Abstract
Morphological, immunohistochemical, and molecular methods often need to be combined for accurate diagnosis and optimal clinical management of sarcomas. Here, we have developed, a new molecular diagnostic assay, for the detection of gene fusions in sarcomas. This targeted multiplexed next-generation sequencing (NGS)-based method utilizes ligation dependent reverse-transcriptase polymerase chain reaction (LD-RT-PCR-NGS) to detect oncogenic fusion transcripts involving 137 genes, leading to 139 gene fusions known to be recurrently rearranged in soft-tissue and bone tumors. 158 bone and soft-tissue tumors with previously identified fusion genes by fluorescent in situ hybridization (FISH) or RT-PCR were selected to test the specificity and the sensitivity of this assay. RNA were extracted from formalin-fixed paraffin-embedded (n = 143) or frozen (n = 15) material (specimen; n = 42 or core needle biopsies; n = 116). Tested tumors encompassed 23 major translocation-related sarcomas types, including Ewing and Ewing-like sarcomas, rhabdomyosarcomas, desmoplastic small round-cell tumors, clear-cell sarcomas, infantile fibrosarcomas, endometrial stromal sarcomas, epithelioid hemangioendotheliomas, alveolar soft-part sarcomas, biphenotypic sinonasal sarcomas, extraskeletal myxoid chondrosarcomas, myxoid/round-cell liposarcomas, dermatofibrosarcomas protuberans and solitary fibrous tumors. In-frame fusion transcripts were detected in 98.1% of cases (155/158). Gene fusion assay results correlated with conventional techniques (FISH and RT-PCR) in 155/158 tumors (98.1%). These data demonstrate that this assay is a rapid, robust, highly sensitive, and multiplexed targeted RNA sequencing assay for the detection of recurrent gene fusions on RNA extracted from routine clinical specimens of sarcomas (formalin-fixed paraffin-embedded or frozen). It facilitates the precise diagnosis and identification of tumors with potential targetable fusions. In addition, this assay can be easily customized to cover new fusions.
Similar content being viewed by others
Introduction
More than 117 and 58 different subtypes of soft-tissue tumors and bone tumors are respectively recognized in the latest 2020 WHO classification of bone and soft-tissue tumors1. They are classified either based on their morphology and histogenesis or/and based on their defining molecular alteration. Diagnosis of the histological subtypes is challenging owing to the significant number of different entities, their rareness and the considerable morphological heterogeneity. Gene fusions that arise from chromosomal rearrangements leading to translocations, insertions, inversions, or interstitial deletions, are involved in up to 30% of sarcomas2. To date, over 200 gene fusions have been reported in these tumors1,2,3,4,5, more than half being recurrent in a specific subtype. Therefore, their identification in routine diagnosis is shifting from reverse-transcriptase polymerase chain reaction (RT-PCR) and fluorescent in situ hybridization (FISH) to next-generation sequencing (NGS) techniques to cover the wide range of fusions. However, RNA sequencing is expensive, time consuming, and the results are highly dependent on the technique used and requires bioinformatic support for analysis.
We have recently developed a NGS based ligation-dependent reverse transcription (LD-RT-PCR-NGS) assay that proved to be highly specific and sensitive for the simultaneous detection of a very wide range of fusion genes in fresh but also formalin-fixed, paraffin-embedded tumors (over 200 tested). Its performance has so far been validated in hematologic neoplasms6 and solid tumors such as salivary gland tumors7 and lung adenocarcinomas8,9. However, the identification of gene fusions is routinely performed for the primary diagnosis of in bone and soft tissue tumors, making these tumors a prime field to apply our technique. Here, we report the performance of this assay in soft-tissue and bone tumors (benign and malignant tumors/sarcomas). We describe its application to the detection of the most frequent fusion genes in a retrospective cohort of 158 soft-tissue and bone tumors, and compared its performance to that of conventional methods (immunohistochemical analysis, FISH and RT-PCR).
Materials and methods
Patients and tumor samples
We tested a retrospective cohort of 158 soft-tissue and bone tumors associated with gene fusion that were sampled in Bergonié Institute (Bordeaux, France), UNICANCER centers (France) and Centre Henri Becquerel (Rouen, France). The samples were from 158 patients including both surgical and biopsy specimen between 2010 and 2019. All tumors were reviewed by an expert pathologist from the NETSARC+ pathology network (French Sarcoma Group, 10) to confirm the diagnosis according to the WHO classification of sarcomas at the time of diagnosis. Ethics approval from the appropriate committees was obtained. Tumors encompassed 23 of the major subtypes of soft-tissue and bone tumors known to be associated with gene fusion described in Table 1. Adequate material [formalin-fixed paraffin-embedded tissue (n = 143) or frozen tissue (n = 15)] was available for subsequent immunohistochemical analysis (IHC) and molecular studies (FISH or RT-PCR) in all cases. Tumor tissue included surgical specimens (n = 42) or core needle biopsies (n = 116). Tumoral cellularity was over 15% in all cases. All sarcomas were then analyzed by LD-RT-PCR-NGS between September 2018 and February 2021.
LD-RT-PCR-NGS
Synthesis of LD-RT-PCR-NGS probe mix
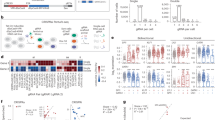
The LD-RT-PCR Fusion assay is a multiplex NGS-based method which has been developed at the Henri Becquerel Cancer Center. It relies on a specific validated custom panel of gene specific probes to detect gene fusions in formalin-fixed paraffin-embedded solid tumor tissue (detailed method described previously6,7,8,9,11). Its principle is illustrated in Fig. 1. Here, we designed a “solid tumors” probe mix, following a comprehensive review of the literature, to detect recurrent chromosomal rearrangements in solid tumors from several organs, including soft-tissue and bone tumors. It encompasses 333 oligonucleotides that target 184 genes, leading to 201 recurrent gene fusions. This mix include the “soft-tissue and bone tumors” mix, encompassing 267 oligonucleotides that were designed specifically to target the 137 most frequent fusion genes in soft-tissue and bone tumors transcripts (listed in Table 2, Fig. 2, Supplementary Table 1).
