Abstract
Conventional methods for the synthesis and analysis of chemical and biological materials often utilize homogeneous bulk environments or surface immobilization on a substrate. Homogeneous bulk environments, however, require large quantities of samples and reagents as well as significant effort to functionalize materials. Liquid–substrate interfaces can also pose problems because adsorption can hinder diffusion and reagent transport during bioanalyses, and sensitive materials, such as proteins, experience denaturation, or other types of deformation. Here, we describe the construction and use of droplet microenvironments created through a combination of surface modification, water-in-oil systems, and aqueous two-phase systems (ATPSs). This integration of an immobilization-free droplet microenvironment with liquid–liquid interfaces, material compartmentalization, directional reagent transport, and small volumes enables unique material functions. Specific examples include ATPS-assisted fabrication of functional microparticles for drug delivery, microscale determination of ATPS phase diagrams, dendritic self-assembly of semiconductive nanoparticles, multiplex immunoassays, and analysis of breast cancer cell migration.
Similar content being viewed by others
Microanalysis at a liquid–substrate interface and recapitulation of the cellular microenvironment
The scale of reactions affects reaction kinetics [1]. When a reactor is microscale, the rapid diffusion of molecules eliminates the need for an external device to expedite mixing and reaction [2, 3]. However, rapid diffusion at the microscale is a double-edged sword for biological systems. For example, thousands of biomolecules in living cells go through chemical and biological processes without a significant loss of material. The cells exploit rapid signaling reactions while simultaneously compartmentalizing the involved biomolecules within organelles to prevent unwanted cross-talk [4]. In other words, the cells can synthesize and functionalize biomaterials in a spatiotemporally controlled manner. In vitro studies, however, often use large-scale homogeneous reactions that may lack the unique effects of microscale compartmentalization [5]. By utilizing microscale compartmentalization, engineered cell-mimetic microenvironments can enhance in vitro material synthesis and bioanalysis.
Meanwhile, surface microanalysis techniques, including labeled and label-free methods, provide useful information on molecular and cellular interactions and often serve as the gold-standard for analytical platforms. Fluorescence-based detection techniques such as enzyme-linked immunosorbent assay (ELISA) [6] are commonly employed in biological studies. Label-free sensing techniques such as a quartz crystal microbalance [7,8,9] and surface plasmon resonance [10, 11] have also been developed to quantify biomolecular interactions with small quantities of samples. These techniques, however, often involve molecular immobilization on a substrate and require precise control of the molecular orientation and patterning to avoid misinterpretation of the data due to molecule-scale disorganization and signal cross-talk [12]. To overcome these challenges and properly interrogate physiological, biochemical, and cellular interactions, it would be useful to have the ability to reduce the reaction size, provide compartmentalization, and reconstitute appropriate microenvironments. This review describes unique droplet-based analytical platforms that recapitulate the cellular microenvironment using liquid–liquid separation and surface modification and enable novel material synthesis and bioanalysis.
Microanalysis in liquid microenvironments using microscale self-organization
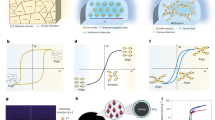
Microenvironments can be constructed through a combination of bottom-up and top-down technologies (Fig. 1a). The bottom-up strategy utilizes attraction, repulsion, and self-organization at the molecular scale to establish microenvironments [13, 14], while the top-down strategy establishes macro-scale guidance. This review focuses on droplet microenvironments prepared from aqueous two-phase systems (ATPS) and oil–water systems. ATPSs are generally comprised of aqueous solutions of two immiscible polymers that spontaneously phase separate above critical concentrations (Fig. 1b) [15]. ATPSs demonstrate distinct molecular partitioning and up to 1000-fold lower interfacial tension than conventional organic solvent–water-based extraction systems. These beneficial characteristics of ATPSs have been widely utilized for the separation of soft materials such as nucleic acids, proteins, and cells [16]. Recent studies demonstrated that ATPS-based microcompartments facilitate chemical reactions [17, 18] and site-specific reagent delivery to cells [19, 20]. Moreover, ATPSs have been exploited for the synthesis of drug carriers with novel properties that conventional methods fail to achieve [21, 22]. These reports highlight the ATPS-based microenvironment as a new platform for material synthesis and analysis that utilizes immiscible liquid–liquid interfaces.
