Abstract
Breast neuroendocrine neoplasms (NENs) constitute a rare histologic subtype that includes both neuroendocrine tumors (NETs) and neuroendocrine carcinomas (NECs). In this study, we aimed to gain insight into the clinical and molecular characteristics of NENs of the breast. NEN and paired distant normal fresh tissues and clinicopathological data were obtained from 17 patients with NENs, and clinicopathological data were collected from 755 patients with invasive breast carcinomas of no special type (IBCs-NST). We compared the clinicopathological characteristics of NENs and IBCs-NST and performed whole-exome sequencing (WES) of both NEN and paired normal tissues. Compared with the IBC-NST patients, the NEN patients had a higher mean age, lower clinical stage, and lower pathological nodal (pN) stage (P < 0.001, P < 0.001, and P = 0.017, respectively). The most frequently mutated gene in NENs was KMT2C (3/17, 17.6%). NENs had copy number variations (CNVs) of 8q, 11q, and 17q amplification and 17q and 11q deletion and harbored the following specific genes related to tumorigenesis: (i) suppressor genes with loss of heterozygosity (LOH) such as ACE (2/17, 11.8%); (ii) tumor driver genes such as GATA3 (2/17, 11.8%); and (iii) susceptibility genes such as MAP3K4 (17/17, 100%) and PDE4DIP (17/17, 100%). The oncogenic/likely oncogenic mutations of NETs in PI3K pathway genes (50.0%, 18.2%; P < 0.001) and MAPK signaling pathway genes (83.3%, 18.2%; P = 0.035) affected higher proportions than those of NECs. In conclusion, this study provides certain clinical and molecular evidence supporting NENs as a distinct subtype of breast cancer and provides some potential molecular features for distinguishing NETs from NECs.
Similar content being viewed by others
Introduction
Neuroendocrine neoplasms (NENs) of the breast constitute an extremely rare histologic subtype that includes neuroendocrine tumors (NETs) and neuroendocrine carcinomas (NECs). The latest WHO classification (2019) aimed to reduce discrepancies and contradictions in their terminology, subgroups, and criteria for histologic grading and staging by unifying the definitions of other organ systems with a uniform classification framework of NENs. The definition of breast NENs in the latest classification provides the most stringent diagnostic criteria compared to previous WHO classifications;1,2,3 the term NEN was characterized by diffuse neuroendocrine (NE) marker expression and NE morphology, regardless of Nottingham histologic grade1,4. However, few studies have investigated NENs based on the latest version of the classification criteria, and various terminologies and diagnostic criteria for breast cancer with NE differentiation have been used in earlier studies5,6,7, resulting in a poor understanding of the clinicopathological and molecular features of NENs.
Although prior studies have provided some clinicopathological features of NEN, many issues remain unsolved. Many studies using the 2003 criteria showed that NENs were more likely to be older, have larger tumors, be estrogen receptor/progesterone receptor (ER/PR)-positive and be human epidermal growth factor receptor 2 (HER2)-negative than invasive breast cancers (IBCs-NST)2,8,9. In addition, some published studies revealed adverse outcomes in NE breast cancers7,10,11,12,13,14,15, and most of the above studies considered NEN as a whole entity rather than dividing it into subgroups. Thus, the clinicopathological factors, prognosis, and treatment strategies for NENs and even the subgroups of NETs and NECs require more data accumulation.
Studies focusing on molecular characteristics provide certain molecular evidence regarding NENs of the breast. Compared with mucinous breast carcinomas (MBCs), breast NECs have different copy number variation (CNV) levels as follows:16,17,18 1q increase and 16q deletion coexist in 28% of NECs; breast NETs are characterized by an enrichment of transcriptional mutations and are considered representative of a spectrum of entities histologically and genetically similar to MBCs. In addition, the majority of breast NETs are distinct from ER-positive/HER2-negative IBCs-NST19. Moreover, although a recent study conducted in 2021 demonstrated that NETs had better disease-specific survival (DSS) and overall survival (OS) than NECs12, the differences between NETs and NECs have not been confirmed at the molecular level.
Due to the use of diverse nomenclature and diagnostic criteria for NENs of the breast in different studies, the clinicopathological features, molecular characteristics and therapeutic strategies of NEN remain largely known. Thus, our study aimed to (i) compare the differences in the clinicopathological characteristics and prognosis between NENs and IBCs-NST, (ii) identify the molecular characteristics of NENs, and (iii) find the differences between NETs and NECs.
Materials and methods
NEN and IBC-NST case collection
Patients diagnosed with NEN from 2013 to 2018 and IBC-NST from 2008 to 2019 were collected from the surgical pathology files of West China Hospital, Sichuan University. The inclusion and exclusion criteria for NEN were as follows:1 (i) cancers with > 90% NEN pattern; (ii) tumor cells with ≥90% chromagranin A (CgA) and/or synaptophysin (Syn) expression; (iii) NENs with low Nottingham grade (grade 1–2) were classified as NETs, and high Nottingham grade NENs were classified as high grade as NECs; (iv) special breast cancers with the expression of NE markers, such as solid papillary carcinoma in situ, MBCs, were excluded.