This assay is, based on an LD-RT-PCR-NGS amplification method adapted for the detection of multiple hybrid mRNAs linked by NGS technology. a Complementary DNA (cDNA) is incubated with oligonucleotide probes complementary to the starts and the ends of the exons fused on hybrid messenger RNAs (mRNA). If the fusion transcript is present, two probes hybridize side by side at the aberrant cDNA junction. After ligation a covalent link is created between the hybridized probes which allows their amplification by PCR using primers complementary to their tails. The two partners are finally identified using NGS technology. To detect the fusion, the left oligonucleotide probes harbor seven additional bases corresponding to a unique molecular identifier (UMI). The right oligonucleotide probes harbor eight additional bases corresponding to the molecular index that enables the patient to be identified. b The P5 and P7 sequences in the extremities enable sequencing by the Miseq Illumina. The P5 adaptor primer contained identical sequences of illumina P5 Sequence and 5′ universal adaptors, while the indexed P7 adaptor primer had reverse-complementary sequences of illumina P7 and 3′ universal adaptor as well as 8 bases index. rcl reverse complement.
These fusions cover 87.9% (58/66) of the histological entities known to be associated with gene fusion, including the main clinically relevant diagnosis (Table 2)1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71. Twenty-one gene fusions are associated with inflammatory myofibroblastic tumors (IMT), 11 with several subtypes of rhabdomyosarcomas (RMS), 10 with round cell sarcomas (RCS), 9 to NTRK rearranged neoplasms (NTRK-N), 9 with Ewing sarcomas (ES), 11 with low grade endometrial stroma sarcomas (ESSLG) and high grade endometrial stroma sarcomas (ESSHG), 9 with myoepitheliomas, 5 with biphenotypic sino-nasal sarcomas (BSNS), 6 with different subtypes of hemangioendotheliomas, 5 with extraskeletal myxoid chondrosarcomas (EMC), 7 with low grade fibromyxoid sarcomas/sclerosing epithelioid sarcomas (LGFMS/SEF), 4 with infantile fibrosarcomas (IF), and 3 with clear cell sarcomas (CCS). Some genes are implicated in several fusions such as EWSR1 (n = 24), ALK (n = 18), FUS (n = 11), NCOA1/2 (n = 9), NTRK1/3 (n = 8), PHF1 (n = 7), FN1 (n = 6), PAX3/7 (n = 6), NFATC1/2 (n = 4), CREB3L1/2 (n = 4), USP6 (n = 5), ROS1 (n = 3), TFCP2 (n = 2).
LD-RT-PCR-NGS probes are DNA oligonucleotides (Eurofins MWG Operon, Ebersberg, Germany) composed of a gene-specific region complementary to the fusion cDNA and fused to a 5′ or 3′ tail. All left probes had the same GTGCCAGCAAGATCCAATCTAGA tail (illu2) + UMI (unique molecular identifier), which is composed of 7 random oligonucleotides + the specific sequence of the left gene partner (5′ to 3′). All right probes had the same TCCAACCCTTAGGGAACCC tail (rc illu1) + the specific right gene partner (5′ to 3′). All 3′ probes were phosphorylated at their 5′ end to allow for the ligation reaction. All probes were pooled at a final concentration of 1 fmol/µl each in Tris 10 mM/EDTA 1 mM to create the LD-RT-PCR-NGS mix and can be distinguished according to the index contained in the P7 adaptor primers.
RNA extraction
One H&E-stained slide from formalin-fixed paraffin-embedded tissue (FFPE) was obtained for each sample and reviewed by an expert pathologist to evaluate tumoral cellularity, which was always greater than 15%. RNA was then isolated from 6 consecutive 10-µm unstained slides using the automated Maxwell®16 Research extraction system (Promega, Madison, WI, USA) and the Maxwell®16 FFPE Plus LEV RNA Purification Kit following the manufacturer’s instructions, and were stored at −80 °C.
RNA from frozen tissues was extracted according to the Chomczynzki method72 using 1.5 ml of Trizol-LS reagent (invitrogen) for 500 µl of cellular lysate. The solution was placed under moderate shaking for 30 minutes to 1 hour and RNA was extracted according to the manufacturer’s instructions (Qiagen). The RNA pellet was resuspended in 50 µl of RNase-free water and stored at –80 °C.
RNA concentration evaluation was performed by NanoDrop™ and varied from 2 to 6000 ng/µL. RNA concentration should be ≥2 ng/µL for LD-RT-PCR assay.
LD-RT-PCR-NGS reaction
The whole procedure was performed in a thermal cycler with a heated lid. RNA samples were first reverse-transcribed into cDNA using the SuperScript™VILO™cDNA Synthesis Kit (Invitrogen, Thermo Fisher Scientific). One microliter of a 5X Vilo reaction mix, 1 µl of H2O and 0.5 µl of the 10X superscript VILO reverse transcriptase were added to 2.5 µl of total RNA. These samples were then heated for 10 min at 25 °C, incubated for 60 min at 42 °C, incubated for 5 min at 85 °C and cooled to 4 °C. The obtained cDNA (5 µl) was then incubated with the LD-RT-PCR-NGS probe mix (1.5 µl). After the addition of 1.5 µl of SALSA-MLPA hybridization buffer (MRC Holland, Amsterdam, The Netherlands), the samples were heated for 2 min at 95 °C and incubated for 1 h at 60 °C to allow the annealing of the LD-RT-PCR-NGS probes. Thirty-two µl of a DNA ligase mix were then added to the reaction (3 µl SALSA-Ligase 65 Buffer A, 3 µl SALSA-Ligase Buffer B, 25 µl water, 1 µl SALSA-Ligase 65 (MRC-Holland)), and the samples were incubated for 15 min at 56 °C and heated at 98 °C for 5 min.
For this PCR amplification step, 5 µl of the ligation products were transferred to new tubes containing 18 µl of a PCR mix (12.5 µl Red’y’Star Mix (Eurogentec, Liege, Belgium), 5.5 µl water and 2 µL of primer PCR mix that contained P5I2+P7 barcodingI1 at 5 µM). Amplification was performed as follows: 6 min at 94 °C; 35 cycles (30 s at 94 °C, 30 s at 58 °C, 30 s at 72 °C); 4 min at 72 °C; and cooled to 4 °C.
