiology.
- Kevin B. Yang
- , Maria Cameranesi
- & Evgeny Nudler
-
Article
| Open Access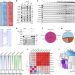 R-loop-dependent promoter-proximal termination ensures genome stability
R-loop-dependent promoter-proximal termination ensures genome stability
SOSS–INTAC stimulates promoter-proximal termination of transcription and attenuates R-loops associated with paused RNA polymerase II to prevent R-loop-induced genome instability.
- Congling Xu
- , Chengyu Li
- & Fei Xavier Chen
-
Article |
 OBOX regulates mouse zygotic genome activation and early development
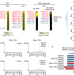
OBOX regulates mouse zygotic genome activation and early development
OBOX, PRD-like homeobox domain transcription factors (OBOX1–OBOX8), are key regulators of mouse zygotic genome activation and early embryogenesis.
- Shuyan Ji
- , Fengling Chen
- & Wei Xie
-
Matters Arising
| Open Access Reply to: Cold induction of nuclear FRIGIDA condensation in Arabidopsis
Reply to: Cold induction of nuclear FRIGIDA condensation in Arabidopsis
- Pan Zhu
- & Caroline Dean
-
Matters Arising
| Open Access Cold induction of nuclear FRIGIDA condensation in Arabidopsis
Cold induction of nuclear FRIGIDA condensation in Arabidopsis
- Zhicheng Zhang
- , Xiao Luo
- & Yuehui He
-
Article |
 Complementary Alu sequences mediate enhancer–promoter selectivity
Complementary Alu sequences mediate enhancer–promoter selectivity
Using RNA in situ conformation sequencing technology, the role of Alu elements in mediating the interaction between enhancers and promoters is shown.
- Liang Liang
- , Changchang Cao
- & Yuanchao Xue
-
Article
| Open Access Cooperation between bHLH transcription factors and histones for DNA access
Cooperation between bHLH transcription factors and histones for DNA access
Cryo-EM structures and analysis provide insight into the mechanisms by which basic helix–loop–helix transcription factors access E-box DNA sequences that are embedded within nucleosomes, and cooperate with other transcription factors.
- Alicia K. Michael
- , Lisa Stoos
- & Nicolas H. Thomä
-
Article
| Open Access Histone modifications regulate pioneer transcription factor cooperativity
Histone modifications regulate pioneer transcription factor cooperativity
Binding of the human pioneer transcription factor OCT4 to nucleosomes containing endogenous DNA sequences causes changes to the nucleosome structure and facilitates the cooperative assembly of multiple pioneer transcription factors, a property that can be affected by histone modifications.
- Kalyan K. Sinha
- , Silvija Bilokapic
- & Mario Halic
-
Article |
 γ-Linolenic acid in maternal milk drives cardiac metabolic maturation
γ-Linolenic acid in maternal milk drives cardiac metabolic maturation
The switch from glucose- to fatty acid-dependent metabolism in cardiomyocytes of newborn mice is governed by γ-linolenic acid in maternal milk, which binds to retinoid X receptors, thereby causing a transcription-dependent metabolic transition.
- Ana Paredes
- , Raquel Justo-Méndez
- & Mercedes Ricote
-
Article |
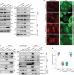 Mitotic bookmarking by SWI/SNF subunits
Mitotic bookmarking by SWI/SNF subunits
Subunits of SWI/SNF act as mitotic bookmarks to safeguard cell identity during cell division.
- Zhexin Zhu
- , Xiaolong Chen
- & Charles W. M. Roberts
-
Article
| Open Access Ageing-associated changes in transcriptional elongation influence longevity
Ageing-associated changes in transcriptional elongation influence longevity
Increases in transcriptional elongation speed with age affect organismal lifespan and ageing-related changes could be reversed with lifespan-extending interventions.