Construction of microenvironments using bottom-up and top-down technologies. a Schematic illustration of a hybrid construction approach. b Photographs and binodal curves for aqueous two-phase systems (ATPSs) comprised of two immiscible polymers in aqueous solutions. Modified and used with permission from ref. 30
In this context, we developed a dextran (DEX: Mw = 500 kg mol−1)—pluronic F127 (F127: Mw = 12.6 kg mol−1) system and characterized the unique poly(lactide-co-glycolide) (PLGA) microparticles synthesized in the system (Fig. 2a) [23]. PLGA has been utilized to generate nano-to-microscale drug carrier particles that exhibit excellent biodegradability and biocompatibility [24, 25]. The microparticles form both single and multiple core domains depending on the PLGA concentration. When emulsified in a DEX—F127 mixture at room temperature, PLGA (Mw = 12 kg mol−1) formed core-shell microparticles with a DEX-rich shell and F127-rich core. The particle diameter ranged from 2 to 10 µm with an average size of ~7 µm. The PLGA microparticles encapsulated hydrophilic and hydrophobic model drugs, rhodamine B and coumarin-6, respectively. The loading efficacy could be adjusted based on the concentration of PLGA. The ability to alter the microparticle morphology and obtain high dual loading levels of hydrophilic and hydrophobic compounds are some of the advantages the ATPS-based method provides over conventional microparticle preparation protocols [26, 27].
Synthesis of PLGA microparticles in DEX–pluronic F127 ATPS. a Emulsification of PLGA (Mw = 12 kDa) in pluronic F127 (Mw = 13 kDa) and DEX (Mw = 500 kDa). b The generated PLGA microparticles (purple) with DEX-rich shells (red) and F127-rich cores (green) showing the different loading of Rhodamine B and Coumarin-6. c Temperature-dependent release of Rhodamine B at 4, 24, and 37 °C. Modified and used with permission from ref. 23
Another interesting feature of microparticles is that F127 containing a poly(propylene oxide) backbone changes from a hydrated state into a liquid crystalline state above the lower critical solution temperature (LCST). When the temperature increases above the LCST (30 °C), the F127 core collapses, and the PLGA microparticles release the encapsulated compounds, demonstrating the potential for temperature change-triggered drug delivery (Fig. 2c). The extent of the temperature-triggered drug release was higher for rhodamine B (20% release) than coumarin-6 (10% release). We speculate that the hydrophilic DEX-rich shell facilitates the release of the hydrophilic rhodamine B. The generated microparticles showed no significant cytotoxicity. We envision that this ATPS-based microparticle synthesis will be applicable to other drug carrier materials.
Microscale dehydration using soybean oil and FC-40
Oil and water are generally immiscible. Cells utilize the immiscibility and produce oil droplets in cytoplasm. The droplets associate with other intracellular organelles and presumably possess biological functions [28, 29]. Interestingly, some organic oils absorb small amounts of water into the oil phase due to the presence of acylglyceride groups [30]. For instance, soybean oil dissolves 0.3 (v/v)% water and has been adopted for microscale dehydration [31]. To take advantage of this unique property, we developed a microscale dehydration system that comprises two immiscible oils: Fluorinert™ FC-40 (FC-40) and soybean oil (Fig. 3a) [32]. Based on the density of FC-40 (1.8 g ml−1), soybean oil (0.92 g ml−1), and water (1.0 g ml−1), a suspended microdroplet (~0.1 µl) stays at the interface of the two oils in a plastic dish. FC-40, a fluorinated water-repellent oil, prevents droplet wetting on the dish surface, while the soybean oil continuously dehydrates the microdroplets at a slow rate (Fig. 3b). As a consequence, the solutes gradually condense as the microdroplet shrinks. A cross-section image taken by a conventional microscope provides the precise volume and solute concentration of the droplet at specific time points.