Products were purified using the Agencourt AMPure XP kit (Beckman Coulter) according to the manufacturer’s instruction. The purified products were assayed with the Qubit DNA HS Assay kit (ThermoFisher Scientific). The different samples were pooled at a concentration of 4 nM to obtain approximately 100,000 reads for each case. They were next sequenced on an Illumina MiSeq instrument (Illumina, San Diego, CA). For each LD-RTPCR-NGS reaction, a control sample with known fusion transcript is included to validate the experiment.
The lower limit of transcript detection was 10 UMis. The lower limit of cellularity for transcript detection was evaluated as 5%. To validate the LD-RT-PCR assay, controls (one positive (EWSR1-FLI1) and one negative RNA) were systematically tested together with routine samples during the whole course of the study.
Data analysis
A customized software was developed in our institution as described previously8. RT-MIS analysis was used to align the sequencing data with the hybridizing sequences of all gene probes and to detect the gene fusion and calculate the number of sequence reads that contained this transcript. The approximate time from receipt of the sample in the laboratory to reporting of results was 2 full time days.
The unique molecular identifiers (UMIs) are an estimation of the number of ligations and the quantity of the fusion transcript expressed in the tumor. To interpret the results, we first need to evaluate if the fusion transcript identified has been associated in the literature with a sarcoma subtype.
-
If the fusion transcript has not been associated in the literature with a sarcoma subtype, the result is rendered as “no known fusion transcript detected in the tumor” (result negative in Table 1). It corresponds to a non-specific artifact fusion.
-
If the fusion transcript has been associated in the literature with a sarcoma subtype, we need to consider the number of UMIs. If the UMI number is <10, the result is negative. Conversely, if this number is ≥10, the fusion transcript is validated as a true fusion.
The background level of non-specific artifact fusion transcript is below 10 UMIs and is composed of fusions not described in the literature.
RT-PCR analysis
Reverse transcription of 5 µg RNA was performed in a total volume of 20 µl with 50 mM Tris-HCl, pH 8.3, 40 mM KCl, 5 mM MgCl2, 0.5% Tween, 0.5 mMdNTP mix, 10 mM dithiothreitol, specific reverse primer (FAM22 or reverse primer b2-microglobulin), 12 U RNAse inhibitor (Promega), and 10U Expand Reverse Transcriptase (Roche Diagnostics, Meylan, France). Samples were incubated at 42 °C for 1 h, then at 95 °C for 5 min.
PCR amplification was performed in duplicate using a 96-well plate (Applied Biosystems, Foster City, CA, USA) with a 50-µl final reaction mixture containing: 300 nM of each primer, 200, and 50 nM, respectively, of probe X and probe b2-microglobulin, and 0.25 U of Amperase UNG in a 2 qPCR Mastermix plus-low Rox (Eurogentec, Herstal, Belgium). The primers specific for each gene were designed to detect possible fusion transcripts (Supplementary Table 2). Thermal cycling conditions were 2 min at 50 °C for Amperase activation, 10 min at 95 1 C for Taq polymerase activation, then 50 cycles of two PCR steps consisting of 30 s at 95 °C, and 1 min at 60 °C. All reactions were performed in the ABI Prism 7500 Sequence Detection System (Applied Biosystems). RT-PCR analysis was performed in 115 tumors.
FISH analysis
FISH was performed on all cases using the following probes according to the diagnostic hypothesis: SS18 [Vysis LSI SS18 (18q11.2)] Dual Color Break Apart Rearrangement FISH Probe Kit, DDIT3 [Vysis LSI DDIT3 (12q13.3)] Dual Color Break Apart Rearrangement FISH Probe Kit, FUS [Zytovision LSI FUS (16p11.2)] Dual Color Break Apart Rearrangement FISH Probe Kit, TFE3 [ZytoLight® SPEC TFE3 (Xp11.23)] Dual Color Break Apart Rearrangement FISH Probe Kit, USP6 [ZytoLight® SPEC USP6 (17p13.2)] Dual Color Break Apart Rearrangement FISH Probe Kit, PDGFB [ZytoLight® SPEC PDGFB (22q13)] and homemade probes for WWTR1 and CAMTA1, Invitrogen (double fusion assay). Dual Color Break Apart Rearrangement FISH Probe Kit. FISH was performed using the ZytoLight SPEC Dual Color Break Apart Probe kit (CliniSciences). Cells were considered rearranged when at least one set of red/orange and green signals were two or more signal diameters apart, or when there was a red/orange single signal without a corresponding green signal in addition to fused and/or broken-apart signals, according to the manufacturer’s instructions. At least 100 tumor nuclei were analyzed, and a case was considered as positive when the number of neoplastic nuclei with a split signal or with a single 3′ signal was at least 15% of the observed neoplastic nuclei. FISH analysis was performed in 33 tumors.
Immunohistochemical analysis
We performed molecular IHC with antibodies highly associated with molecular alteration: STAT61 and pan-TRK73. Immunohistochemical staining was performed on 4‐μm‐thick formalin‐fixed paraffin‐embedded whole‐tissue sections with pan‐TRK rabbit monoclonal antibody, which reacted to a homologous region of TRK A, TRK B and TRK C near the C‐terminus (clone EPR17341; Abcam, Cambridge, MA, USA; dilution, 1:250 for 20 min. Immunostaining was performed on a Dako (Omnis) automated staining platform using the antigen retrieval method (EnVision FLEX Target Retrieval Solution, High pH (Agilent/Dako). The staining pattern, percentage of positive tumor cells and staining intensity were reviewed and recorded for all cases. Immunoreactivity was graded according to the percentage of cells with staining (0, <5%; 1+, 5–24%; 2+, 25–49%; 3+, 50–74%; or 4+, 75–100%) and the staining intensity (weak, moderate, or strong). The staining pattern (cytoplasmic, nuclear, and/or membranous) was also noted. Positive pan‐TRK staining was defined as immunoreactivity in at least 5% of cells.
Immunohistochemical staining was performed with STAT6 rabbit monoclonal antibody (abcam) clone [YE361] Abcam, Cambridge, MA, USA; dilution, 1:50 for 20 min. Immunostaining was performed on a Dako (Omnis) automated staining platform using the antigen retrieval method (EnVision FLEX Target Retrieval Solution, low pH (Agilent/Dako). IHC analysis was performed in 9 tumors (STAT6 in 8 and pan-TRK in one).
