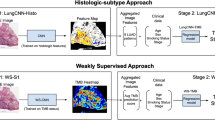
Abstract
Correctly diagnosing a rare childhood cancer such as sarcoma can be critical to assigning the correct treatment regimen. With a finite number of pathologists worldwide specializing in pediatric/young adult sarcoma histopathology, access to expert differential diagnosis early in case assessment is limited for many global regions. The lack of highly-trained sarcoma pathologists is especially pronounced in low to middle-income countries, where pathology expertise may be limited despite a similar rate of sarcoma incidence. To address this issue in part, we developed a deep learning convolutional neural network (CNN)-based differential diagnosis system to act as a pre-pathologist screening tool that quantifies diagnosis likelihood amongst trained soft-tissue sarcoma subtypes based on whole histopathology tissue slides. The CNN model is trained on a cohort of 424 centrally-reviewed histopathology tissue slides of alveolar rhabdomyosarcoma, embryonal rhabdomyosarcoma and clear-cell sarcoma tumors, all initially diagnosed at the originating institution and subsequently validated by central review. This CNN model was able to accurately classify the withheld testing cohort with resulting receiver operating characteristic (ROC) area under curve (AUC) values above 0.889 for all tested sarcoma subtypes. We subsequently used the CNN model to classify an externally-sourced cohort of human alveolar and embryonal rhabdomyosarcoma samples and a cohort of 318 histopathology tissue sections from genetically engineered mouse models of rhabdomyosarcoma. Finally, we investigated the overall robustness of the trained CNN model with respect to histopathological variations such as anaplasia, and classification outcomes on histopathology slides from untrained disease models. Overall positive results from our validation studies coupled with the limited worldwide availability of sarcoma pathology expertise suggests the potential of machine learning to assist local pathologists in quickly narrowing the differential diagnosis of sarcoma subtype in children, adolescents, and young adults.
Similar content being viewed by others
Introduction
Soft-tissue sarcomas (STS) are rare tumor types representing approximately 0.7% of all newly diagnosed cancers per year1. Despite the low relative incidence in adults, STS are comparatively overrepresented in childhood malignancies, accounting for 6–8% of all childhood cancers in patients under 15 years of age1,2. Pediatric sarcoma is generally subdivided into rhabdomyosarcoma (RMS) and non-RMS STS1,2. The plurality of new sarcoma diagnoses are RMS (approximately 350 per year)3 which has two major subtypes, embryonal RMS (eRMS) and alveolar (aRMS), and three rare but recognized subtypes, pleomorphic RMS (which is recognized in adults but may occur in children as well), botryoid RMS, and spindle cell/sclerosing RMS3. eRMS accounts for 70–80% of childhood RMS cases, aRMS accounts for 20–30%, and pleomorphic and spindle-cell account for a slim remaining fraction3. The remaining STS diagnoses (500–550 annually in the United States)1 are distributed amongst the 16+ NRSTS subtypes, including epithelioid sarcoma (EPS) and clear cell sarcoma (CCS)4. Between 2001 and 2010, the worldwide incidence of soft-tissue sarcoma in children aged 0–14 years was 18,323 cases (8.9 worldwide incidences per million person-years, age-standardized), and 7,011 in young persons aged 15–19 years (12.9 worldwide incidences per million person-years, age-standardized)5. Importantly, over 90% of potential childhood cancer patients each year live in low-income or middle-income countries6.
Common amongst all childhood STS malignancies is the importance of subtype and diagnosis on selecting chemotherapeutic and radiotherapeutic treatment strategy1,2,3. Correspondingly, correct diagnosis of sarcoma subtype is critical for clinical decision-making. However, due to the individual rarity of the numerous childhood STS tumors let alone the rarity in adult populations, histopathological diagnosis of childhood STS has historically demonstrated inconsistencies between individual pathologists and between medical institutions7,8,9. Among a subset of pediatric cancer classifications, STS cases demonstrate the highest misdiagnosis rate (72% incorrect diagnosis for STS, 52% incorrect diagnosis across all surveyed pediatric malignancies)9. Time to diagnosis was also elevated versus childhood leukemia (median 50.5 days to diagnosis for STS versus 18.5 days for leukemia)9. Furthermore, RMS misdiagnosis/disagreement in diagnosis rates can be as high as 40–50% for RMS7, highlighting the frequency of incorrect diagnosis in childhood STS.
Advances in the fields of molecular profiling and artificial intelligence have introduced new technologies to support diagnosis and prognosis of new cancer patients10,11,12,13. A recent approach focused on the use of hematoxylin and eosin (H&E) stained histopathology slides to predict prognosis in lung cancer patients10. This initial approach was subsequently expanded to include merged multi-omics datasets with pathology images to identify molecular mechanisms associated with pathological findings, thereby offering additional insight into patient prognosis11. A similar approach has been applied to a cohort of skin lesion images to visually diagnose benign versus malignant and low-risk versus high-risk melanoma through the use of smartphone-embedded cameras12. Recent work has leveraged existing convolutional neural network (CNN) deep-learning frameworks and models to apply transfer learning to the classification of lung cancer subtype and presence of a small set of biologically-relevant mutations based solely on histopathology image13. These efforts have demonstrated both the acute strength of machine learning models to assist in pathological diagnosis, and the focused application of novel image-based learning frameworks toward adult malignancies. The prevalence of adult neoplasms provides significantly larger histopathology cohorts which corresponds to a comparatively deeper body of clinical experts able to diagnose adult cancer patients.
Given the high rates of misdiagnosis in pediatric STS and the limited availability of deep expertise both within and outside economically-advanced nations, we applied the CNN-based histopathology diagnosis framework13 to a cohort of pediatric and young adult histopathology tissue slides originating from aRMS, eRMS and CCS cancer patients. Here, we report the results from CNN model generation, testing and validation for an externally-sourced RMS histopathology slide cohort. We also report robustness testing of the generated CNN model as well as murine histopathology applications arising from the trained CNN model and associated learning framework.