Microscale dehydration system that utilizes a soybean oil—FC-40 interface. a Solute condensation at the super hydrophobic interface of soybean oil and FC-40. b Kinetics of droplet shrinking at the oil interface. c Microscale determination of the phase diagram of 8k PEG—10k DEX ATPS. The inside panels show the phase morphology before and after phase separation. Critical concentrations determined by the dehydration method in the droplet (solid line) and the conventional dilution method in bulk solution (dashed line). Modified and used with permission from ref. 30
We demonstrated the microscale determination of phase diagrams for the following ATPSs: 500 k DEX—35k polyethylene glycol (PEG: Mw = 35 kg mol−1), 10k DEX—8 k PEG, 500 k DEX—105 k polyvinyl alcohol (PVA: Mw = 105 kg mol−1), 35 k PEG—105 k PVA, and 66 k bovine serum albumin (BSA: Mw = 66 kg mol−1)—35 k PEG systems. In this method, a microdroplet containing dilute phase-forming polymers was dispensed at the oil interface. Because of the dilute concentrations of the polymers, there is no phase separation at this stage. The microdroplet slowly shrinks as a function of time, and the microscale phase separation begins when the droplet reaches a critical concentration (Fig. 3c). The critical concentration was estimated based on a cross-sectional image of the microdroplet. This microscale method produced phase diagrams that agreed well with those generated using conventional macroscopic techniques but used significantly less material (a few micrograms versus grams of material). The reduction in materials by six orders of magnitude opens the door to test the phase formation of precious biomolecules only available in small quantities. Recent studies revealed that ATPSs comprised of biomolecules in vivo play an important role in biological events such as transcription [33, 34] and translation [35, 36]. Since such biomolecules are often precious and available only in limited quantities, microanalysis techniques shed light on the quantification of biological ATPSs.
Self-assembly of semiconductive nanoparticles in a dehydration system
Next, we hypothesized that a shrinking microenvironment could facilitate the self-assembly of inorganic nanoparticles. We subjected a dilute solution of semiconductive cadmium telluride (CdTe) nanoparticles [37] to a dehydration system and compared the nanoparticle self-assembly in a droplet and bulk solution (Fig. 4a) [38]. Interestingly, the nanoparticles in the droplet solution first aggregated at the droplet interface (day 1) and slowly evolved into branched dendrites with sizes from 50 µm to 1000 µm (day 3). In contrast, the bulk solution formed compact dendrites without a branched extension (day 3). A spectroscopic analysis revealed that the extended dendrites exhibited a distinct blueshift (λem = 515 nm) compared with that of the prepared CdTe solution (λem = 550 nm). In contrast, the compact dendrites showed a slight redshift (λem = 555 nm). The microscopic analysis showed that the fractal dimension of the extended dendrites in the droplet was higher (D = 1.9) than that of the compact dendrites (D = 1.7) in the bulk solution (Fig. 4b). The EDX analysis implied that the distinguishable optical property was probably due to different chemical compositions in the extended and compact dendrites.
Self-assembly of CdTe nanoparticles in a microscale dehydration system. a Fluorescent images of the compact and extended dendrites observed in bulk and in a droplet on day 3. The emission spectra of compact and extended dendrites collected at t = 0 and day 3. The excitation wavelength was 480 nm. b TEM observation of the compact and extended dendrites. c SEM observation of the extended dendrites in the presence of 500 mM NaCl (left) and light (right) on day 3. d SEM observation of a bulk mixture of soybean oil and CdTe solution on day 3. Modified from ref. 36
We also showed that the dendritic morphogenesis was sensitive to the droplet microenvironment (Fig. 4c). When the ionic strength increased, dendritic evolution failed to occur, and the compact dendrite remained, presumably due to reduced interparticle repulsion. Conversely, when we irradiated the microdroplet with light, the extended dendrites disassembled piece-by-piece. We assumed that the irradiation process removes the surface stabilizers on the nanoparticles and causes repulsions between the nanoparticles, resulting in the disassembly. To understand the underlying mechanism, we mixed soybean oil in the bulk solution and observed the dendritic extension (Fig. 4d). Inspired by the fatty acid transfer between cell organelles, we speculated that residual fatty acids are transferred to the droplet through the large surface-to-volume droplet interface and attach to the nanoparticle surface, prompting dendritic evolution by increasing the steric hindrance. The obtained semiconductive dendrites with a high fractal dimension and large surface area can be attractive light-harvesting materials. More broadly, the developed microscale dehydration system is a versatile platform for microscale self-assembly.