Results
Cohort description
Pathological data are summarized in Table 1. All diagnosis had been validated by expert sarcomas pathologists part of a national sarcoma pathology review network (RRePS, NETSARC + in France). The final cohort included 158 tumors, that covered 23 subtypes of bone and soft-tissue tumors including myxoid/round-cell liposarcomas (MLPS) (n = 17), ES (n = 14), synovial sarcomas (SS) (n = 13), desmoplastic small round-cell tumors (DSRCT) (n = 13), CCS (n = 8), gastrointestinal clear-cell sarcomas (CCS-GI) (n = 3), EMC (n = 9), IF (n = 9), alveolar rhabdomyosarcomas (ARMS) (n = 9), BSNS (n = 8), dermatofibrosarcomas protuberans (DFSP) (n = 8), round cell sarcomas (RCS) (n = 8), endometrial stromal sarcomas low grade (ESSLG) (n = 8), solitary fibrous tumors (SFT) (n = 8), alveolar soft-part sarcomas (ASPS) (n = 5), nodular fasciitis (NF) (n = 4), epithelioid hemangioendotheliomas (EHE) (n = 5), LGFMS/SEF (n = 3), perivascular epithelioid-cell neoplasms (PEComa) (n = 2), spindle-cell/epithelioid rhabdomyosarcomas (SERMS) (n = 1), endometrial stromal sarcomas high grade (ESSHG) (n = 1), mesenchymal chondrosarcomas (MC) (n = 1), and NTRK-rearranged spindle cell neoplasms (NTRK-N) (n = 1) (Table 1 and Fig. 4). The microscopic features of the main tumor types, FISH and IHC data are illustrated in Fig. 3.
Representative H&E pictures of tested sarcomas showing a ARMS with a PAX3-FOXO1 fusion (case no 1) showing proliferation of dyscohesive round cells with characteristic alveolar architecture (x200); b PEComa associated with a SFPQ-TFE3 fusion (case no 10). Tumor with sheet-like growth pattern. Spindle cells present clear cytoplasm and round or spindle nuclei (x200); c ASPS associated with ASPSCR1-TFE3 fusion (case no 12). Tumor shows large eosinophilic tumor cells with abundant cytoplasm arranged in compact nests (x200); d Representative picture of TFE3 immunohistochemical staining in an ASPS harboring a ASPSCR1-TFE3 fusion (case no 14). Tumor shows nuclear staining (x200); e Representative H&E pictures of CCS associated with EWSR1-ATF1 fusion (case no 25). Tumor shows plump spindle cells with pale cytoplasm arranged in solid pattern (x200); f DSRCT associated with EWSR1-WT1 fusion (case no 36). Tumor shows characteristic morphology with nests in desmoplastic stroma. Tumor cells are uniform, small, round with minimal nuclear pleomorphism. (x200); g EHE associated with WWTR1-CAMTA1 fusion (case no 58). Tumor is composed of nest and cords of epithelioid tumoral cells in sclerotic matrix (x200); h ES associated with EWSR1-FLI1 fusion (case no 71). Tumor is composed of small round cells arranged in solid pattern with occasional rosettes (x400). Insert shows nucleus with EWSR1 FISH fission; i Representative H&E of EMC with EWSR1-NR4A3 (case no 76) (x200). Spindle tumor cells with eosinophilic cytoplasm arranged in fascicles with myxoid matrix; j low grade ESS associated with JAZF1-SUZ12 fusion (case no 95). Tumor is composed of small uniform tumor cells with scant cytoplasm and oval nuclei arranged in sheets. Focal whorling of tumor cells is visible around small vessels (x200); k LGFMS/SEF associated with FUS-CREB3L2 fusion (case no 103). Bland spindle tumor cells are arranged in fascicles in myxoid matrix (x100); l BCOR-sarcoma associated with BCOR-CCNB3 fusion (case no 128), small round tumor cells are arranged in solid sheets separated by scant stroma (x200); m SFT associated with NAB2-STAT6 fusion (case no 137), hypercellular part of tumor is composed of spindle or ovoid cells admixed with hyalinized staghorn blood vessels (x200); n Representative picture of STAT6 immunohistochemical staining of SFT (case no 137). Tumor shows nuclear staining (x200); o Representative H&E picture of SS associated with SS18-SSX fusion (case no 145). This monophasic proliferation forms fascicles of monomorphic round or spindle cells (X200).
Results and failure rates of the LD-RT-PCR-NGS assay compared to conventional techniques in soft-tissue and bone tumors
LD-RT-PCR-NGS detected in-frame fusion transcripts in 155/158 tumors (98.1%) (Table 1):(FUS-DDIT3 (n = 17), EWSR1-FLI1 (n = 13), SS18-SSX (n = 13), EWSR1-WT1 (n = 13), PAX3-FOXO1 (n = 9), EWSR1-ATF1 (n = 9), EWSR1-NR4A3 (n = 9), ETV6-NTRK1 (n = 9), PAX3-MAML3 (n = 8), BCOR-CCNB3 (n = 8), JAZF1-SUZ12 (n = 8), COL1A1-PDGFB (n = 8), NAB2-STAT6 (n = 8), ASPRC1-TFE3 (n = 5), MYH9-USP6 (n = 4), WWTR1-CAMTA1 (n = 3), EWSR1-CREB1 (n = 2), SFPQ-TFE3 (n = 2), FUS-TFCP2 (n = 1), EWSR1-ERG (n = 1), YWHAE-NUTM2 (n = 1), FUS-CREB3L2 (n = 1), HEY1-NCOA2 (n = 1), TPM3-NTRK1 (n = 1), YAP1-TFE3 (n = 1) (Table 1). LD-RT-PCR-NGS failed to identify gene fusion in three tumors (1.9%). Conventional techniques were interpretable for all tumors (100%). Immunohistochemical analysis was performed in 9 tumors (STAT6 in 8 and pan-TRK in one), FISH analysis in 39 tumors, and RT-PCR in 114 tumors (Table 2).
Comparison of the results: LD-RT-PCR-NGS versus conventional techniques (immunohistochemistry, FISH, RT-PCR)
LD-RT-PCR-NGS (assay detected the known fusion gene in 155 of the 158 cases (98.1%, sensitivity) (Table 3). Among the three tumors negatives with the LD-RT-PCR-NGS assay, two cases were LGFMS showing a FUS gene rearrangement on the break-apart FISH, and the last case was an EHE showing a CAMTA1-WWTR1 gene fusion identified by double fusion FISH. None of these cases had been previously studied by PCR. LD-RT-PCR-NGS did not detect any unexpected fusion gene in any of samples, (100% specificity). Considering the NETSARC+ diagnosis, the concordance rate was 100% (150/150) for 21 tumor types, 80% (4/5) for diagnosing EHE and 33% (1/3) for diagnosing LGFMS/SEF.