Methods and materials
Human digital pathology datasets
Digital histopathology tissue images from a total of 274 human subjects were included in this study (Supplementary Table 1). Of these, 146 were male, 98 were female, and 30 with unknown sex. Subjects ranged in age from 0 to 56 years of age with an average age less than 11 years (median of 6 years) for the subjects with age information available. These subjects included 83 patients diagnosed with aRMS, 101 with eRMS, 43 with CCS, one (1) with botryoid RMS, one (1) with CIC-rearranged sarcoma, 11 with epithelioid sarcoma, 14 with mixed type RMS, 14 with RMS not otherwise specified (NOS) and six (6) with spindle cell RMS (Supplementary Table 1). Institutional Review Boards at all collaborating institutions (Children’s Cancer Therapy Development Institute, University of California - Davis, The Royal Marsden Hospital, University Hospitals Leuven/Leuven Cancer Institute, Oregon Health & Science University (OHSU), Rhode Island Hospital) approved the sharing and use of histopathology images for this study. Children’s Oncology Group (COG) images were centrally reviewed. Rhabdomyosarcoma TMA images were previously centrally reviewed14. CCS images were from the European Organization for Research and Treatment of Cancer (EORTC) 90101 CREATE Phase II trial15 (TRAC project 384), and were centrally reviewed. In total, 424 digital histology slides were collected for use in this work. All histopathology images were provided with hematoxylin & eosin (H&E) staining performed at the originating institutions or were stained at the OHSU Histopathology Shared Resource core facility. Clinical metadata and institution of origin are provided in Supplementary Table 1 and experiment input sets are provided in Supplementary Table 2. Sample classical and variant STS images are provided in Fig. 1.
Murine digital pathology datasets
Mouse tumors from genetically engineered animals were developed and processed as previously described16,17,18. A total of 318 murine histopathology slides from 133 mice genetically engineered mouse models comprised the murine pathology dataset used in this study. The murine histopathology cohort consists of 206 aRMS tissue slides from 62 mice, 54 eRMS tissue slides from 13 mice, and 58 normal muscle tissue slides from 58 mice. All murine histopathology images were provided with hematoxylin & eosin (H&E) staining performed at OHSU and Legacy Emmanuel (Randall) Children’s hospital. Genotype data on the included murine cohort is provided in Supplementary Table 3. Mouse histopathology was reviewed by co-author Dr. Brian Rubin.
Cooperative group human tissue microarrays
Human rhabdomyosarcoma tissue arrays 3000_30_P8968_92, 3000_30_P8968_143, 3000_30_P8967_115, 3000_30_P8967_140, 3000_31_p1542_25, 3000_31_p1542_75 and 3000_31_P1542_50 were obtained from the Children’s Oncology Group.
Slide scanning
We used the ScanScope XT histopathology slide scanning platform (Leica, Wetzlar, Germany) running the manufacturer’s software (version 102.0.0.33) to digitize pathology images. We digitized images at 40x magnification to enable use of 5x and 20x resolution images.
Custom mouse-human tissue microarrays
Four samples of formalin-fixed paraffin embedded (FFPE) human RMS (1 x aRMS, 3 x eRMS) and forty-six (46) murine model sarcomas were used to construct a custom tissue microarray (TMA). The murine RMS cohort comprised representative developmental stages and genotypes, including early myoblast (origin), postnatal stem cell (origin), maturing myofiber (origin), Pax3:Foxo1-expressing, Trp53 wild type or mutated and Rb1 wildtype or mutated16,18,19. The TMA was stained with a standard hematoxylin & eosin (H&E) stain for histological verification. Co-author AM verified each tumor as non-rhabdomyosarcoma and rhabdomyosarcoma. The latter was further divided into aRMS, eRMS, pleomorphic RMS, and RMS NOS.
Clinical data and genotype information on TMA resources are provided in Supplementary Table 4.
Pre-processing of digital histopathology slide images
Machine Learning is a computational tool used to identify patterns in data. In the context of image pattern processing, convolutional neural network (CNN) models are among the most popular deep learning neural network architectures due to development of high-accuracy models across numerous image types and problem spaces. CNN uses a grid-like topology to classify images through multiple decision-making layers, and are frequently applied to both static and dynamic visual image data.
We used the open source DeepPATH software13 to create our CNN models. This software is written in the Python language and has a collection of software modules that enable all aspects of CNN model development, beginning with image pre-processing and including training, validation, and testing of CNN models for histopathology image classification. The adapted pre-processing consists of color normalization using the Vahadane methodology20 to adjust for color inconsistencies between immunohistochemical staining operations or slide scanning platforms12, segmenting the image into non-overlapping image sub-regions (termed tiles) of size 512 × 512 pixels at different image magnification (5x and 20x magnification), removing extraneous blank tiles generated during digitization and segmenting, and oversampling the resulting tile cohort data by rotating the tiles by 90-degree rotations and image mirroring.
We preprocessed the histopathology images by performing image segmentation, adjusting the magnification to 5x or 20x, performing color normalization, and eliminating blank tiles within the histopathology images. The complete pre-processing stage results in histopathology images color-normalized, magnified to proper resolution, segmented, and cleared of blank tiles.
Cross-validation, training, and testing of STS CNN model using digital histopathology slide images
We adapted the DeepPATH CNN software suite13 to create the human-trained CNN model for histopathological analysis. The adapted training/testing/validation model creation schematic is presented in Fig. 2A.
A Schematic of training, validation, and testing steps of CNN model construction using aRMS, eRMS, and CCS histopathology training set slides. B Testing using external aRMS, eRMS, and CCS histopathology slides. C CNN robustness testing using a discrete model-naïve STS histopathology cohort. D Robustness testing using anaplasia-presenting aRMS and eRMS histopathology slides. E Human-trained CNN model testing using murine-origin RMS histopathology images.