- Cédric Debès
- , Antonios Papadakis
- & Andreas Beyer
-
Article |
 Large-scale mapping and mutagenesis of human transcriptional effector domains
Large-scale mapping and mutagenesis of human transcriptional effector domains
A high throughput recruitment assay testing the transcriptional activity of more than 100,000 protein fragments tiling across most human chromatin regulators and transcription factors maps the locations and strengths of activation, repression and bifunctional domains, and identifies the sequences necessary for these functions.
- Nicole DelRosso
- , Josh Tycko
- & Lacramioara Bintu
-
Article
| Open Access H3K4me3 regulates RNA polymerase II promoter-proximal pause-release
H3K4me3 regulates RNA polymerase II promoter-proximal pause-release
Acute loss of H3K4me3 does not have detectable effects on transcriptional initiation, but leads to a widespread decrease in transcriptional output, an increase in RNA polymerase II pausing and slower elongation
- Hua Wang
- , Zheng Fan
- & Kristian Helin
-
Article |
 Structural basis of Rho-dependent transcription termination
Structural basis of Rho-dependent transcription termination
Structures presented in this study confirm decades of genetic and biochemical evidence for the mechanism of Rho-dependent termination in bacteria.
- Vadim Molodtsov
- , Chengyuan Wang
- & Richard H. Ebright
-
Article |
 Structural basis for intrinsic transcription termination
Structural basis for intrinsic transcription termination
Structural studies of Escherichia coli transcription intrinsic termination complexes representing distinct intermediates using cryo-electron microscopy provide insights into the steps and mechanism of transcription termination.
- Linlin You
- , Expery O. Omollo
- & Yu Zhang
-
Article |
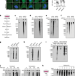 R-loop-derived cytoplasmic RNA–DNA hybrids activate an immune response
R-loop-derived cytoplasmic RNA–DNA hybrids activate an immune response
RNA–DNA hybrids are immunogenic species that can aberrantly accumulate in the cytoplasm after R-loop processing, linking R-loop accumulation to cell death through the innate immune response.
- Magdalena P. Crossley
- , Chenlin Song
- & Karlene A. Cimprich
-
Article
| Open Access A transcriptional switch controls sex determination in Plasmodium falciparum
A transcriptional switch controls sex determination in Plasmodium falciparum
A non-genetic mechanism of sex determination in the human malaria parasite, Plasmodium falciparum, is described, and the male development 1 gene is identified as a potential target for interventions that block malaria transmission.
- A. R. Gomes
- , A. Marin-Menendez
- & A. M. Talman
-
Article |
 Programmable RNA sensing for cell monitoring and manipulation
Programmable RNA sensing for cell monitoring and manipulation
RNA sensing-mediated payload expression provides a specific, versatile, simple and generalizable means of detecting and manipulating animal cells with broad potential applications.
- Yongjun Qian
- , Jiayun Li
- & Z. Josh Huang
-
Article |
 A mechanism for oxidative damage repair at gene regulatory elements
A mechanism for oxidative damage repair at gene regulatory elements
The nuclear mitotic apparatus protein NuMA helps to protect genes from oxidative damage by occupying regions around transcription start sites, binding DNA repair factors and promoting transcription following damage.
- Swagat Ray
- , Arwa A. Abugable
- & Sherif F. El-Khamisy
-
Article |
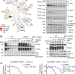 CDK11 regulates pre-mRNA splicing by phosphorylation of SF3B1
CDK11 regulates pre-mRNA splicing by phosphorylation of SF3B1
CDK11 associates with SF3B1 and phosphorylates threonine residues at the N terminus of SF3B1 during spliceosome activation, and the inhibition of CDK11 blocks the activation and leads to widespread intron retention and the accumulation of non-functional spliceosomes on pre-mRNAs and chromatin.
- Milan Hluchý
- , Pavla Gajdušková
- & Dalibor Blazek
-
Article |
 Recording gene expression order in DNA by CRISPR addition of retron barcodes
Recording gene expression order in DNA by CRISPR addition of retron barcodes
Retro-Cascorder, a system for time-ordered recording of transcriptional output, uses retrons as a tag to mediate DNA barcode acquisition in a CRISPR array.