Discussion
The identification of new recurrent gene fusions is continuously increasing (over 200 at this time) in soft-tissue and bone tumors2,3 and the identification of gene fusion has been shown to have a clinical impact in the management of sarcomas patients74. In this setting, molecular testing of sarcomas represent a daunting challenge for pathology laboratories to cover all possible fusions. The striking diversity involved in sarcomas has yielded many laboratories to switch to NGS-based screening techniques, putting aside techniques such as FISH and RT-PCR. In line with these assays, we describe a new LD-RT-PCR-NGS method that interrogates multiple genes at transcript level simultaneously and identifies fusion partners and exons participating in gene fusion. In addition, LD-RT-PCR-NGS is a simple assay which requires limited laboratory handling, such as reverse transcription, hybridization of the probes, ligation, and PCR amplification. The amplification products are purified and loaded on a next-generation sequencer, and the results are analyzed automatically using a dedicated bioinformatics pipeline. Therefore, the assay is easy to implement in a diagnostic workflow in the many molecular diagnostic laboratories that have already adopted NGS in their routine diagnostic workflow.
We have already implemented and clinically validated it to detect multiple gene fusions in leukemias6 and solid tumors such as salivary gland tumors7 and lung tumors8,9. Since the technique has not yet been used in soft-tissue and bone tumors, we adapted this assay to the detection of sarcoma specific rearrangements using panel using 267 primers designed to target 137 genes corresponding to 139 fusions known to be associated with soft-tissue and bone tumors. The present study assessed the capability of LD-RT-PCR fusion assay to detect fusion transcripts in bone and soft-tissue tumors in formalin-fixed paraffin embedded (FFPE) material in a routine diagnostic set-up and compared its performance with conventional techniques (FISH, RT-PCR, specific molecular immunohistochemistry). Although not all gene fusions described until now were part of our mix, the most frequently encountered were included, allowing our method to detect in-frame fusion transcripts in 155/158 tumors (98.1%) and to differentiate a wide range of soft-tissue and bone tumors (23 subtypes) in a single experiment.
The concordance rate between LD-RT-PCR-NGS and conventional techniques is very high: 155/158 tumors (98.1%). We analyzed frequent and rarer subtypes of translocation-related soft-tissue and bone tumors. LD-RT-PCR fusion assay confirmed the presence of the gene fusion in all but three cases (155/158). Regarding the three false-negative cases (cases (#57, #104, and #105), molecular confirmation had been performed by FISH on 5 µm FFPE tissue sections. Failure of LD-RT-PCR-NGS might be due to low RNA quality or quantity (all were archival FFPE samples), a complex translocation (insertion of intron between exon–exon junction), alternative breakpoints located in regions that are not covered by the gene-specific primers in our panel, an inadequate amount of ligation qualified as noise in the bioinformatics analyses, or to a tumor with gene fusion but no expression of the transcript. In addition, it is notorious that LGFMS (cases 104 and 105) are difficult to study by PCR75. Considering NETSARC+ diagnosis, the concordance rate was 100% (150/150) for 21 tumor types, 80% (4/5) for diagnosing EHE and 33% (1/3) for diagnosing LGFMS/SEF. Concerning the three false-negative cases (cases (#57, #104 and #105), RNA quality control (molecular monitoring of ABL gene expression levels - ipsogen®WT1 ProfileQuant, Qiagen®) showed low quality. As no remaining tumoral tissue (FFPE or frozen) was available, we could not perform others RNA sequencing, whole transcriptome sequencing or targeted RNA-seq, techniques that need of nucleic acid extraction to identify fusion transcripts.
The advantage of LD-RT-PCR-NGS over other technologies is its ability to multiplex and detect both common and potentially novel gene fusions, even starting from moderate quality samples (if two probes of the novel fusion are part of the assay). The unguided detection of many fusion genes in one test reduces the need to run multiple individual FISH or RT-PCR tests for single genes or single fusion variants. Therefore it shortens turnaround time and reduces labor costs8 while allowing the complete characterization of the fusion gene partners (exon–exon junction), which could potentially be of importance for clinical management. Furthermore, NGS can analyze specific nucleotide sequences, so there is no limit on the number of probes in one reaction. In addition, any newly described gene fusion can be subsequently added to the mix. Importantly, our data confirm that LD-RT-PCR-NGS is a robust technique that is easily applicable to FFPE samples, but also cytology slides and cell blocks, even with limited amounts of material are available9. Tumor samples successfully passing the sequencing analysis range from 95.1%9 to 100% (7 and the present study), i.e., more than the 92.9% rate by FISH analysis7.
The LD-RT-PCR fusion assay also has some drawbacks. First, the number of partner genes and break points is potentially very high and new partner genes are regularly identified, which would imply the regular update of our gene fusion panel. Second, the targeted NGS fusion gene assays are not always able to detect such fusion gene variants, a step that is highly dependent on knowledge of the junction on the fusion mRNA: i.e., if the fusion exon–exon junction requires an exon that is not in our panel, it will not be detected. The specific break point in each partner gene can be variable, resulting in a variety of exon–exon fusion combinations at the transcript level. Finally, the data analysis also requires specialized bioinformatics pipelines and sequence analysis techniques.
Although simple and robust, FISH testing can only detect one gene or one fusion gene at a time, requiring a sequential time-consuming strategy to screen tumor samples. RT-PCR can screen simultaneously several fusion genes in a single assay but the capacity of multiplexing is much limited with this technique compared to LD-RT-PCR.
Even if RNA-seq or Anchored multiplex PCR NGS assays are blinded techniques that can detect multiple and new gene fusions, fusion, splicing and exon skipping in a single assay compared to LD-RT-PCR that can only detect gene fusion and exon skipping. Nonetheless, they are more expensive, complicated with a longer turn-round time techniques. Interpretation is more difficult and requests expert bioinformatics pipeline. It requests higher amount of material, higher RNA quantity and quality.