First, to address disparities in the size of the aRMS, eRMS, and CCS training tile set, we used image rotation and reflection of base images to generate new aRMS, eRMS, and CCS images until the resulting “oversampled” tile sets were approximately equivalent in size between all groups (Fig. 3A).
A Histopathology slide count (left) and resulting tile set (right) for the CNN model training, validation and testing. B Histopathology slide count and resulting tile set for the external validation aRMS, eRMS, and CCS slide cohort. C Histopathology slide count and resulting tile set for model-naïve STS robustness test. D Histopathology slide count and resulting tile set for anaplastic RMS robustness test. E Histopathology slide count and resulting tile set for the murine RMS cohort.
Next, to determine if the DeepPATH CNN software suite was appropriate for the task of STS diagnosis, we performed 5-fold cross validation testing, a process often used to determine if a computational model is appropriate for specific datasets or specific problems. We split the aRMS, eRMS, and CCS slide cohorts into five distinct sets and then trained five independent CNN models, where each model was trained on four of the five sets and the withheld fifth set was used to quantify model performance.
Having confirmed the applicability of the CNN framework to STS diagnostic applications, we created a new singular CNN model using the complete aRMS, eRMS and CCS image set. The singular CNN model was trained using a slide cohort comprising approximately 70% of the overall tile set for each disease type, validated for overall performance using a slide set consisting of 15% of the overall tile set to select the best performing CNN model, and subsequently tested using a slide set consisting of 15% of the overall tile set. Class likelihood scores were determined using a softmax output layer, with final CNN diagnosis class assigned by the maximum softmax score, i.e., the diagnosis class with the highest overall softmax score.
Training, testing, validation, and all subsequent experiments were performed in a Linux CentOS 7 environment run on a high-performance compute system using Broadwell Xeon processors and two NVIDIA QUADRO P6000 graphics processing units (GPUs). GPUs are often used in image processing due to the computational complexity required.
Diagnosis of external histopathology slide cohort
The process underlying disease specificity testing using non-trained sarcoma histopathology slides is described in Fig. 2B. In brief, histopathology images from institutions that did not provide images for model training (denoted external images) were pre-processed using the previously described pre-processing steps. No oversampling was performed in this experiment. Following pre-processing, the slides were classified by the human-trained CNN model.
Determining disease specificity in trained model
The process underlying disease specificity testing using non-trained sarcoma histopathology slides is described in Fig. 2C. In brief, histopathology images from multiple model-naïve STS subtypes (i.e., sarcoma subtypes on which the CNN model was not trained) were pre-processed using the previously described pre-processing steps. No oversampling was performed in this experiment. Following pre-processing, the slides were assigned random true classifications, and were then classified by the human-trained CNN model.
Anaplastic tissue testing with trained model
The process underlying classification of histopathology slides determined to have pathological hallmarks of anaplasia using the trained CNN model is described in Fig. 2D. In brief, RMS images pathologically determined to have present, diffuse, or focal anaplasia were pre-processed using the previously described pre-processing steps. No oversampling was performed in this experiment. Following pre-processing, the slides were classified by the human-trained CNN model.
Mouse prediction using human-trained model
The process underlying application of the human-trained CNN model to predict diagnosis in the murine RMS histopathology slide cohort is described in (Fig. 2E). In brief, murine-origin histopathology images were pre-processed using the previously described pre-processing steps. No oversampling was performed in this experiment. Following pre-processing, the slides were classified by the human-trained CNN model.
Training and testing of murine-origin RMS CNN model using digital histopathology slide images
To create the mouse-trained CNN model for histopathological analysis, we adapted the DeepPATH CNN software suite13 built atop the Inception v3 platform. The adapted training/testing/validation model creation schematic is identical to the schematic presented in Fig. 2A, using mouse-origin aRMS, eRMS, and mSKM images in place of human-origin tissues.
First, to address disparities in the size of the aRMS, eRMS, and normal muscle training tile set, we used image rotation and reflection to oversample the base images as previously described, until the resulting oversampled tile sets were approximately equivalent in size between all groups (Fig. 3E). Then, two CNN models were trained using the same 70% training/15% validation/15% testing method as described previously, with one model trained on murine RMS and murine skeletal muscle (mSKM) tissues, and the other model trained only on murine RMS tissues. Finally, CNN validation and testing were performed on both mouse-trained CNN models.
Python modules and other software tools used in CNN training/validation/testing
The following software packages were used for the described purposes.
-
0d_SortTiles.py (DeepPATH software suite): assignment of tiles or images into training/validation/testing sets while maintaining the balance of tiles or images within each group based on specified distributions, generally 70% training, 15% testing, and 15% validation.
-
nc_imagenet_eval.py (DeepPATH): image validation and testing/classification using the trained CNN model.
-
0b_tileLoop_deepzoom4.py (DeepPATH): preprocessing of histopathology images through image segmentation, magnification adjustment, and elimination of blank tiles.
-
Normalize_Vahadane.py (DeepPATH): color normalization of histopathology images through the Vahadane method using a selected baseline image.
-
0h_ROC_MultiOutput_BootStrap.py (DeepPATH): generation of ROC AUC scores and graphs.
-
Python Imaging Library (Pillow): enables rotation and reflection of image tiles for oversampling.
-
nSplit function (DeepPATH): segments slide or image datasets into distinct sets for cross-validation studies.
-
Inception v3 Deep Learning Model: base deep learning model used to develop new CNN models for STS differential diagnosis.
Python 3.6.8 was used for Python-based computational work.
Statistics
ROC AUC scores were performed in Python or by ROC analysis in GraphPad Prism (version 8). Class likelihood scores were statistically compared using the Kruskal–Wallis test with Dunnet’s multiple comparisons test in GraphPad Prism. ROC curve comparison was performed in Excel using the Hanley-McNeil method using the Kendall tau method for calculating the underlying ROC curve correlation.