Microanalysis at surface-assisted liquid–liquid interfaces
Microfabrication through the generation of microchannels or micropatterning also assists in the construction of microenvironments [39, 40]. Since biological events usually occur under compliant conditions, soft lithography [41] using elastomers has commonly been employed. In particular, polydimethylsiloxane (PDMS), a transparent, gas-permeable, biocompatible polymer, prevails in soft lithography and enables both chemical and physical modifications to create customized PDMS platforms [42,43,44]. We adopted this technique to implement liquid–liquid patterning and create surface-assisted bioanalytical platforms.
First, we developed immobilization-free and cross-talk-free multiplexed ELISA using a 500k DEX—35 k PEG system [45,46,47]. DEX microdroplets (~1 µl) containing capture and detection antibodies were pre-dehydrated in each micro-basin in a customized microwell plate fabricated through microfabrication (Fig. 5a). The rehydration of the DEX phase facilitates reagent mixing through spontaneous convection once PEG solutions containing antigens are added. The DEX phase compartmentalizes the antibodies while allowing the small antigens to diffuse in. As a result, fluorescent signals are detected only in the dedicated DEX droplet containing the proper antibodies (Fig. 5b). In contrast, reagent cross-talk routinely occurs with conventional multiplexed ELISA in the absence of DEX-PEG ATPS because the antibodies are not compartmentalized in droplets. This technique successfully detected multiple cytokines from a standard dilution series and analyzed a macrophage cell culture supernatant in a single assay (Fig. 5c). These results not only illustrate practical biosensing, they also demonstrate the benefits of compartmentalized cellular bioreactions.
Multiplex ELISA in DEX-PEG ATPS. a Compartmentalization of the capture and detection antibodies in the DEX phase in the presence (left) and absence (right) of PEG. b Multiplex detection of cytokines. Antigens in a BSA solution showing no signal (left) and antigens in a PEG solution providing a distinct signal (right). Comparison of the signal-to-noise ratio between ATPS ELISA and conventional ELISA. c Detection of cytokine release from macrophage cell culture samples with/without LPS-induced activation. Modified and used with permission from ref. 44
Reconstruction of a cancer microenvironment using surface-assisted hydrogel patterning
In tumor microenvironments, cancer cells sense chemokine gradients and migrate along the gradients in a process known as chemotaxis [48]. In the example of the chemokine CXCL12 in breast cancer cell migration, CXCR4 (CXCL12 receptor)-expressing cells migrate toward CXCL12-expressing cells [49]. CXCL12 has several isoforms, CXCL12-α, -β, and -γ, with positive C-terminal amino acids that strongly bind to oppositely charged glycosaminoglycans (affinity strength: CXCL12-α < -β < -γ) [50]. Interestingly, the expression of CXCL12-β and -γ may be correlated to later stages of breast cancer [51]. Previous studies utilizing PDMS microchannels addressed the role of each isoform in 2D cell migration [52, 53]. However, the role of each isoform in cancer chemotaxis in the presence of 3D chemical gradients was elusive. We interrogated the role of CXCL12-α and -β in the migration of breast cancer cells using a droplet-patterning strategy [54, 55].