Fresh NEN tissues and matched NEN distant normal tissues were extracted from the biobank of West China Hospital, Sichuan University, but fresh tissue from the control group was not available. Clinicopathologic data, including age, menstrual status, laterality, clinical tumor (cT) stage, clinical nodal (cN) stage, clinical (TNM) stage, pathological tumor (pT) stage, pathological node (pN) stage, and follow-up information of patients with NENs were collected from the Breast Cancer Patient Follow-up database of West China Hospital, Sichuan University.
Histological review and immunohistochemistry (IHC) for NEN and IBC-NST
Hematoxylin and eosin (H&E)-stained slides were reviewed by 3 pathologists under a multi-head microscope (BW, YW, and XK) to morphologically confirm whether each case met the diagnostic criteria of the 5th edition of the WHO classification in 20191.
Immunohistochemical staining was performed using the selected most representative paraffin block using the following antibodies: ER, PR, HER2, Ki67, CgA, and Syn. See Supplementary Table 1 for the details of the antibodies.
ER and PR expression was assessed based on the 2010 American society of clinical oncology/college of American pathologists (ASCO/CAP) guideline recommendations20. HER2 status was assessed according to the ASCO/CAP clinical practice guideline focused update21. A cutoff of 20% was used for the evaluation of Ki67 expression22. Tumor cells expressing ≥90% CgA and/or Syn were considered positive1.
Comparison of clinicopathological features in NEN and IBC-NST patients
SPSS 25.0 (SPSS, Chicago, IL, USA) statistical analysis software was used to compare the differences in the clinicopathological characteristics among the groups (NETs vs. NECs and NENs vs. IBCs-NST). The descriptive statistics of the demographic and clinicopathologic factors were calculated, and the differences among the groups were evaluated by the chi-square or Fisher exact test, as appropriate. Kaplan-Meier curves and statistics (log-rank) were used for survival data representation and analysis23. Propensity score matching (PSM)24 was used to balance the important characteristics of the NEN and IBC-NST patients (ratio 1:5). Two-tailed P values less than 0.05 were considered to indicate statistical significance.
DNA extraction and quality assessment of NEN fresh tissues
DNA was purified from fresh NEN tissues and matched normal tissues using a Qiagen kit (QIAamp System, Germany) and amplified by polymerase chain reaction (PCR). DNA quality control included the assessment of degradation and contamination by 0.1% agarose gel electrophoresis. In addition, the DNA concentration was measured (Qubit® DNA Assay Kit, Invitrogen, USA), and only qualified fresh samples were used for whole-exome sequencing (WES).
Whole-exome sequencing of 17 paired NEN and normal fresh tissues
According to the above DNA evaluation results, samples with DNA concentrations ≥20 ng/µL and total amounts >0.6 µg were used to generate a library (Agilent Technologies, CA, USA). Then, the genomic DNA was randomly broken into fragments of 180–280 bp in length (Covaris, Massachusetts, USA). The remaining short DNA fragments were converted into blunt ends by exonuclease/polymerase activities. When a poly(A) tail was added to the short DNA fragments, they were linked by adapter oligonucleotides. The ligated DNA fragments were enriched by PCR. The library with a specific index was subjected to liquid phase hybridization with up to 543,872 biotin-labeled probes. Then, 334,378 exons of 20,965 genes were captured by magnetic beads with streptomycin, which was linearly enriched by PCR. Finally, the DNA was purified (Beckman Coulter, Beverly, USA) and quantified (Agilent Bioanalyzer 2100 system), and then, the DNA was prepared for sequencing.
The DNA samples were clustered according to the manufacturer’s instructions (HiSeq PE Cluster Kit, Illumina). Then, DNA libraries of appropriate sizes were sequenced on the Illumina HiSeq PE150 platform.
Bioinformatics analyses
The analyses mainly included the following three aspects: data quality evaluation, NEN molecular characteristics analyses, and comparative analyses of NET and NEC characteristics. The quality evaluation of the raw data included sequencing data filtering, sequencing error rate distribution checking, sequencing data quality distribution, and reference sequence alignment analysis (sequencing depth and coverage distribution). The NEN molecular characteristics analyses included DNA mutation analyses, loss of heterozygosity (LOH) analysis, driver gene, and susceptibility gene identification. See Supplementary Materials and methods for the details of the mutation signature analysis. The differences in the gene pathways enriched by somatic mutations after excluding germline mutations between the NETs and NECs were also compared. In the comparative pathway analysis, the genes were obtained from PathCards (https://pathcards.genecards.org/).
The details of raw sequencing data cleaning are shown in the Supplementary Materials and methods.