Results
Training of clear cell sarcoma and rhabdomyosarcoma histopathology CNN model
The primary goal of this study was to develop a CNN deep learning model to serve as a differential diagnosis engine to support histopathological classification of human-origin RMS and CCS. The computational framework used to train, test and validate the RMS/CCS CNN model is described in (Fig. 2), and the input dataset for all experiments is provided in (Fig. 3). Note that the main research objective addressed in this study is differential diagnosis between the primary RMS subtypes, with CCS serving as an outgroup largely based on availability of histopathology images. The training/testing/validation set used to generate the set of CNN models consists of 119 aRMS histopathology samples (divided into 82 training, 19 validation, and 18 testing images), 103 eRMS histopathology samples (divided into 71 training, 17 validation, and 15 testing images), and 15 CCS samples (divided into 10 training, 2 validation, and 3 testing images). The set allotment was performed such that images and image sub-regions (termed tiles) used for model training are distinct from images and tiles used for model testing and validation. The cohort of RMS histopathology slides was collected from the Cancer Registry for Familial and Sporadic Tumors (CuRe-FAST) tumor bank at the Children’s Cancer Therapy Development Institute, University Hospitals Leuven/Leuven Cancer Institute (KU Leuven), Rhode Island Hospital, the Children’s Oncology Group (COG) Biorepository, the University of California at Davis, Oregon Health & Science University (OHSU), and the Royal Marsden Hospital (Supplementary Table 1). The comparatively small number of CCS histopathology slides used in this study is due to the paucity of available resources. All cases were locally and centrally reviewed (Supplementary Table 1) and then underwent secondary histology review by co-author sarcoma pathologist Sonja Chen.
With the slide cohorts curated and secondarily reviewed, we pre-processed each histopathology slide via color normalization20, segmenting the image into non-overlapping image tiles of size 512 × 512 pixels at different image magnification (5x and 20x magnification), removing extraneous blank tiles generated during digitization and segmenting, and oversampling the resulting tile cohort data by 90-degree increment tile rotations and image mirroring.
Using the 20x magnification tile set, oversampling was deemed necessary due to significant differences in tile count by disease type (baseline 22,419 aRMS tiles, 64,519 eRMS tiles, 30,766 CCS tiles), which would likely lead to training and validation biases. Oversampling resulted in 557,505 total tiles (179,352 aRMS from 119 histopathology slides from 61 patients, 177,183 eRMS from 103 histopathology slides from 63 patients, and 190,737 CCS from 15 histopathology slides from 13 patients, Fig. 3A, Supplementary Table 2).
With the tile cohort prepared, we next generated baseline CNN models for 5x and 20x magnification tile sets and determined that the 20x magnification dataset outperformed the 5x magnification model in initial comparative studies (Supplementary Table 5) and in 5-fold cross validation studies (Supplementary Fig. 1). Correspondingly, the remaining analyses in this study are performed exclusively with tiles generated from images at 20x magnification.
Next, to validate the applicability of CNN to the RMS/CCS histopathology dataset, we performed 5-fold cross-validation of the histopathology slide cohort as previously described (Fig. 3, Supplementary Table 6). The 5-fold trained CNN models performed well in one vs. all receiver operating characteristic (ROC) area under the curve analysis, with average AUC scores weighted by disease cohort size (Fig. 3A) and disease-level ROC AUC scores for each tile set (Fig. 3B–F) above the true positive rate (TPR)/true negative rate (TNR) threshold. Median and minimum micro-average AUC scores are 0.91 and 0.70 respectively, both above the 0.50 TPR/TNR threshold AUC. The positive cross validation results demonstrated the applicability of the CNN model to the problem of STS differential diagnosis.
We next trained a spectrum of CNN models using 70% of the image tiles from each disease type, reserving 15% for validation and 15% for testing. Using the validation AUC scores for each of the trained CNN models, we selected the top performing model for downstream use (Fig. 4, Supplementary Tables 7, 8). We tested the selected singular model using the withheld test data cohort at both the tile level (minimum AUC score 0.93, Fig. 4A) and slide level (minimum AUC score 0.89, Fig. 4B), affirming model performance during the 5-fold cross validation test. Additionally, the likelihood distribution scores of tiles classified during the testing step (Fig. 4C–H) demonstrate the ability of the trained CNN model to provide strong numerical support of the correct class, i.e., the high AUC scores are not an artifact of a low class membership threshold. Across all disease-types, tile-level and slide-level membership likelihood scores for the correct class are statistically significantly higher than incorrect classes (i.e., aRMS vs. others, eRMS vs. others, and CCS vs. others, p < 0.0001 all cases, Fig. 4C–H). Sample histopathology images and associated class membership heatmaps are presented in Fig. 5, and comparative pathologist-predicted and CNN-predicted tile images are presented in Fig. 6. Overall results are presented in Supplementary Tables 7, 8. Overall multi-class Matthews Correlation Coefficient for testing slides is 0.622.
A Tile-level ROC curves and AUC scores of CCS, aRMS, and eRMS tiles in slide set withheld from model training. B Slide-level ROC curves and AUC scores of CCS, aRMS, and eRMS slide set withheld from model training. C Likelihood distributions resulting from CNN classification of tiles from pathologist-diagnosed CCS slides, with the top (All) representing softmax likelihood scores of all testing images, middle (CNN Agreement) representing likelihood distributions when pathologist diagnosis and softmax-assigned CNN diagnosis match, and bottom (CNN Misdiagnosis) represents likelihood distributions when pathologist diagnosis and softmax-assigned CNN diagnosis are mismatched. Minimum and median correct softmax tile scores were 0.3416 and 0.9252, respectively. D As per (C), but for tiles from pathologist-diagnosed aRMS slides. Minimum and median correct softmax tile scores were 0.4026 and 0.9130, respectively. E As per (C), but for tiles from pathologist-diagnosed eRMS slides. Minimum and median correct softmax tile scores were 0.3316 and 0.9191, respectively. F As per (C), but for pathologist-diagnosed CCS slides. Minimum and median correct softmax tile scores were 0.7095 and 0.8717, respectively. G As per (C), but for pathologist-diagnosed aRMS slides. Minimum and median correct softmax tile scores were 0.5025 and 0.8654, respectively. H As per (C), but for pathologist-diagnosed eRMS slides. Minimum and median correct softmax tile scores were 0.7095 and 0.6175, respectively.