Non-adhesive polyacryamide (PAA)-coated PDMS surfaces were fabricated using a heterogeneous surface coating technique [56]. Cell-laden hydrogel droplets were guided to adjacent positions with a defined spacing between the droplets (Fig. 6a). The system enables the control of three parameters: the spacing between groups of patterned cells, gel composition, and cell type (Fig. 6b). The attachment assay showed significantly reduced cell attachment on the PAA-coated regions and ensured that the patterned cells migrated through the 3D hydrogel matrix rather than onto the 2D PDMS surface (Fig. 6c). Moreover, the 3D gel matrix allowed the formation and maintenance of 3D chemical gradients when a chemokine-binding component such as MatrigelTM was included. We patterned both CXCL12-secreting and CXCR4-expressing variants of human breast cancer cells (MDA-MB-231) in a collagen I – MatrigelTM matrix and measured the migration rate at day 1 (Fig. 7a). CXCL12-β showed a higher migration rate than CXCL12-α, while the rate of both CXCL12 isoforms was statistically distinct from that of the control. Notably, the migration rate of CXCL12-β was indistinguishable among non-secreting cells in the absence of Matrigel (Fig. 7b). The CXCL12-binding assay showed that CXCL12-β has a significantly stronger binding affinity for Matrigel than CXCL12-α and no differential binding affinity for collagen I (Fig. 7c). The results are consistent with the observed migration behavior. We inferred that heparan sulfate, a negatively charged glycosaminoglycan contained in MatrigelTM, boosts the binding of CXCL12-β through electrostatic interactions and facilitates the gradient formation and maintenance.
Gel–gel patterning on a PAA-coated PDMS surface for a cancer migration study. a PDMS mask-guided PAA-coating of a PDMS surface and hydrogel patterning. b Spatial patterning of human brain microvascular endothelial cell (HBMEC)-laden Matrigel (top) and NIH3T3 cell-laden collagen I gel with 200 µm and 400 µm gel-to-gel spacing, respectively. c Cell attachment assay of MDA-MB-231 cells on a cell culture plate, PAA-coated PDMS, and oxidized PAA-coated PDMS surfaces. Nuclei and cytosol were stained blue and green, respectively. The bar indicates individual pairwise differences (***p value <0.005). Modified and used with permission from ref. 52
Migration assay of CXCR4-expressing cells in 3D gel matrices. a Comparison of the migration in collagen gel—Matrigel™ on day 1 in the presence of CXCL12 isoforms. CXCR4+ cells (green) and CXCL12-secreting cells (red) on day 0 and day 1. b Comparison of the migration under various conditions. c Comparison of the binding affinity of CXCL12 isoforms for collagen gel and Matrigel™. Statistical demarcations above a bar indicate a pairwise comparison to all others in that group. The bar indicates individual pairwise differences (*p value <0.05, **p value <0.001, ***p value <0.005, and ****p value <0.0001). Modified with permission from ref. 52
A computational model revealed the differences in the gradient advancement between different CXCL12 isoforms (Fig. 8). The gradient of CXCL12-β is maintained in the relevant regions longer than that of CXCL12-α. The micropatterning capability of this procedure also allowed testing of the role of gel-to-gel spacing. When the spacing was doubled, no significant migration was directed for either CXCL12 isoform, further demonstrating the importance of the microenvironment in regulating cell behavior. These results not only corroborate prior 2D studies of the importance of CXCL12 isoforms, but also emphasize the critical role of 3D chemical gradient formation through chemokine–matrix interactions. That is, in the presence of MatrigelTM binding of CXCL12-β, there is no need for chemokine scavenger cells that express CXCR7 [57] to generate and maintain gradients sufficient for chemotaxis. Altogether, the results address important questions about physiological roles of CXCL12 isoform and matrix interactions as well as illustrate the versatility of our patterning strategy.
Gradient evolution of CXCL12 isoforms in the presence of Matrigel™. Concentrations of CXC12-α (top) and CXCL12-β (bottom) as a function of the distance from the patterned position every 8 h. Modified and used with permission from ref. 52
Summary and outlook
These developed platforms provide unique research opportunities ranging from microscale self-assembly to biomedical applications. We leveraged top-down and bottom-up technologies to design sophisticated liquid-based platforms. The liquid-based microenvironments allowed microscale characterizations of ATPS phase diagrams and syntheses of functional materials that conventional bulk conditions fail to achieve. The surface-assisted microenvironments revealed the underlying mechanism of liquid–liquid patterning and addressed the role of CXCL12 isoforms in cancer migration.