Results
Comparison of the characteristics in NEN and IBC-NST patients
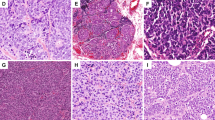
In total, 17 patients with NEN and 755 patients with IBC-NST were assessed in this study. Among the NEN cases, 35.3% (6/17) were NETs, and 64.7% (11/17) were NECs. Representative cases of NEN are shown in Fig. 1A–C. The clinicopathological characteristics, treatment strategies, and prognostic information of the 17 NEN patients are shown in Table 1.
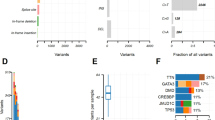
A Low-grade of NEN (H&E, Nottingham grade 2). IHC of NEN showing diffuse, uniform immunoreactivity of CgA B and Syn C. D Mutation spectrum analysis showing that the most frequent mutation type in the NENs was C > T/G > A, followed by T > C/A > G, and mutation type with the lowest frequency was T > A/A > T. Each row in the figure represents a sample, and each column represents a mutation type. The darker the mutation type, the higher proportion of the sample. E Proportion of each mutation feature in different samples. Nonnegative matrix factorization (NMF) cluster analyses revealed three different mutation signatures. The Y-axis indicates the proportion of acquired mutation signatures. Signature B was found to be identical to COSMIC mutation feature 2 with a cosine similarity greater than 0.9, and it was found in 70.6% (12/17) of the NENs in this study. F Frequency mutation gene analysis showing that the most frequently mutated genes in NENs were KMT2C (3/17, 17.6%), ACE (2/17, 11.8%), ANKRD18B (2/17, 11.8%), and CACNA1B (2/17, 11.8%). The X-axis on the bottom reflects frequency mutations, the X-axis on the top reflects the gene name, and the Y-axis reflects the sample name. The different colors in the map represent different types of mutations. This figure shows only some genes with high mutation frequencies, and the full list is shown in Supplementary Table 3.
The baseline characteristics and comparison results of the NENs and IBCs-NST are shown in Table 2. The mean age of the NEN patients (50.0 years) was higher than that of the IBC-NST patients (47.6 years) (P < 0.001). Low clinical tumor stage (cT1 and cT2), lymph node stage (cN0), and clinical stage (stage I and stage II) were more common in the NEN group than in the IBC-NST group (P < 0.001, P < 0.001, P = 0.001). More NET patients had pN0 stage than stage-matched IBC-NST patients (83.3% vs.26.1%, P = 0.017). NEC patients exhibited higher PR expression than that in IBC-NST patients (90.9% vs. 68.2%; P = 0.030).
The patients with NEN were matched 1:5 to IBC-NST, and 10 NEN and 42 IBC-NST patients were included in the Kaplan-Meier analysis. The Kaplan-Meier analysis showed that there were no significant differences in the OS or disease-free survival (DFS) between the NEN and IBC-NST patients, as shown in Supplementary Fig. 1A, B.
Sequencing data quality assessment
The ratio of the high-quality data (clean reads) of the 17 paired samples to the original test data (raw data) was greater than 90%; the error rate was less than 0.1%; and the quality of the sequencing data was distributed at Q30 (≥80%). The data quality output details of the NENs are shown in Supplementary Table 2. The analyses of the sequencing depth and coverage distribution of the NENs showed that the sequencing depth was mainly concentrated at 100×; the coverage depth of each chromosome was mainly distributed at 100× (Supplementary Fig. 2).
Mutation spectrum analyses related to the characteristics of NENs
The genetic mutation spectrum analysis of the NENs showed that the mutation type with the highest frequency was C > T/G > A (100%, 17/17), followed by T > C/A > G. The nonnegative matrix factorization (NMF) cluster analyses revealed three different mutation signatures: Signature A (near signature 1, filter with cosine similarity = 0.82), Signature. B (near signature 2, filter with cosine similarity = 0.91), and Signature C (near signature 6, filter with cosine similarity = 0.70). If a filter has a cosine similarity > 0.9, the mutation feature of this sample and the known mutation feature are the same (Fig. 1D, E).
Frequency mutation gene analysis of the NENs showed that KMT2C was the most frequently mutated gene (17.6%, 3/17), followed by ACE (2/17, 11.8%), ANKRD18B (2/17, 11.8%), and CACNA1B (2/17, 11.8%) (Fig. 1F). This figure only partially shows some genes with mutation frequencies; the results of all mutated genes are shown in Supplementary Table 3.
The high-frequency CNV analysis showed that the NENs had amplification of 8q, 11q,17q, and 19q and deletion of 17q and 11q. The CNV distribution in each NEN sample and the high-frequency CNVs of all NENs are shown in Fig. 2A, B, respectively. The tumor clonal structure analysis showed that a high proportion of the NENs (9/17, 52.9%) were heterogeneous tumors. NENs with and without heterogeneity are shown in Supplementary Fig. 3.