Color map scores are assigned to individual tiles within the slide images based on class membership likelihood values. Red is membership in class aRMS, blue is membership in class eRMS, black is membership in class CCS. A Original aRMS histopathology image. B Heatmap-coded aRMS histopathology image. C Original eRMS histopathology image. D Heatmap-coded eRMS histopathology image. E Original CCS histopathology image. F Heatmap-coded CCS histopathology image. Scale bars represent 5 mm.
Tile images generated from 512 × 512 pixel sub-regions of individual histopathology. Pathologist Diagnosed represents the disease subtype for the entire histopathology slide determined by pathologist analysis, while CNN Predicted Class represents the class predicted by the CNN model for the individual tile. Numbers in bottom right corner are likelihood scores assigned to the tile by the CNN model for the CNN predicted class. Scale bars = 50 µm.
Classification of external CCS, aRMS and eRMS slide cohort
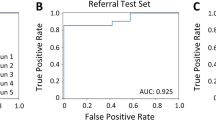
Having generated and validated an RMS/CCS CNN model, we next applied the singular CNN model to an external histopathology slide cohort (Fig. 3B, Supplementary Table 2), consisting of 30 CCS tissue slides, 14 aRMS tissue slides and 17 eRMS tissue slides. We pre-processed the external histopathology image cohort and then used the trained CNN model to predict class membership of the new slide cohort (Fig. 7). The trained CNN outperformed 0.50 TPR/TFR threshold across all disease cohorts, with AUC scores of 0.64, 0.89, and 0.61 for CCS, aRMS, and eRMS respectively. Class likelihood distributions for CCS, aRMS and eRMS cohorts show numerical support of predicted class membership (Fig. 7B–D). CCS correct class likelihood scores were significantly higher than aRMS likelihood scores (p < 0.0001) but were indistinguishable from eRMS likelihood scores (p > 0.05, Fig. 7B). However, both aRMS and eRMS correct class likelihood scores were significantly higher than other class likelihood scores (p < 0.05, Fig. 7C, D). Taken together, the CNN prediction results on external histopathology slides demonstrate the ability of the RMS/CCS trained CNN model to classify unseen histopathology specimens, although show the need for improvement with respect to CCS and eRMS disease classification. Overall results are presented in Supplementary Tables 9, 10. Overall multi-class Matthews Correlation Coefficient for external testing slides is 0.213.
A Slide-level ROC curves and AUC scores of externally-sourced CCS, aRMS, and eRMS slide set. B Likelihood distributions resulting from CNN classification of tiles from pathologist-diagnosed CCS slides, with the top (All) representing softmax likelihood scores of all testing images, middle (CNN Agreement) representing likelihood distributions when pathologist diagnosis and softmax-assigned CNN diagnosis match, and bottom (CNN Misdiagnosis) represents likelihood distributions when pathologist diagnosis and softmax-assigned CNN diagnosis are mismatched. Minimum and median correct softmax tile scores were 0.4328 and 0.6575, respectively. C As per (B), but for pathologist-diagnosed aRMS slides. Minimum and median correct softmax tile scores were 0.4083 and 0.5458, respectively. D As per (B), but for pathologist-diagnosed eRMS slides. Minimum and median correct softmax tile scores were 0.4588 and 0.6685, respectively.
Specificity of human STS-trained CNN on model-naïve STS disease image cohort
We trained the RMS/CCS CNN model used in this study based on availability of histopathology slides and resulting tile set size, which excluded a cohort of histopathology slides from multiple sarcoma subtypes: Botryoid RMS, sarcoma with a CIC-family gene fusion, epithelioid sarcoma, mixed-type RMS, RMS classified as not otherwise specified (NOS) and spindle-cell RMS (Fig. 3C, Supplementary Table 2). To investigate the specificity of the trained CNN model with respect to histopathology slides from untrained malignancies, we used the RMS/CCS CNN model to classify the non-RMS, non-CCS sarcoma slide cohort (Supplementary Fig. 2). The epithelioid sarcoma cohort did not specifically resemble either trained malignancy (p > 0.05 between CCS and eRMS cohort, Supplementary Fig. 2A), the mixed histology RMS cohort demonstrated a significant similarity to eRMS (p < 0.0001, eRMS vs. other, Supplementary Fig. 2B), the cohort of tissue samples with subtypes listed as NOS (not otherwise specified) tended towards being predicted as aRMS (p < 0.05, aRMS vs other, Supplementary Fig. 2C), and the remaining slide cohort comprised of various sarcoma types showed significant similarity to eRMS (p < 0.0001, eRMS vs. other, Supplementary Fig. 2D). Taken together, the varied prediction patterns for non-RMS, non-CCS sarcoma histopathology images suggest the currently generated CNN model performs well predicting classes for trained diseases but cannot currently discriminate against untrained disease images. Overall results are presented in Supplementary Table 11.
Predictive accuracy of human RMS-trained CNN in the presence of anaplasia
Anaplastic tumors, which are defined as having large, lobate, hyperchromatic nuclei (at least three times the size of neighboring nuclei) or atypical (obvious, multipolar) mitotic figures, are histopathologically-anomalous cases and may skew model training and classification toward atypical cases. Correspondingly, the singular CNN model used for the current study was trained only on histopathology slides determined to not have evidence of anaplasia. We subsequently classified the withheld anaplastic histopathology samples (4 aRMS tissue slides and 27 eRMS tissue slides, Fig. 3D, Supplementary Table 2) using the singular trained CNN model to determine classification rates of RMS tissues with evidence of anaplasia (Supplementary Fig. 3). Prediction of both anaplastic aRMS and eRMS outperformed the TPR/TNR 0.50 threshold with aRMS and eRMS ROC AUC scores at 0.73 and 0.74, respectively (Supplementary Fig. 3A), demonstrating that presence of anaplasia in aRMS or eRMS does not impair the accurate diagnose via the CNN model.