- Santi Bhattarai-Kline
- , Sierra K. Lear
- & Seth L. Shipman
-
Article |
 Differential cofactor dependencies define distinct types of human enhancers
Differential cofactor dependencies define distinct types of human enhancers
The systematic categorization of human enhancers by their cofactor dependencies provides a conceptual framework to understand the sequence and chromatin diversity of enhancers and their roles in different gene-regulatory programmes.
- Christoph Neumayr
- , Vanja Haberle
- & Alexander Stark
-
Article |
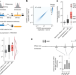 Compatibility rules of human enhancer and promoter sequences
Compatibility rules of human enhancer and promoter sequences
A new high-throughput assay applied to 1,000 enhancers and 1,000 promoters in human cells reveals how different classes of enhancers and promoters control RNA expression.
- Drew T. Bergman
- , Thouis R. Jones
- & Jesse M. Engreitz
-
Article |
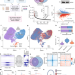 Gibbin mesodermal regulation patterns epithelial development
Gibbin mesodermal regulation patterns epithelial development
Characterization of Gibbin, encoded by AHDC1, offers insights into the epidermal and mesodermal patterning phenotypes seen in Xia–Gibbs and related syndromes in humans, which derive from abnormal mesoderm maturation as a result of gene-specific DNA methylation decisions.
- Ann Collier
- , Angela Liu
- & Anthony E. Oro
-
Article |
 Transcriptional coupling of distant regulatory genes in living embryos
Transcriptional coupling of distant regulatory genes in living embryos
In Drosophila, there are extensive physical and functional associations of distant paralogous genes, including co-regulation by shared enhancers and co-transcriptional initiation over distances of nearly 250 kilobases.
- Michal Levo
- , João Raimundo
- & Michael S. Levine
-
Article
| Open Access Gene regulation by gonadal hormone receptors underlies brain sex differences
Gene regulation by gonadal hormone receptors underlies brain sex differences
A study maps neuronal genomic targets of oestrogen receptor-α and shows how they coordinate brain sexual differentiation, concluding that the genome remains responsive to hormonal changes after structural dimorphisms have been established.
- B. Gegenhuber
- , M. V. Wu
- & J. Tollkuhn
-
Article
| Open Access Nonlinear control of transcription through enhancer–promoter interactions
Nonlinear control of transcription through enhancer–promoter interactions
The transcriptional effect of an enhancer depends on its contact probabilities with the promoter through a nonlinear relationship, and enhancer strength determines absolute transcription levels as well as the sensitivity of a promoter to CTCF-mediated transcriptional insulation.
- Jessica Zuin
- , Gregory Roth
- & Luca Giorgetti
-
Article |
 Basis of narrow-spectrum activity of fidaxomicin on Clostridioides difficile
Basis of narrow-spectrum activity of fidaxomicin on Clostridioides difficile
Structural analysis of Clostridioides difficile RNA polymerase in complex with fidaxomicin combined with biochemical, genetic and bioinformatic analyses identifies a key residue that determines fidaxomicin sensitivity.
- Xinyun Cao
- , Hande Boyaci
- & Elizabeth A. Campbell
-
Article |
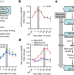 Crucial role and mechanism of transcription-coupled DNA repair in bacteria
Crucial role and mechanism of transcription-coupled DNA repair in bacteria
Integrated structure–function studies show that transcription-coupled DNA repair (TCR)—rather than global genomic repair—is responsible for most chromosomal repair events in bacteria, and that TCR mainly occurs independently of the Mfd translocase.
- Binod K. Bharati
- , Manjunath Gowder
- & Evgeny Nudler
-
Article |
 Aldehyde-driven transcriptional stress triggers an anorexic DNA damage response
Aldehyde-driven transcriptional stress triggers an anorexic DNA damage response
Endogenous formaldehyde accumulation reveals Cockayne syndrome in mice and stimulates production of the anorexiogenic peptide GDF15 in proximal tubule cells.