A CNV distribution in each NEN sample. The X-axis is chromosomes 1-22, and the Y-axis is the NEN sample names. In the figure, red and blue represent somatic CNVs as follows: red represents an increase in the copy number, and blue represents a decrease in the copy number. The darker the color, the greater the change in the copy number. B High-frequency CNV analysis showed amplification of 8q, 11q, 17q, and 19q and deletion of 11q and 17q in all NEN cases. The X-axis is chromosomes 1–22; the Y-axis is the score of the high-frequency CNV segment by GISTIC software. The higher the score is, the higher the frequency of CNV in this segment. AMP (red) indicates an increase in the copy number, and DEL (blue) indicates a decrease in the copy number.
Loss of heterozygosity and driver gene analyses in NEN tissues
The LOH analysis results showed that 88.2% (15/17) of the NEN patients exhibited LOH in tumor suppressor genes. Both NEN-4 and NEN-14 (2/17, 11.8%) showed LOH in the suppressor gene ACE on chromosome 17. Most suppressor genes with LOH in NENs were located on chromosome 11 (11/16, 94.1%), chromosome 3 (14/17, 82.4%), the X chromosome (12/17, 70.6%), chromosome 6 (10/17, 58.8%), and chromosome 8 (9/17, 52.9%).
The predictive analysis of the driver genes indicated that GATA3 (2/17, 11.8%), KMT2C (2/17, 11.8%), TRRAP (2/17, 11.8%), and SPEN (2/17, 11.8%) were the driver genes of NENs (Fig. 3A).
A Landscape of driver genes in NENs. The X-axis is the sample, the Y-axis is the gene, the top is the number of gene mutations in each sample, and the right is the number of samples per mutant gene. This analysis was only based on somatic mutations in NEN tissues. B Landscape of susceptibility genes in NENs. Susceptibility genes were screened from germline mutations. The X-axis is the NEN-matched normal sample name, the Y-axis is the gene, the top is the number of gene mutations in each sample, and the right is the number of samples per mutant gene.
Susceptibility gene analyses in tumor distant normal tissues
The following susceptibility genes were found in the NEN-matched normal tissues: MAP3K4 (17/17, 100%), PDE4DIP (17/17, 100%), NCOR2 (15/17, 88.2%), BPTF (14/17, 82.4%), MAP3K1 (14/17, 82.4%), and MUC20 (14/17, 82.4%) (Fig. 3B).
Analysis related to the treatment of NENs
This study compared somatic mutations in NENs to previously known databases such as PD FDA (KEGG-based targeted drug data), PharmGKB, DrugBank, My Cancer Genome, and therapeutic trial findings. In total, 76.5% (13/17) of NENs may harbor targetable mutations, rendering them candidates for targeted therapy. (Supplementary Fig. 4 and Supplementary Table 4).
The mutant genes that may be related to drug resistance in NENs were PIK3CA (2/17, 11.8%), CAT (1/17, 5.9%), AKTI (1/17, 5.9%), TP53 (1/17, 5.9%), and ABCC6 (1/17, 5.9%) (Supplementary Table 5).
Comparative analyses of NETs and NECs
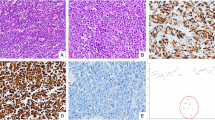
The comparative analysis of NETs and NECs showed that Ki67 expression and Nottingham grade in NECs were significantly higher than those in NETs (P = 0.029) (Fig. 4A, B), but there was no significant difference in OS and DFS between NETs and NECs (P = 0.540, P = 0.377, respectively; Supplementary Fig. 1C, D).
A, B Comparison of the baseline characteristics between NETs and NECs. C, D Landscape of frequency mutated genes in NETs and NECs. E, F Comparative analyses of 7 pathways between NETs and NECs. Comparative analysis of NETs and NECs showed that Ki67 expression in NECs was significantly higher than that in NETs. ACE (33.3%, 2/6) was the most frequently mutated gene in the NETs, while KMT2C (18.2%, 2/11) was the most frequently mutated gene in the NECs. The oncogenic/likely oncogenic mutations of NETs in PI3K pathway genes (P < 0.001) and MAPK signaling pathway genes (P = 0.035) affected higher proportions than those of NECs. *P < 0.05, Fisher’s exact test. The X-axis is the sample type, and the Y-axis is the percentage of NENs with genetic mutations in each genetic pathway.
The mutated gene with the highest frequency in the NETs was ACE (2/6, 33.3%), followed by KMT2C (1/6, 16.7%) and ANKRD18B (1/6, 16.7%); the mutated gene with the highest frequency in the NECs was KMT2C (2/11, 18.2%), followed by CSMD1(2/11, 18.2%) and KRT32(2/11, 18.2%). The most common mutation types in NETs and NECs were nonsense mutations (3/6, 50.0%) and missense mutations (8/11, 72.7%), respectively. (Fig. 4C, D)
The analysis of the differences in gene pathways between NETs and NECs showed that the pathway with the highest gene mutation rate in NETs is MAPK (83.3%), followed by RTK/ras (50.0%), PI3K/AKT (33.3%), chromatin remodelers (33.3%), retinoblastoma (RB) (33.3%), and p53 (0.0%); the pathway with the highest gene mutation rate in NECs is chromatin remodelers (54.5%), followed by RB (27.3%), PI3K/AKT (18.2%), MAPK (18.2%), RTK/ras (18.2%) and p53 (18.2%) (Figs. 4E, F and Table 2). The genes contained in the gene pathways in this study are shown in Supplementary Table 6. Oncogenic/likely oncogenic mutations in chromatin remodelers, RB pathway genes, p53 pathway genes, and RTK/Ras signaling pathway genes affected similar proportions of the NETs and NECs (P = 0.620, P = 1.000, P = 0.515, P = 0.280, respectively); the oncogenic/likely oncogenic mutations of NETs in PI3K pathway genes (3/6, 50.0%; 2/11, 18.2%; P < 0.001) and MAPK signaling pathway genes (5/6, 83.3%; 2/11, 18.2%; P = 0.035) affected higher proportions than those of NECs.