RNA-seq or Anchored multiplex PCR NGS assays are blinded techniques that can detect multiple and new gene fusions, known gene fusion, splicing and exon skipping in a single assay compared to LD-RT-PCR that can only detect gene fusion and exon skipping9. Nonetheless, these techniques are more expensive, request longer turn-round time, higher amount of material, higher RNA quantity and quality, require higher sequencing depth compared to LD-RT-PCR. In addition, their interpretation is more difficult and request expert bioinformatics pipeline.
In addition, LD-RT-PCR fusion assay can easily identify chimeric protein kinases PKs that are the therapeutic target of many molecular-targeted drugs specific to translocations (ALK, ROS1, NTRK3, MET fusions and the tyrosine kinase receptor PDGFRB).
Given the challenge that the diagnosis of soft-tissue and bone tumors represents and the clinical impact of molecular methods in sarcoma diagnosis, our study supports that the LD-RT-PCR fusion assay is a sensitive and specific method to detect most gene fusions involved in soft-tissue and bone tumors that could be implemented in routine clinical settings. This multiplexed NGS-based LD-RT-PCR molecular approach could become an excellent screening method for the unguided detection of fusion transcripts and for classifying soft-tissue and bone tumors on FFPE material. The present findings show that it could become a first-line diagnostic test with the potential to replace or complement other more widely used molecular techniques.
References
WHO Classification of Tumors Editorial Board. Soft Tissue and Bone Tumors (International Agency for Research on Cancer, 2020).
Goldblum, J. R., Folpe, A. L, Weiss, S. W. &. Einzinger and Weiss’s Soft Tissue Tumors 7th edn, (Elsevier, 2020).
Mitelman F., Johansson B., Mertens F. eds Mitelman Database of Chromosome Aberrations and Gene Fusions in Cancer. Available from: URL: http://cgap.nci.nih.gov/Chromosomes/Mitelman (2018).
Mertens, F., Johansson, B., Fioretos, T. & Mitelman, F. The emerging complexity of gene fusions in cancer. Nat. Rev. Cancer 15, 371–381 (2015).
Mertens, F., Antonescu, C. R. & Mitelman, F. Gene fusions in soft tissue tumors: recurrent and overlapping pathogenetic themes. Genes Chromosom. Cancer 55, 291–310 (2016).
Ruminy, P. et al. Multiplexed targeted sequencing of recurrent fusion genes in acute leukaemia. Leukemia 30, 757–760 (2016).
Lanic, M.-D. et al. Ligation-dependent RT-PCR: a new specific and low-cost technique to detect gene fusion in salivary gland tumors. Abstract from USCAP 2020: Head and Neck Pathology (1235–1315). Mod. Pathol. 33, 1338–1408 (2020).
Piton, N. et al. Ligation-dependent RT-PCR: a new specific and low-cost technique to detect ALK, ROS, and RET rearrangements in lung adenocarcinoma. Lab. Investig. 98, 371–379 (2018).
Piton, N. et al. An improved assay for detection of theranostic gene translocations and MET exon 14 skipping in thoracic oncology. Lab. Investig. 101, 648–660 (2021).
NETSARC+, RRePS - Réseau de Référence en Pathologie des Sarcomes des tissus mous et des viscères. Available at: https://rreps.sarcomabcb.org/.
Angot, E. et al. A new simple low-cost multiplexed targeted sequencing assay to detect recurrent fusion genes in sarcomas. J. Clin. Oncol. 2015 USCAP Annual Meeting, Vol 33. N° 15_suppl (May 20 supplement) (2015).
WHO Classification of Tumors Editorial Board. Female Genital Tumors. Lyon (France): International Agency for Research on Cancer (2020).
WHO Classification of Tumors Editorial Board. Digestive System Tumors. Lyon (france): International Agency for Research on Cancer (2019).
Travis, W. D. et al. (eds) WHO Classification of Tumors of Lung, Pleura, Thymus and Heart 4th edn (IARC, 2015).
Elder, D. E., Massi, D., Scolyer, R. A. & Willemze, R. (eds) WHO Classification of Tumors of Skin Tumors 4th edn (IARC, 2018).
El-Naggar, A. K., Chan, J. K. C., Takata, T., Grandis, J. R. & Slootweg, P. J. The fourth edition of the head and neck World Health Organization blue book: editors’ perspectives. Hum. Pathol. 66, 10–12 (2017).
Siegfried, A. et al. RREB1-MKL2 fusion in biphenotypic “oropharyngeal” sarcoma: New entity or part of the spectrum of biphenotypic sinonasal sarcomas? Genes Chromosom. Cancer 57, 203–210 (2018).
Le Loarer, F. et al. Clinicopathologic and molecular features of a series of 41 biphenotypic sinonasal sarcomas expanding their molecular spectrum. Am. J. Surg. Pathol. 43, 747–754 (2019).
Arbajian, E. et al. A benign vascular tumor with a new fusion gene: EWSR1-NFATC1 in hemangioma of the bone. Am. J. Surg. Pathol. 37, 613–616 (2013).
Dickson, B. et al. Dermatofibrosarcoma protuberans with a novel COL6A3-PDGFD fusion gene and apparent predilection for breast. Genes Chromosom. Cancer 57, 437–445 (2018).
Dadone-Montaudié, B. et al. Alternative PDGFD rearrangements in dermatofibrosarcomas protuberans without PDGFB fusions. Mod. Pathol. 31, 1683–1693 (2018).
Dickson, B. C. et al. Ectomesenchymal chondromyxoid tumor: a neoplasm characterized by recurrent RREB1-MKL2 fusions. Am. J. Surg. Pathol. 42, 1297–1305 (2018).
Dickson, B. C., Swanson, D., Charames, G. S., Fletcher, C. D. & Hornick, J. L. Epithelioid fibrous histiocytoma: molecular characterization of ALK fusion partners in 23 cases. Mod. Pathol. 31, 753–762 (2018).
Suurmeijer, A. J. H. et al. Variant WWTR1 gene fusions in epithelioid hemangioendothelioma-A genetic subset associated with cardiac involvement. Genes Chromosom. Cancer 59, 389–395 (2020).
Sumegi, J. et al. A novel t(4;22)(q31;q12) produces an EWSR1-SMARCA5 fusion in extraskeletal Ewing sarcoma/primitive neuroectodermal tumor. Mod. Pathol. 24, 333–342 (2011).