We envision future microscale analytical and reaction platforms moving further toward bio-inspired microenvironments. As we work to reconstruct more sophisticated and higher-order physiology in vitro, e.g., the immune system, and as diagnostics become more information-rich and multiplexed, we believe compartmentalized microenvironments enabled by novel methods and materials will be critical. This review described several strategies that combine top-down and bottom-up technologies to approach that goal.
References
Hill CG, Root TW. Introduction to chemical engineering kinetics and reactor design. NJ, USA: John Wiley & Sons; 2014.
Burns J, Ramshaw C. Development of a microreactor for chemical production. Chem Eng Res Des. 1999;77:206–11.
Squires TM, Quake SR. Microfluidics: fluid physics at the nanoliter scale. Rev Mod Phys. 2005;77:977.
Phillips R, Kondev J, Theriot J, Garcia H. Physical biology of the cell. 2nd ed. NY, USA: Garland Science; 2012.
Rieger E, Blankenburg J, Grune E, Wagner M, Landfester K, Wurm FR, Controlling the polymer microstructure in anionic polymerization by compartmentalization. Angew Chem Int Ed. 2018;57:2483–7.
Engvall E, Perlmann P. Enzyme-linked immunosorbent assay (ELISA) quantitative assay of immunoglobulin G. Immunochemistry. 1971;8:871–4.
Yoshimine H, Kojima T, Furusawa H, Okahata Y. Small mass-change detectable quartz crystal microbalance and its application to enzymatic one-base elongation on DNA. Anal Chem. 2011;83:8741–7.
Kojima T. Combined reflectometric interference spectroscopy and quartz crystal microbalance detect differential adsorption of lipid vesicles with different phase transition temperatures on SiO2, TiO2, and Au surfaces. Anal Chem. 2017;89:13596–602.
Kojima T. Surface modification enhanced reflection intensity of quartz crystal microbalance sensors upon molecular adsorption. Anal Sci. 2018;34:363–8.
Homola J, Yee SS, Gauglitz G. Surface plasmon resonance sensors. Sens Actuators B. 1999;54:3–15.
Homola J. Surface plasmon resonance sensors for detection of chemical and biological species. Chem Rev. 2008;108:462–93.
Lu B, Smyth MR, O’Kennedy R. Tutorial review. Oriented immobilization of antibodies and its applications in immunoassays and immunosensors. Analyst. 1996;121:29R–32R.
Whitesides GM, Boncheva M. Beyond molecules: self-assembly of mesoscopic and macroscopic components. Proc Natl Acad Sci USA. 2002;99:4769–74.
van Esch JH. Supramolecular chemistry: more than the sum of its parts. Nature. 2010;466:193–4.
Albertsson P-Å. Chromatography and partition of cells and cell fragments. Nature. 1956;177:771–4.
Hatti-Kaul R. Aqueous two-phase systems: methods and protocols. NY, USA: Humana Press; 2000.
Aumiller WM Jr, Davis BW, Keating CD. Phase separation as a possible means of nuclear compartmentalization. Int Rev Cell Mol Biol. 2013;307:109–49.
Kishimura A. Development of polyion complex vesicles (PICsomes) from block copolymers for biomedical applications. Polym J. 2013;45:892–7.
Tavana H, Jovic A, Mosadegh B, Lee Q, Liu X, Luker K, Luker G, Weiss S, Takayama S. Nanolitre liquid patterning in aqueous environments for spatially defined reagent delivery to mammalian cells. Nat Mater. 2009;8:736–41.
Armstrong JP, Olof SN, Jakimowicz MD, Hollander AP, Mann S, Davis SA, Miles MJ, Patil AJ, Perriman AW. Cell paintballing using optically targeted coacervate microdroplets. Chem Sci. 2015;6:6106–11.