Discussion
NEN patients tend to be older, have a lower clinical stage, be hormone receptor (HR)-positive, HER2-negative and have a low Ki67 proliferation index12. Breast cancers are diagnosed at various ages in different parts of the world;25,26,27,28 the peak age of breast cancer in some Asian and African countries was even over 10 years earlier than that in European or American countries26,27,28,29,30. As a result, factors impacting the age of onset of NENs, such as geography and ethnicity, should also be a considered in addition to the tumor subtype. In addition, most NENs and IBCs-NST with NE differentiation (IBC-NST-NE) expressed ER and PR, lacked HER2 expression, and had a low Ki67 proliferation index according to previous studies7,15,31, which is consistent with our finding in NENs. Regarding NECs, a recent study investigating small cell neuroendocrine carcinomas (SCNECs) showed that 30–50% of cases were ER-positive, and none of the cases were HER2-positive;32,33 some studies found PR expression in SCNECs, while the others did not32,33,34.
Our study found that KMT2C was the most frequently mutated gene in breast NENs; the mutation appears in both breast NENs and other types of breast cancer, while the mutation rate may differ between the two types above. A study revealed that the frequency of KMT2C mutations in IBCs-NST-NE was higher than that in NETs19. In addition, another study investigating stomach NEC and mixed NEN found alterations in KMT2C in NE components only35, and another recent study showed that KMT2C was the most frequently mutated gene in inflammatory breast cancer36, suggesting that a mutation of KMT2C may play an important role in NE differentiation in breast cancer. Moreover, KMT2C was identified as a driver gene with relatively high mutation rates in our study, indicating that KMT2C may play a driving role in the progression of breast cancer. Interestingly, mutant KMT2C may be related to resistance to HR therapy37, as a previous study showed that in an HR-rich environment, KMT2C can regulate enhancers and promote tumor proliferation38, possibly playing an important role in ER-positive breast cancer. Two cases of NEN in our study were ER-negative, which is consistent with the results of the above.
Compared with IBCs-NST, NENs have a unique gene mutation spectrum as follows: lower PIK3CA and TP53 mutation rates and higher FOXA1 and TBX3 mutation rates39,40. The mutation rate of GATA3 in our study was slightly lower than that in Marchiò’s study (16.7%) but was close to that reported by Fresia Pareja (10%)19,40. Noticeably, the mutation rate of GATA3 in our study did not significantly differ from that in IBC-NST but was higher than that in IBCs-NST in the study by Marchiò39,40. In addition, Lavigne10 found that the mutation rate of PIK3CA in primary NECs was consistent with the rate in The Cancer Genome Atlas (TCGA) database (9%) but lower than that in our study (11.8%). Although IBCs-NST-NE may account for 12.5% of all IBCs-NST6, studies comparing NENs and this group of tumors are limited.
NENs have some intrinsic CNV features; the incidence of some CNVs may provide some evidence suggesting that NEN is an independent subtype. Both our and Marchiò’s studies40 found 8q amplification in NENs, and Bao et al41 discovered multiple gene region CNVs, which were partially similar to our findings and related to lymph node metastasis; the variation of 8q in NENs may be a high-frequency CNV and potential target for tumor therapy. In addition, breast NETs were found to harbor a lower frequency of 1q gains and 16q losses than ER-positive/HER2-negative breast cancers16,19. Compared with that in luminal breast cancer and lobular carcinoma, the incidence of 1q amplification and 16q deletion in NENs was lower and differed from that in MBCs with NE differentiation17,18, supporting that NEN is an independent breast cancer subtype. Additionally, breast tumors with 16q deletion were related to a small tumor size, low histological grade, decreased lymph node metastasis, and high ER expression, suggesting that 16q deletion is a good prognostic marker for breast cancer42,43.