Additionally, predicted class likelihood scores demonstrate significantly higher likelihood scores for the correct classes versus incorrect classes for both anaplastic aRMS vs. others (p < 0.05 for both, Supplementary Fig. 3B) and anaplastic eRMS vs. others (p < 0.001 all cases, Supplementary Fig. 3C). Overall results are presented in Supplementary Tables 12, 13. Overall multi-class Matthews Correlation Coefficient for external testing slides is 0.287.
Classification of murine RMS histopathology images using human RMS-trained CNN model
We also applied the singular human RMS/CCS CNN model to a cohort of murine aRMS and eRMS histopathology slides originating from genetically engineered mouse models of RMS. We pre-processed the image cohort (Fig. 3E, Supplementary Table 2), then used the trained CNN model to predict human class membership of the murine slide cohort (Supplementary Fig. 4). The CNN model outperformed the TPR/TNR 0.50 threshold, with the ROC AUC for aRMS and eRMS at 0.72 and 0.67, respectively (Supplementary Fig. 4A), and the class membership likelihood for murine eRMS slides demonstrates strong membership in the correct class (Supplementary Fig. 4C). Notably, the class membership likelihood scores for murine aRMS are less clearly weighted towards aRMS (Supplementary Fig. 4B), suggesting that murine aRMS histopathology slides may loosely resemble human aRMS in the context of individual subregions, but more fully resemble human aRMS in the context of whole histopathology images. Murine aRMS correct class likelihood scores are not significantly higher than either eRMS or CCS likelihood scores (p > 0.05, Supplementary Fig. 4B). However, murine eRMS correct class likelihood scores were significantly higher than both aRMS and CCS class likelihood scores (p < 0.0001 both cases, Supplementary Fig. 4C). Overall results are presented in Supplementary Tables 14, 15. Overall multi-class Matthews Correlation Coefficient for external testing slides is 0.01.
Training of CNN model of murine rhabdomyosarcoma and normal murine muscle tissue histopathology images
Separate from the human RMS/CCS CNN model, we used a cohort of 318 murine histopathology slides comprised of 206 aRMS tissue slides from 62 mice, 54 eRMS tissue slides from 13 mice, and 58 normal muscle tissue slides from 58 mice to create a CNN model to classify murine RMS. Similar to the human created model, we normalized tissue slide color, segmented the slide images into 512 × 512 pixel tiles at a 20x magnification, removed extraneous blank tiles, and oversampled the murine aRMS, eRMS, and normal muscle tissue tile sets to produce 447,192 total tiles (157,595 aRMS, 136,584 eRMS, and 153,013 normal muscle, Fig. 3E, Supplementary Table 2). We subsequently trained and validated two murine RMS CNN models, one model generated with RMS histopathology images and normal murine skeletal muscle (mSKM) images (Supplementary Fig. 5A–D) and the other model generated only with murine RMS images (Supplementary Fig. 5E–G). We then ranked the generated models by using validation set prediction scores and tested the top-ranked CNN model using the withheld testing tile set using the previously described 70/15/15% tile set segmentation approach (Fig. 1A, using murine-origin instead of human-origin histopathology slides).
The AUC ROC scores for murine aRMS, eRMS, and normal tissue slides are 0.97, 0.98, and 1.0, respectively, for the withheld tile set classified by the top-ranked murine-trained CNN model. Additionally, class likelihood scores for the correct class versus incorrect class are statistically significant across all disease cohorts (p < 0.0001 for aRMS vs. others, eRMS vs. others, normal vs. others).
Notably, CNN model performance from the ROC AUC score perspective and likelihood score perspective was statistically indistinguishable between the models trained with and without normal mSKM tissue (p = 0.7514 and 0.8157 for murine aRMS and murine eRMS ROC curves, respectively21). However, the CNN model including mSKM outperformed the CNN model without mSKM based on MCC, F1 score, and Accuracy metrics (0.62, 0.87, 0.83 vs. 0.56, 0.87, 0.81, respectively), suggesting that inclusion of histologically normal tissue may improve practical classifier performance, but will require a larger image cohort and study to fully validate improvement in predictive accuracy. Sample histopathology images and associated class membership heatmaps are presented in Supplementary Fig. 6. Overall, the results demonstrate the strong predictive power of the murine histopathology-trained CNN model to predict murine disease classification. Tabular results are presented in Supplementary Tables 16, 17.
Discussion
The results from our study demonstrate the applicability of CNN models to numerical differential diagnoses for soft-tissue sarcomas. Sarcoma tissues are at elevated risk of misclassification by histopathology analysis due to the few cases seen by generalized anatomical pathologists in primary centers. By leveraging a cohort of 424 centrally-reviewed sarcoma histopathology slides (Supplementary Table 1), we then trained a CNN model to recognize three sarcoma subtypes (CCS, aRMS, eRMS) with high ROC AUC scores on an unseen dataset (0.96, 0.93, and 0.95 tile-level scores and 1.0, 0.90, and 0.98 slide-level scores for CCS, aRMS, and eRMS, respectively) and significant differences in class membership likelihood scores (p < 0.0001 for all correct class vs. incorrect class likelihood scores). Positive predictive results were further affirmed by accurately classifying additional aRMS and eRMS histopathology datasets (AUC ROC scores 0.98 and 0.84, respectively), and the trained CNN model demonstrated robustness with respect to atypical morphological features collectively known as anaplasia. The decrease in predictive accuracy can potentially be attributed to differences in histopathology slide scanning and slide quality between different institutions. Additionally, visualization of tile-level CNN scores identified a pattern of increased CNN classification variation around the borders of histopathology images, which stabilizes towards the image center of mass. This observation merits further investigation to improve classification accuracy. Nonetheless, the collective results of our study demonstrate the potential of trained CNN models to assist in differential diagnosis of rare cancers such as pediatric sarcoma.