Zhang Y, Wu F, Yuan W, Jin T. Polymersomes of asymmetric bilayer membrane formed by phase-guided assembly. J Control Release. 2010;147:413–9.
Ma S, Thiele J, Liu X, Bai Y, Abell C, Huck WT. Fabrication of microgel particles with complex shape via selective polymerization of aqueous two‐phase systems. Small. 2012;8:2356–60.
Yeredla N, Kojima T, Yang Y, Takayama S, Kanapathipillai M. Aqueous two phase system assisted self-assembled PLGA microparticles. Sci Rep. 2016;6:27736.
Makadia HK, Siegel SJ. Poly lactic-co-glycolic acid (PLGA) as biodegradable controlled drug delivery carrier. Polymers. 2011;3:1377–97.
Mundargi RC, Babu VR, Rangaswamy V, Patel P, Aminabhavi TM. Nano/micro technologies for delivering macromolecular therapeutics using poly (D, L-lactide-co-glycolide) and its derivatives. J Control Release. 2008;125:193–209.
Danhier F, Ansorena E, Silva JM, Coco R, Le Breton A, Préat V. PLGA-based nanoparticles: an overview of biomedical applications. J Control Release. 2012;161:505–22.
Español L, Larrea A, Andreu V, Mendoza G, Arruebo M, Sebastian V, Aurora-Prado MS, Kedor-Hackmann ER, Santoro MIR, Santamaria J. Dual encapsulation of hydrophobic and hydrophilic drugs in PLGA nanoparticles by a single-step method: drug delivery and cytotoxicity assays. RSC Adv. 2016;6:111060–9.
Gao Q, Goodman JM. The lipid droplet—a well-connected organelle. Front Cell Dev Biol. 2015;3:49.
Thiam AR, Farese RV Jr, Walther TC. The biophysics and cell biology of lipid droplets. Nat Rev Mol Cell Biol. 2013;14:775.
He M, Sun C, Chiu DT. Concentrating solutes and nanoparticles within individual aqueous microdroplets. Anal Chem. 2004;76:1222–7.
Bajpayee A, Edd JF, Chang A, Toner M. Concentration of glycerol in aqueous microdroplets by selective removal of water. Anal Chem. 2010;82:1288–91.
Kojima T, Takayama S. Microscale determination of aqueous two phase system binodals by droplet dehydration in oil. Anal Chem. 2013;85:5213–8.
Hnisz D, Shrinivas K, Young RA, Chakraborty AK, Sharp PA. A phase separation model for transcriptional control. Cell. 2017;169:13–23.
Mitrea DM, Kriwacki RW. Phase separation in biology; functional organization of a higher order. Cell Commun Signal. 2016;14:1.
Schisa JA, Pitt JN, Priess JR. Analysis of RNA associated with P granules in germ cells of C. elegans adults. Development. 2001;128:1287–98.
Schwartz JC, Wang X, Podell ER, Cech TR. RNA seeds higher-order assembly of FUS protein. Cell Rep. 2013;5:918–25.
Srivastava S, Santos A, Critchley K, Kim K-S, Podsiadlo P, Sun K, Lee J, Xu C, Lilly GD, Glotzer SC. Light-controlled self-assembly of semiconductor nanoparticles into twisted ribbons. Science. 2010;327:1355–9.
Kojima T, Hirai K, Zhou Y, Weerappuli P, Takayama S, Kotov NA. Nanoparticle assemblies into luminescent dendrites in shrinking microdroplets. Langmuir. 2016;32:12468–75.
Weibel DB, DiLuzio WR, Whitesides GM. Microfabrication meets microbiology. Nat Rev Microbiol. 2007;5:209–18.
Abe K, Suzuki K, Citterio D. Inkjet-printed microfluidic multianalyte chemical sensing paper. Anal Chem. 2008;80:6928–34.
Whitesides GM, Ostuni E, Takayama S, Jiang X, Ingber DE. Soft lithography in biology and biochemistry. Annu Rev Biomed Eng. 2001;3:335–73.