The specific genes identified in this study, such as suppressor genes and tumor susceptibility genes, suggest that tumorigenesis is a multistep process. First, we found that NEN had a loss of heterozygosity in the suppressor gene ACE on chromosome 17 as previously reported by a study which may be related to more aggressive breast cancer44. Second, the top three mutated susceptibility genes in our study were MAP3K4, PDE4DIP, and NCOR2. Mutation of MAP3K4 can regulate filamin expression and neuronal migration45, suggesting that the neuroendocrine function in the NENs may be obtained from neurons immigrating from other sites; MAP3K4 also plays an important role in multiple signaling pathways46,47,48,49. Although the significance of PDE4DIP in breast cancer is still unknown, recent studies have revealed that PDE4DIP has a high predictive value for neuroblastoma prognosis50. NCOR2 mutation was found in ER-positive breast cancer with local recurrence51 and may be related to tamoxifen resistance in primary breast cancer52.
Studies on pancreatic carcinomas have revealed the different genetic phenotypes of NETs and NECs, which can be used clinically to distinguish these two tumors with quite distinct therapeutic strategies and outcomes53,54,55. However, to date, research comparing the mutated genes involved in different gene pathways in breast NETs and NECs has not been reported. In this study, the percentages of NETs with gene mutations involved in the PI3K and MAPK signaling pathways were significantly higher than those of NECs, suggesting that the detection of genes in those two pathways may be a novel method to discriminate breast NETs and NECs. Interestingly, the MAPK pathway is highly ranked in the NEC component of gastric mixed NENs56, and MYC family pathways were commonly found in prostate NECs57 and cervical NECs58 but not in breast NENs, suggesting that the pathogenic mechanism of breast NENs may differ from that of NE tumors of other systems.
Our study has some limitations. First, the number of samples is still limited due to the low incidence of NEN under strict diagnostic criteria. Second, the fresh tissues used for WES were more difficult to obtain than paraffin samples.
In conclusion, comprehensive clinical and genomic analyses of breast NEN revealed that NENs are an entity distinct from IBCs-NST in clinicopathological characteristics, show that the tumorigenesis of NENs may be a multistep process resulting from multiple genetic alterations, and present some potential molecular features to distinguish breast NETs from NECs.
Data availability
The sequencing data generated during the current study are available in the NCBI Sequence Read Archive (SRA) database (PRJNA741833) and can be accessed at the following link: https://dataview.ncbi.nlm.nih.gov/object/PRJNA741833?reviewer=knskiu0jsm5h4lin58l2m7rejg.
References
WHO Classification of Tumors Editorial Board. WHO Classification of Tumors. Breast Tumors. 5th ed. IARC: Lyon, (2019).
Tavassoli F. A. & Devilee P. World Health Organisation classification of tumors. Pathology and genetics of tumors of the breast and female genital organs. IARC: Lyon, (2003).
Lakhani S. R., Ellis I. O., Schnitt S. J., Tan P. H. & van de Vijver M. J. World Health Organisation classification of tumors of the Breast. 4th ed. IARC: Lyon, (2012).
Tsang, J. Y. & Tse, G. M. Breast cancer with neuroendocrine differentiation: an update based on the latest WHO classification. Mod Pathol 34, 1062–1073 (2021).
Kwon, S. Y. et al. Neuroendocrine differentiation correlates with hormone receptor expression and decreased survival in patients with invasive breast carcinoma. Histopathology 64, 647–659 (2014).
Miremadi, A. et al. Neuroendocrine differentiation and prognosis in breast adenocarcinoma. Histopathology 40, 215–222 (2002).
Bogina, G. et al. Neuroendocrine differentiation in breast carcinoma: clinicopathological features and outcome. Histopathology 68, 422–432 (2016).
Wei, B. et al. Invasive neuroendocrine carcinoma of the breast: a distinctive subtype of aggressive mammary carcinoma. Cancer 116, 4463–4473 (2010).
Zhang, Y. et al. Invasive neuroendocrine carcinoma of the breast: a prognostic research of 107 Chinese patients. Neoplasma 60, 215–222 (2013).
Lavigne, M. et al. Comprehensive clinical and molecular analyses of neuroendocrine carcinomas of the breast. Mod Pathol 31, 68–82 (2018).
Roininen, N. et al. Primary neuroendocrine breast carcinomas are associated with poor local control despite favourable biological profile: a retrospective clinical study. BMC Cancer 17, 1–7 (2017).
Yang, L. et al. Validation of prognostic significance of the proposed uniform classification framework in neuroendocrine neoplasms of the breast. Breast Cancer Res Treat 186, 403–415 (2021).
Kawasaki, T. et al. Neuroendocrine cells associated with neuroendocrine carcinoma of the breast: nature and significance. J Clin Pathol 65, 699–703 (2012).
Kimura, N., Yoshida, R., Shiraishi, S., Pilichowska, M. & Ohuchi, N. Chromogranin A and chromogranin B in noninvasive and invasive breast carcinoma. Endocr Pathol 13, 117–122 (2002).
Lai, B. S. et al. The Clinical Significance of Neuroendocrine Features in Invasive Breast Carcinomas. Oncologist 25, e1318–e1329 (2020).
Natrajan, R. et al. Loss of 16q in high grade breast cancer is associated with estrogen receptor status: Evidence for progression in tumors with a luminal phenotype? Genes Chromosome Canc 48, 351–365 (2009).