Urbini, M. et al. HSPA8 as a novel fusion partner of NR4A3 in extraskeletal myxoid chondrosarcoma. Genes Chromosom. Cancer 56, 582–586 (2017).
Church, A. J. et al. Recurrent EML4-NTRK3 fusions in infantile fibrosarcoma and congenital mesoblastic nephroma suggest a revised testing strategy. Mod. Pathol. 31, 463–473 (2018).
Bender, J. et al. Refractory and metastatic infantile fibrosarcoma harboring LMNA-NTRK1 fusion shows complete and durable response to crizotinib. Cold Spring Harb Mol Case Stud. 5, a003376 (2019).
Huson, S. M. et al. Infantile fibrosarcoma with TPM3-NTRK1 fusion in a boy with Bloom syndrome. Fam. Cancer https://doi.org/10.1007/s10689-020-00221-1 (2020).
Flucke, U. et al. TFG-MET fusion in an infantile spindle cell sarcoma with neural features. Genes Chromosom. Cancer 56, 663–667 (2017).
Maruggi, M., Malicki, D. M., Levy, M. L. & Crawford, J. R. A novel KIF5B-ALK fusion in a child with an atypical central nervous system inflammatory myofibroblastic tumour. BMJ Case Rep. 21, 2018 (2018).
Tanaka, M. et al. Inflammatory myofibroblastic tumors of the lung carrying a chimeric A2M-ALK gene: report of 2 infantile cases and review of the differential diagnosis of infantile pulmonary lesions. Hum. Pathol. 66, 177–182 (2017).
Cools, J. et al. Identification of novel fusion partners of ALK, the anaplastic lymphoma kinase, in anaplastic large-cell lymphoma and inflammatory myofibroblastic tumor. Genes Chromosom. Cancer 34, 354–362 (2002).
Preobrazhenskaya, E. V. et al. Gene rearrangements in consecutive series of pediatric inflammatory myofibroblastic tumors. Pediatr. Blood Cancer 67, e28220 (2020).
Haimes, J. D. et al. Uterine inflammatory myofibroblastic tumors frequently harbor ALK fusions with IGFBP5 and THBS1. Am. J. Surg. Pathol. 41, 773–780 (2017).
Honda, K. et al. Durable response to the ALK inhibitor alectinib in inflammatory myofibroblastic tumor of the head and neck with a novel SQSTM1-ALK fusion: a case report. Investig. New Drugs 37, 791–795 (2019).
Lopez-Nunez, O. et al. Infantile inflammatory myofibroblastic tumors: clinicopathological and molecular characterization of 12 cases. Mod. Pathol. 33, 576–590 (2020).
Quade, B. J. et al. Fusion transcripts involving HMGA2 are not a common molecular mechanism in uterine leiomyomata with rearrangements in 12q15. Cancer Res. 63, 1351–1358 (2003).
Panagopoulos, I., Gorunova, L., Bjerkehagen, B. & Heim, S. Fusion of the genes EWSR1 and PBX3 in retroperitoneal leiomyoma with t(9;22)(q33;q12). PLoS ONE. 10, e0124288 (2015).
Dickson, B. C. et al. Novel EPC1 gene fusions in endometrial stromal sarcoma. Genes Chromosom. Cancer 57, 598–603 (2018).
Murshed, K. A. & Ammar, A. Hybrid sclerosing epithelioid fibrosarcoma/low-grade fibromyxoid sarcoma arising in the small intestine with distinct HEY1-NCOA2 gene fusion. Pathology 52, 607–610 (2020).
Mohamed, M., Fisher, C. & Thway, K. Low-grade fibromyxoid sarcoma: clinical, morphologic and genetic features. Ann. Diagn. Pathol. 28, 60–67 (2017).
Antonescu, C. R. et al. A distinct malignant epithelioid neoplasm with GLI1 gene rearrangements, frequent S100 protein expression, and metastatic potential: expanding the spectrum of pathologic entities with ACTB/MALAT1/PTCH1-GLI1 fusions. Am. J. Surg. Pathol. 42, 553–560 (2018).
Karanian, M. et al. SRF fusions other than with RELA expand the molecular definition of SRF-fused perivascular tumors. Am. J. Surg. Pathol. 44, 1725–1735 (2020).
Dahlén, A. et al. Activation of the GLI oncogene through fusion with the beta-actin gene (ACTB) in a group of distinctive pericytic neoplasms: pericytoma with t(7;12). Am. J. Pathol. 164, 1645–1653 (2004).
Hofvander, J. et al. RNA sequencing of sarcomas with simple karyotypes: identification and enrichment of fusion transcripts. Lab. Investig. 95, 603 (2015).
Sloan, E. A. et al. Intracranial mesenchymal tumor with FET-CREB fusion—a unifying diagnosis for the spectrum of intracranial myxoid mesenchymal tumors and angiomatoid fibrous histiocytoma-like neoplasms. Brain Pathol. 31, e12918 (2020).
White, M. D., McDowell, M. M., Pearce, T. M., Bukowinski, A. J. & Greene, S. Intracranial myxoid mesenchymal tumor with rare EWSR1-CREM translocation. Pediatr. Neurosurg. 54, 347–353 (2019).
Olson, N. et al. A novel case of an aggressive superficial spindle cell sarcoma in an adult resembling fibrosarcomatous dermatofibrosarcoma protuberans and harboring an EML4-NTRK3 fusion. J. Cutan. Pathol. 45, 933–939 (2018).
Yamazaki, F. et al. Novel NTRK3 fusions in fibrosarcomas of adults. Am J Surg Pathol 43, 523–530 (2019).
Tallegas, M. et al. Novel KHDRBS1-NTRK3 rearrangement in a congenital pediatric CD34-positive skin tumor: a case report. Virchows Arch. 474, 111–115 (2019).
Michal, M., Hájková, V., Skálová, A. & Michal, M. STRN-NTRK3-rearranged mesenchymal tumor of the uterus: expanding the morphologic spectrum of tumors with NTRK fusions. Am. J. Surg. Pathol. 43, 1152–1154 (2019).
Lu, Y. et al. Novel CTNNB1-USP6 fusion in intravascular fasciitis of the large vein identified by next-generation sequencing. Virchows Arch. 477, 455–459 (2020).