- Lee Mulderrig
- , Juan I. Garaycoechea
- & Ketan J. Patel
-
Article
| Open Access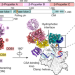 Structural basis of human transcription–DNA repair coupling
Structural basis of human transcription–DNA repair coupling
The authors resolve the structure of five complexes containing RNA polymerase II and the CSA and CSB proteins, offering insight into how the repair of DNA lesions is coupled to transcription.
- Goran Kokic
- , Felix R. Wagner
- & Patrick Cramer
-
Article |
 BANP opens chromatin and activates CpG-island-regulated genes
BANP opens chromatin and activates CpG-island-regulated genes
BANP is identified as the transcription factor that binds the CGCG element in a DNA-methylation-dependent manner, opens chromatin and activates a class of essential CpG-island-regulated genes.
- Ralph S. Grand
- , Lukas Burger
- & Dirk Schübeler
-
Article |
 Shape of promoter antisense RNAs regulates ligand-induced transcription activation
Shape of promoter antisense RNAs regulates ligand-induced transcription activation
The authors describe a role for the non-coding antisense transcripts produced at promoters in regulating ligand-induced activation of gene transcription.
- Fan Yang
- , Bogdan Tanasa
- & Michael G. Rosenfeld
-
Article |
 Enhancer release and retargeting activates disease-susceptibility genes
Enhancer release and retargeting activates disease-susceptibility genes
Disruption of a promoter can release its partner enhancer to activate other promoters in the same contact domain, and this process, named ‘enhancer release and retargeting’, can often lead to gene alterations that cause disease.
- Soohwan Oh
- , Jiaofang Shao
- & Michael G. Rosenfeld
-
Article |
 Structure of the human Mediator–RNA polymerase II pre-initiation complex
Structure of the human Mediator–RNA polymerase II pre-initiation complex
The structure of a recombinant 20-subunit version of human Mediator bound to the transcription pre-initiation complex is determined, providing insight into the regulation of RNA polymerase II initiation.
- Srinivasan Rengachari
- , Sandra Schilbach
- & Patrick Cramer
-
Article |
 Structures of mammalian RNA polymerase II pre-initiation complexes
Structures of mammalian RNA polymerase II pre-initiation complexes
The high-resolution structure of the mammalian pre-initiation complex in different functional states provides detailed insights into the mechanism of transcription initiation.
- Shintaro Aibara
- , Sandra Schilbach
- & Patrick Cramer
-
Article |
 A high-resolution protein architecture of the budding yeast genome
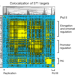
A high-resolution protein architecture of the budding yeast genome
A ChIP–exo method is used to define the genome-wide positional organization of proteins associated with gene transcription, DNA replication, centromeres, subtelomeres and transposons, revealing distinct protein assemblies for constitutive and inducible gene expression.
- Matthew J. Rossi
- , Prashant K. Kuntala
- & B. Franklin Pugh
-
Article |
 Systematic analysis of binding of transcription factors to noncoding variants
Systematic analysis of binding of transcription factors to noncoding variants
An ultra-high-throughput multiplex protein–DNA binding assay is used to assess binding of 270 human transcription factors to 95,886 noncoding variants in the human genome, providing data to improve prediction of the effects of noncoding variants on transcription factor binding and thereby increase understanding of molecular pathways involved in diverse human traits and genetic diseases.
- Jian Yan
- , Yunjiang Qiu
- & Bing Ren
-
Article |
 Small-molecule inhibitors of human mitochondrial DNA transcription
Small-molecule inhibitors of human mitochondrial DNA transcription
Inhibitors of mitochondrial transcription that target human mitochondrial RNA polymerase provide a chemical biology tool for studying the role of mitochondrial DNA expression in a wide range of pathologies.
- Nina A. Bonekamp
- , Bradley Peter
- & Nils-Göran Larsson
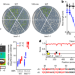
 The temperature sensor TWA1 is required for thermotolerance in Arabidopsis
The temperature sensor TWA1 is required for thermotolerance in Arabidopsis