Lacroix-Triki, M. et al. Mucinous carcinoma of the breast is genomically distinct from invasive ductal carcinomas of no special type. J Pathol 222, 282–298 (2010).
Horlings, H. M. et al. Genomic profiling of histological special types of breast cancer. Breast Cancer Res Treat 142, 257–269 (2013).
Pareja, F. et al. Neuroendocrine tumors of the breast: a genomic comparison with mucinous breast cancers and neuroendocrine tumors of other anatomic sites. J Clin Pathol 0, 1–8 (2020).
Hammond, M. E., Hayes, D. F., Wolff, A. C., Mangu, P. B. & Temin, S. American society of clinical oncology/college of american pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Oncol Pract 6, 195–197 (2010).
Wolff, A. C. et al. Human Epidermal Growth Factor Receptor 2 Testing in Breast Cancer: American Society of Clinical Oncology/College of American Pathologists Clinical Practice Guideline Focused Update. Arch Pathol Lab Med 142, 1364–1382 (2018).
Goldhirsch, A. et al. Personalizing the treatment of women with early breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann Oncol 24, 2206–2223 (2013).
Breslow N. E. Introduction to Kaplan and Meier (1958) Nonparametric Estimation from Incomplete Observations. Breakthroughs in Statistics (1992).
Porro, G., King, G. & Iacus, S. M. cem: Software for Coarsened Exact Matching. J Stat Softw 30, 1–27 (2009).
Lee, J. H. et al. Population-based breast cancer statistics in Korea during 1993-2002: incidence, mortality, and survival. J Korean Med Sci 22(Suppl), 11–16 (2007).
DeSantis, C. E. et al. Breast cancer statistics, 2019. CA Cancer J Clin 69, 438–451 (2019).
Lei, S. et al. Breast cancer incidence and mortality in women in China: temporal trends and projections to 2030. Cancer Biol Med 18, 900–909 (2021).
Lei S., et al. Global patterns of breast cancer incidence and mortality: A population-based cancer registry data analysis from 2000 to 2020. Cancer Commun, 1-12 (2021).
Sun, K. X. et al. Incidence trend and change in the age distribution of female breast cancer in cancer registration areas of China from 2000 to 2014. Chi J Prev Med 52, 567–572 (2018).
Hori, M. et al. Cancer incidence and incidence rates in Japan in 2009: a study of 32 population-based cancer registries for the Monitoring of Cancer Incidence in Japan (MCIJ) project. Jpn J Clin Oncol 45, 884–891 (2015).
Sapino, A. et al. Expression of apocrine differentiation markers in neuroendocrine breast carcinomas of aged women. Mod Pathol 14, 768–776 (2001).
Shin, S. J., DeLellis, R. A., Ying, L. & Rosen, P. P. Small cell carcinoma of the breast: a clinicopathologic and immunohistochemical study of nine patients. Am J Surg Pathol 24, 1231–1238 (2000).
McCullar, B. et al. Correction to: Genomic landscape of small cell carcinoma of the breast contrasted to small cell carcinoma of the lung. Breast Cancer Res Treat 173, 243 (2019).
Sadanaga, N. et al. Clinical characteristics of small cell carcinoma of the breast. Oncol Rep 19, 981–985 (2008).
Ishida, S. et al. Neuroendocrine carcinoma and mixed neuroendocrine-non-neuroendocrine neoplasm of the stomach: a clinicopathological and exome sequencing study. Hum Pathol 110, 1–10 (2021).
Luo, R. et al. Whole-exome sequencing identifies somatic mutations and intratumor heterogeneity in inflammatory breast cancer. NPJ breast cancer 7, 72 (2021).
Priedigkeit, N. et al. Acquired mutations and transcriptional remodeling in long-term estrogen-deprived locoregional breast cancer recurrences. Breast Cancer Res 23, 1–14 (2021).
Mathioudaki, A. et al. Targeted sequencing reveals the somatic mutation landscape in a Swedish breast cancer cohort. Sci Rep 10, 19304 (2020).
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumors. Nature 490, 61–70 (2012).
Marchiò, C. et al. The genetic landscape of breast carcinomas with neuroendocrine differentiation. J Pathol 241, 405–419 (2017).
Bao, L. et al. Coexisting genomic aberrations associated with lymph node metastasis in breast cancer. J Clin Invest 128, 2310–2324 (2018).
Hislop, R. G. et al. Karyotypic aberrations of chromosomes 16 and 17 are related to survival in patients with breast cancer. Br J Surg 89, 1581–1586 (2002).
Hansen, L. L., Yilmaz, M., Overgaard, J., Andersen, J. & Kruse, T. A. Allelic loss of 16q23.2-24.2 is an independent marker of good prognosis in primary breast cancer. Cancer Res 58, 2166–2169 (1998).