Bennett, J. A. et al. Uterine PEComas: a morphologic, immunohistochemical, and molecular analysis of 32 tumors. Am. J. Surg. Pathol. 42, 1370–1383 (2018).
Panagopoulos, I., Lobmaier, I., Gorunova, L. & Heim, S. Fusion of the genes WWTR1 and FOSB in pseudomyogenic hemangioendothelioma. Cancer Genomics Proteom. 16, 293–298 (2019).
Bridge, J. A., Sumegi, J., Royce, T., Baker, M. & Linos, K. A novel CLTC-FOSB gene fusion in pseudomyogenic hemangioendothelioma of bone. Genes Chromosomes Cancer 60, 38–42 (2021).
Sumegi, J. et al. Recurrent t(2;2) and t(2;8) translocations in rhabdomyosarcoma without the canonical PAX-FOXO1 fuse PAX3 to members of the nuclear receptor transcriptional coactivator family. Genes Chromosomes Cancer 49, 224–236 (2010).
Antonescu, C. R., Agaram, N. P., Sung, Y. S., Zhang, L. & Dickson, B. C. Undifferentiated round-cell sarcomas with novel SS18-POU5F1 fusions. Genes Chromosom. Cancer 59, 620–626 (2020).
Bissonnette, C. et al. An EWSR1-CREB3L1 fusion gene in extraskeletal undifferentiated round cell sarcoma expands the spectrum of genetic landscape in the “Ewing-Like” undifferentiated round cell sarcomas. Int. J. Surg. Pathol. 29, 109–116 (2021).
Antonescu, C. R., Agaram, N. P., Sung, Y. S., Zhang, L. & Dickson, B. C. Undifferentiated round cell sarcomas with novel SS18-POU5F1 fusions. Genes Chromosom. Cancer 59, 620–626 (2020).
Alholle, A. et al. Genetic analyses of undifferentiated small round cell sarcoma identifies a novel sarcoma subtype with a recurrent CRTC1-SS18 gene fusion. J. Pathol. 245, 186–196 (2018).
Tamura, R. et al. Novel MXD4-NUTM1 fusion transcript identified in primary ovarian undifferentiated small round cell sarcoma. Genes Chromosom. Cancer 57, 557–563 (2018).
Van Treeck, B. J. et al. NUTM1-rearranged colorectal sarcoma: a clinicopathologically and genetically distinctive malignant neoplasm with a poor prognosis. Mod. Pathol. 34, 1547–1557 (2021).
Pižem, J. et al. FUS-NFATC2 or EWSR1-NFATC2 fusions are present in a large proportion of simple bone cysts. Am. J. Surg. Pathol. 44, 1623–1634 (2020).
Debelenko, L. V., McGregor, L. M., Shivakumar, B. R., Dorfman, H. D. & Raimondi, S. C. A novel EWSR1-CREB3L1 fusion transcript in a case of small cell osteosarcoma. Genes Chromosom. Cancer 50, 1054–1062 (2011).
Panagopoulos, I., Gorunova, L., Viset, T. & Heim, S. Gene fusions AHRR-NCOA2, NCOA2-ETV4, ETV4-AHRR, P4HA2-TBCK, and TBCK-P4HA2 resulting from the translocations t(5;8;17)(p15;q13;q21) and t(4;5)(q24;q31) in a soft tissue angiofibroma. Oncol Rep. 36, 2455–2462 (2016).
Argani, P. et al. Novel MEIS1-NCOA2 gene fusions define a distinct primitive spindle cell sarcoma of the kidney. Am. J. Surg. Pathol. 42, 1562–1570 (2018).
Kao, Y. C. et al. Recurrent MEIS1-NCOA2/1 fusions in a subset of low-grade spindle cell sarcomas frequently involving the genitourinary and gynecologic tracts. Mod. Pathol. 34, 1203–1212 (2021).
Agaram, N. P. et al. A molecular study of synovial chondromatosis. Genes Chromosom. Cancer 59, 144–151 (2020).
Panagopoulos, I., Brandal, P., Gorunova, L., Bjerkehagen, B. & Heim, S. Novel CSF1-S100A10 fusion gene and CSF1 transcript identified by RNA sequencing in tenosynovial giant cell tumors. Int J. Oncol. 44, 1425–1432 (2014).
Hofvander, J. et al. Undifferentiated pleomorphic sarcomas with PRDM10 fusions have a distinct gene expression profile. J. Pathol. 249, 425–434 (2019).
Chomczynski, P. & Sacchi, N. Single step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 162, 156–159 (1987).
Brčić, I. et al. Broadening the spectrum of NTRK rearranged mesenchymal tumors and usefulness of pan-TRK immunohistochemistry for identification of NTRK fusions. Mod. Pathol. 34, 396–407 (2021).
Italiano, A. et al. Clinical effect of molecular methods in sarcoma diagnosis (GENSARC): a prospective, multicentre, observational study. Lancet Oncol. 17, 532–538 (2016).
Guillou, L. et al. Translocation-positive low-grade fibromyxoid sarcoma: clinicopathologic and molecular analysis of a series expanding the morphologic spectrum and suggesting potential relationship to sclerosing epithelioid fibrosarcoma: a study from the French Sarcoma Group. Am. J. Surg. Pathol. 31, 1387–1402 (2007).
Acknowledgements
The authors would like to thank Jean-Michel Coindre and the members of NETSARC+, Ray Cooke for copyediting the manuscript, and La Ligue contre le Cancer.
Funding information
None.
Author information
Authors and Affiliations
Contributions
All authors contributed to the writing and approval of this paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Ethics approval was obtained from the appropriate committees (NETSARC+ ).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Lanic, MD., Le Loarer, F., Rainville, V. et al. Detection of sarcoma fusions by a next-generation sequencing based–ligation-dependent multiplex RT-PCR assay. Mod Pathol 35, 649–663 (2022). https://doi.org/10.1038/s41379-021-00980-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-021-00980-x
This article is cited by
-
An unusual case of primary splenic soft part alveolar sarcoma: case report and review of the literature with emphasis on the spectrum of TFE3-associated neoplasms
Diagnostic Pathology (2024)
-
Insights for precision oncology from the integration of genomic and clinical data of 13,880 tumors from the 100,000 Genomes Cancer Programme
Nature Medicine (2024)