Our study also highlighted limitations of the available datasets and robustness of the current approach. Due to the rarity of pediatric sarcoma amongst the population of cancer patients and the corresponding paucity of histopathology tissues, the current CNN model is trained on a smaller dataset than ideal for serving as a differential diagnosis clinical support tool. The current dataset is likely neither large enough nor diverse enough to truly reflect the biological and morphological diversity of the disease on study, highlighted in part by the reduced performance in the presence of anaplastic histopathology samples. Additionally, our investigation of prediction behavior on histopathology images from non-trained diseases highlights the limited robustness of our model in particular, and CNN modeling in general, on predictions made for unknown image classes. Robustness of the CNN model can be greatly improved by expanding training sets to include curated and centrally-reviewed pathology images from a broader spectrum of diseases with similar tissues- and organs-of-origin. Additional robustness testing will be applied to inclusion of non-cancerous histopathology images. In the context of rhabdomyosarcoma, normal muscle tissue is theoretically well-suited to identifying entrapped muscle fibers or surrounding normal muscle tissue, both of which are challenges in rhabdomyosarcoma diagnosis. However, the current CNN model is well-suited to the specific goal of differential diagnosis of subtypes of pediatric sarcoma.
We also note the difference in performance associated with different image magnification levels between the current study and a previous study on lung cancer13. While the original study demonstrated that models trained on 5x magnification images outperformed models trained on 20x magnification images, our results suggest the opposite to be true for childhood STS. We hypothesize the difference to be in part due to the small size of our current histopathology dataset, as 5x image magnification produces fewer tiles for model training and testing. We furthermore hypothesize the improved performance under 20x magnification may also be inherent to the challenging nature of childhood STS pathological diagnosis, which currently results in frequent misdiagnosis and disagreement in diagnosis in real-world use. Subsequently, the cytomorphological details, and specifically the nuclear characteristics, such as the nuclear shape and the chromatin texture, are better identified and evaluated through 20x magnification for diffuse infiltrating neoplasms such as various sarcoma subtypes. Correspondingly, the underlying reason behind the improvement under higher magnification may be histopathological in nature.
The course of the study also demonstrated the challenges of processing histopathology resources originating from multiple organizations prepared by different researchers/technicians. Variability in histopathology tissue quality, scanning technologies, and scanner operator protocol results in downstream impact in CNN model training and validation. Developing standardized scanning protocols or centralizing histopathology tissue imaging for digital applications would help in alleviating this challenge. Unfortunately, centralizing operations would also negatively impact the accessibility for institutions seeking to leverage the CNN differential diagnosis engine, leaving methodology standardization as the preferred path to improve quality of digital histopathology resources.
Overall, our study demonstrates that deep-learning CNN for RMS differential diagnosis is a promising additional tool to assist pathologists in their classification and subclassification of histopathology images of RMS tissue samples. As extensions of the current work, we will prioritize expanding the image cohort to include more histopathology images of currently trained sarcomas, as well as images of pediatric/young adult sarcomas we have not trained the CNN model to recognize. Additionally, we will broaden image sharing collaborations to include institutions of all sizes around the world, with the goal of enabling a web-accessible differential diagnosis engine (www.cc-tdi.org/machine-learning) to support histopathologic diagnosis for clinicians who may not have access to modern pathology analysis resources.
Materials availability
De-identified histopathology images are available online in.svs format through a material transfer agreement.
Change history
16 May 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41379-022-01098-4
References
Tinkle, C. L., Fernandez-Pineda, I., Sykes, A., Lu, Z., Hua, C.-H., Neel, M. D. et al. Nonrhabdomyosarcoma soft tissue sarcoma (NRSTS) in pediatric and young adult patients: Results from a prospective study using limited-margin radiotherapy. Cancer 123, 4419–4429 (2017).
Sangkhathat, S. Current management of pediatric soft tissue sarcomas. World J Clin Pediatr 4, 94–105 (2015).
Skapek, S. X., Ferrari, A., Gupta, A. A., Lupo, P. J., Butler, E., Shipley, J. et al. Rhabdomyosarcoma. Nat Rev Dis Primers 5, 1–1 (2019).
Spunt, S. L., Skapek, S. X. & Coffin, C. M. Pediatric nonrhabdomyosarcoma soft tissue sarcomas. Oncologist 13, 668–678 (2008).
Steliarova-Foucher, E., Colombet, M., Ries, L. A. G., Moreno, F., Dolya, A., Bray, F. et al. International incidence of childhood cancer, 2001–10: a population-based registry study. Lancet Oncol 18, 719-731 (2017).
Force, L. M., Abdollahpour, I., Advani, S. M., Agius, D., Ahmadian, E., Alahdab, F. et al. The global burden of childhood and adolescent cancer in 2017: an analysis of the Global Burden of Disease Study 2017. Lancet Oncol 20, 1211-1225 (2019).
Qualman, S. J., Coffin, C. M., Newton, W. A., Hojo, H., Triche, T. J., Parham, D. M. et al. Intergroup Rhabdomyosarcoma Study: update for pathologists. Pediatr Dev Pathol 1, 550–561 (1998).
Fouda, A., Mansour, A. & Al-Tonbary, Y. The many faces of Ewing sarcoma: Difficult to diagnose pediatric cases. Hematol Oncol Stem Cell Ther 2, 411–417 (2009).
Chen, J. & Mullen, C. A. Patterns of diagnosis and misdiagnosis in pediatric cancer and relationship to survival. J Pediatr Hematol Oncol 39, e110–e115 (2017).
Yu, K.-H., Zhang, C., Berry, G. J., Altman, R. B., Ré, C., Rubin, D. L. et al. Predicting non-small cell lung cancer prognosis by fully automated microscopic pathology image features. Nat Commun 7, 12474 (2016).