Singh, A. et al. Association of AGTR1 (A1166C) and ACE (I/D) Polymorphisms with Breast Cancer Risk in North Indian Population. Transl Oncol 11, 233–242 (2018).
Sarkisian, M. R. et al. MEKK4 signaling regulates filamin expression and neuronal migration. Neuron 52, 789–801 (2006).
Stevens, M. V., Parker, P., Vaillancourt, R. R. & Camenisch, T. D. MEKK4 regulates developmental EMT in the embryonic heart. Dev Dyn 235, 2761–2770 (2006).
Chi, H., Sarkisian, M. R., Rakic, P. & Flavell, R. A. Loss of mitogen-activated protein kinase kinase kinase 4 (MEKK4) results in enhanced apoptosis and defective neural tube development. Proc Natl Acad Sci U S A 102, 3846–3851 (2005).
Halfter, U. M., Derbyshire, Z. E. & Vaillancourt, R. R. Interferon-gamma-dependent tyrosine phosphorylation of MEKK4 via Pyk2 is regulated by annexin II and SHP2 in keratinocytes. Biochem J 388, 17–28 (2005).
Abell, A. N. et al. MAP3K4/CBP-regulated H2B acetylation controls epithelial-mesenchymal transition in trophoblast stem cells. Cell stem cell 8, 525–537 (2011).
Shao, F., Wang, Z. & Wang, S. Identification of MYCN-Related Gene as a Potential Biomarker for Neuroblastoma Prognostic Model by Integrated Analysis and Quantitative Real-Time PCR. DNA Cell Biol 40, 332–347 (2021).
Priedigkeit, N. et al. Acquired mutations and transcriptional remodeling in long-term estrogen-deprived locoregional breast cancer recurrences. Breast Cancer Res 23, 1–14 (2021).
Leung, M. H. et al. A Splice Variant of NCOR2, BQ323636.1, Confers Chemoresistance in Breast Cancer by Altering the Activity of NRF2. Cancers 12, 1–23 (2020).
Nagtegaal, I. D. et al. The 2019 WHO classification of tumors of the digestive system. Histopathology 76, 182–188 (2020).
Simbolo, M. et al. Gene Expression Profiling of Pancreas Neuroendocrine Tumors with Different Ki67-Based Grades. Cancers 13, 1–13 (2021).
Arakelyan, J., Zohrabyan, D. & Philip, P. A. Molecular profile of pancreatic neuroendocrine neoplasms (PanNENs): Opportunities for personalized therapies. Cancer 127, 345–353 (2021).
Sun, L. et al. Chromosomal and molecular pathway alterations in the neuroendocrine carcinoma and adenocarcinoma components of gastric mixed neuroendocrine-nonneuroendocrine neoplasm. Mod Pathol 33, 2602–2613 (2020).
Aggarwal, R., Zhang, T., Small, E. J. & Armstrong, A. J. Neuroendocrine prostate cancer: subtypes, biology, and clinical outcomes. JNCCN 12, 719–726 (2014).
Pei, X. et al. The next generation sequencing of cancer-related genes in small cell neuroendocrine carcinoma of the cervix. Gynecol Oncol 161, 779–786 (2021).
Acknowledgements
The bioinformatics analysis and further analyses of the sequencing data were assisted by J.Z. and X.Z. from Novogene Bioinformatics Technology Company. L.Y. and F.L. provided graphics and language assistance. This study was funded by the National Natural Science Foundation of China (81172536) and Key Research and Development Project of Department of Science & Technology in Sichuan Province (2018SZ0138).
Author information
Authors and Affiliations
Contributions
B.W. and Y.W. designed the study. B.W. and H.B. managed and supervised the project. Y.W., X.K., X.Z., Q.J. and J.Y. prepared the samples. Y.W. and B.W. reviewed the slides and confirmed the pathological diagnosis. Y.W., H.B. and B.W. analyzed and interpreted the data. Y.W. wrote the manuscript, which was reviewed by all co-authors. All the authors participated in the writing of the paper and ultimately approved the version for submission and publication.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was reviewed and approved by the clinical ethics committee of West China Hospital, Sichuan University (2021-644). The study was performed in accordance with the approval of Biomedical Ethics Review Committee.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Subject ontology breast cancer, whole-exome sequencing (WES), neuroendocrine differentiation, gene, mutation.
Supplementary information
Rights and permissions
About this article
Cite this article
Wei, Y., Ke, X., Yu, J. et al. Clinical and genomic analyses of neuroendocrine neoplasms of the breast. Mod Pathol 35, 495–505 (2022). https://doi.org/10.1038/s41379-021-00965-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-021-00965-w
This article is cited by
-
MAP3K4 kinase action and dual role in cancer
Discover Oncology (2024)
-
Genetic and immunohistochemical profiling of small cell and large cell neuroendocrine carcinomas of the breast
Modern Pathology (2022)
-
Comprehensive analysis of clinicopathologic features and p53 mutation in neuroendocrine neoplasms of the breast: experience from a large academic center
Breast Cancer Research and Treatment (2022)