McDonald JC, Whitesides GM. Poly (dimethylsiloxane) as a material for fabricating microfluidic devices. Acc Chem Res. 2002;35:491–9.
Delamarche E, Juncker D, Schmid H. Microfluidics for processing surfaces and miniaturizing biological assays. Adv Mater. 2005;17:2911–33.
Sunkara V, Park D-K, Hwang H, Chantiwas R, Soper SA, Cho Y-K. Simple room temperature bonding of thermoplastics and poly (dimethylsiloxane). Lab Chip. 2011;11:962–5.
Frampton JP, White JB, Simon AB, Tsuei M, Paczesny S, Takayama S. Aqueous two-phase system patterning of detection antibody solutions for cross-reaction-free multiplex ELISA. Sci Rep. 2014;4:4878.
Eiden L, Yamanishi C, Takayama S, Dishinger JF. Aqueous two-phase system rehydration of antibody–polymer microarrays enables convenient compartmentalized multiplex immunoassays. Anal Chem. 2016;88:11328–34.
Simon AB, Frampton JP, Huang N-T, Kurabayashi K, Paczesny S, Takayama S. Aqueous two-phase systems enable multiplexing of homogeneous immunoassays. Technology. 2014;2:176–84.
Kay RR, Langridge P, Traynor D, Hoeller O. Changing directions in the study of chemotaxis. Nat Rev Mol Cell Biol. 2008;9:455–63.
Burger JA, Kipps TJ. CXCR4: a key receptor in the crosstalk between tumor cells and their microenvironment. Blood. 2006;107:1761–7.
Murphy JW, Cho Y, Sachpatzidis A, Fan C, Hodsdon ME, Lolis E. Structural and functional basis of CXCL12 (stromal cell-derived factor-1α) binding to heparin. J Biol Chem. 2007;282:10018–27.
Ray P, Stacer AC, Fenner J, Cavnar SP, Meguiar K, Brown M, Luker KE, Luker GD. CXCL12-γ in primary tumors drives breast cancer metastasis. Oncogene. 2015;34:2043–51.
Torisawa Y-S, Mosadegh B, Bersano-Begey T, Steele JM, Luker KE, Luker GD, Takayama S. Microfluidic platform for chemotaxis in gradients formed by CXCL12 source-sink cells. Integr Biol. 2010;2:680–6.
Cavnar S, Ray P, Moudgil P, Chang S, Luker K, Linderman J, Takayama S, Luker G. Microfluidic source-sink model reveals effects of biophysically distinct CXCL12 isoforms in breast cancer chemotaxis. Integr Biol. 2014;6:564–76.
Kojima T, Moraes C, Cavnar SP, Luker GD, Takayama S. Surface-templated hydrogel patterns prompt matrix-dependent migration of breast cancer cells towards chemokine-secreting cells. Acta Biomater. 2015;13:68–77.
Lesher-Perez SC, Kim G-A, Kuo C-H, Leung BM, Mong S, Kojima T, Moraes C, Thouless M, Luker G, Takayama S. Dispersible oxygen microsensors map oxygen gradients in three-dimensional cell cultures. Biomater Sci. 2017;5:2106–13.
Kojima T, Takayama S. Patchy surfaces stabilize dextran–polyethylene glycol aqueous two-phase system liquid patterns. Langmuir. 2013;29:5508–14.
Miao Z, Luker KE, Summers BC, Berahovich R, Bhojani MS, Rehemtulla A, Kleer CG, Essner JJ, Nasevicius A, Luker GD. CXCR7 (RDC1) promotes breast and lung tumor growth in vivo and is expressed on tumor-associated vasculature. Proc Natl Acad Sci USA. 2007;104:15735–40.
Acknowledgements
The authors thank Stephen Robinson for his proofreading of the manuscript.
Funding
This work was supported by the National Institute of Health (CA196018 and HL136141).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Kojima, T., Takayama, S. Microanalysis using surface modification and biphasic droplets. Polym J 50, 699–709 (2018). https://doi.org/10.1038/s41428-018-0050-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41428-018-0050-x