Yu, K. H., Berry, G. J., Rubin, D. L., Ré, C., Altman, R. B. & Snyder, M. Association of omics features with histopathology patterns in lung adenocarcinoma. Cell Syst 5, 620–627 (2017).
Esteva, A., Kuprel, B., Novoa, R. A., Ko, J., Swetter, S. M., Blau, H. M. et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542, 115–118 (2017).
Coudray, N., Ocampo, P. S., Sakellaropoulos, T., Narula, N., Snuderl, M., Fenyö, D. et al. Classification and mutation prediction from non–small cell lung cancer histopathology images using deep learning. Nat Med 24, 1559–1567 (2018).
Rudzinski, E. R., Anderson, J. R., Lyden, E. R., Bridge, J. A., Barr, F. G., Gastier-Foster, J. M. et al. Myogenin, AP2β, NOS-1, and HMGA2 are surrogate markers of fusion status in rhabdomyosarcoma: a report from the soft tissue sarcoma committee of the children’s oncology group. Am J Surg Pathol 38, 654–659 (2014).
Schöffski, P., Wozniak, A., Stacchiotti, S., Rutkowski, P., Blay, J. Y., Lindner, L. H. et al. Activity and safety of crizotinib in patients with advanced clear-cell sarcoma with MET alterations: European Organization for Research and Treatment of Cancer phase II trial 90101 CREATE. Ann Oncol 28, 3000–3008 (2017).
Abraham, J., Prajapati, S. I., Nishijo, K., Schaffer, B. S., Taniguchi, E., Kilcoyne, A. et al. Evasion mechanisms to Igf1r inhibition in rhabdomyosarcoma. Mol Cancer Ther 10, 697–707 (2011).
Nishijo, K., Chen, Q. R., Zhang, L., McCleish, A. T., Rodriguez, A., Cho, M. J. et al. Credentialing a preclinical mouse model of alveolar rhabdomyosarcoma. Cancer Res 69, 2902–2911 (2009).
Abraham, J., Nunez-Alvarez, Y., Hettmer, S., Carrio, E., Chen, H. I., Nishijo, K. et al. Lineage of origin in rhabdomyosarcoma informs pharmacological response. Genes Dev 28, 1578–1591 (2014).
Kikuchi, K., Taniguchi, E., Chen, H. H., Svalina, M. N., Abraham, J., Huang, E. T. et al. Rb1 loss modifies but does not initiate alveolar rhabdomyosarcoma. Skelet Muscle 3, 27 (2013).
Vahadane, A., Peng, T., Sethi, A., Albarqouni, S., Wang, L., Baust, M. et al. Structure-preserving color normalization and sparse stain separation for histological images. IEEE Trans Med Imaging 35, 1962–1971 (2016).
Hanley, J. A. & McNeil, B. J. A method of comparing the areas under receiver operating characteristic curves derived from the same cases. Radiology 148, 839–843 (1983).
Acknowledgements
We are grateful to Dr. Gregory Jones for use of the NVIDIA GPUs used in this study. We thank Dr. Christopher Fletcher for critical comments on this study and its scope. We appreciate Dr. Atiya Mansoor for her assistance with mouse pathology image analysis. Akira Nakagawara was invaluable in study concept discussions. We are appreciative of the following collaborators for providing histopathology images: Dr. Cheryl M. Coffin, Dr. Andrew E. Horvai, Dr. Zoltan Sápi, Dr. Francisco Jose Ortuño, Dr. Jalid Sehouli, Dr. Gottfried Lemperle, Dr. Hiroshi Moritake, Dr. Manoj P. Rai, Dr. Keisuke Yoshida, Dr. Markus Eckstein, Dr. Seiko Diane Yamada, Dr. Jessica G. Labadie, Dr. John Lennon Silva Cunha, Dr. Hernán Javier Cutuli, Dr. Shintaro Sugita, Dr. Bernardo Lopez-Andrade, Dr. Nadir Paksoy, Dr. Dalia Y. Ibrahim, and Dr. Güler Yavas.
Funding
This work was supported by the Brighter Days for Childhood Cancer Foundation, Graef Family Foundation, Golf Fights Cancer, Lovenheim Family Fund, Nevonen Family, Sara’s Cure Foundation, Team Megan Bugg and Braver Stronger Smarter Inc. Support for this project includes contributions in honor and/or memory of Io, Brad, Myke, Holly, Mark, Danny, Carson, Thomas, Kelsey, Xander, and Kevin.
Author information
Authors and Affiliations
Contributions
A.O.F., M.L., N.E.B., and C.K. performed writing, review, and revision of the paper, O.W., J.M.J., M.M.C., N.E., S.E.K., A.D.W., R.P., and J.L. performed preparation of resources and materials, A.O.F., M.L., C.Y.S., G.S., and N.E.B. performed statistical and computational modeling and analysis, Y.I., N.M., M.E., S.K., M.I., K.P., P.L., H.Z., A.A., E.J.D., D.G., A.W., P.S., J.L., P.L.H., R.L.J., and M.D. provided data and materials for analysis, A.W., P.S., C.-J.L., P.L.H., R.L.J., and M.D. provided technical and material support. All authors read and approved the final paper.
Corresponding authors
Ethics declarations
Competing interests
C.K. is a co-founder of Artisan Biopharma, a wholly-owned public benefit corporation of the Children’s Cancer Therapy Development Institute (cc-TDI). C.K. through cc-TDI also has research frameworks or collaborations with Roche Genentech, Eli Lilly, and Novartis. The remaining authors declare no conflicts of interest.
Ethics approval and consent to participate
All histopathology images were obtained and shared under IRB approval from their originating institutions.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: Due to a formatting error in an author name.
Supplementary information
Rights and permissions
About this article
Cite this article
Frankel, A.O., Lathara, M., Shaw, C.Y. et al. Machine learning for rhabdomyosarcoma histopathology. Mod Pathol 35, 1193–1203 (2022). https://doi.org/10.1038/s41379-022-01075-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-022-01075-x