Abstract
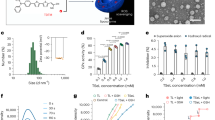
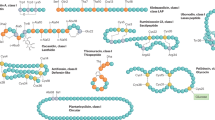
The prevalence of antimicrobial-resistant Cutibacterium acnes in acne patients has increased owing to inappropriate antimicrobial use. Commensal skin bacteria may play an important role in maintaining the balance of the skin microbiome by producing antimicrobial substances. Inhibition of Cu. acnes overgrowth can prevent the development and exacerbation of acne vulgaris. Here, we evaluated skin bacteria with anti-Cu. acnes activity. Growth inhibition activity against Cu. acnes was tested using 122 strains isolated from the skin of healthy volunteers and acne patients. Comparative genomic analysis of the bacterium with or without anti-Cu. acnes activity was conducted. The anti-Cu. acnes activity was confirmed by cloning an identified gene cluster and chemically synthesized peptides. Cu. avidum ATCC25577 and 89.7% of the Cu. avidum clinical isolates (26/29 strains) inhibited Cu. acnes growth. The growth inhibition activity was also found against other Cutibacterium, Lactiplantibacillus, and Corynebacterium species, but not against Staphylococcus species. The genome sequence of Cu. avidum showed a gene cluster encoding a novel bacteriocin named avidumicin. The precursor protein encoded by avdA undergoes post-translational modifications, supposedly becoming a circular bacteriocin. The anti-Cu. acnes activity of avidumicin was confirmed by Lactococcus lactis MG1363 carrying avdA. The C-terminal region of the avidumicin may be essential for anti-Cu. acnes activity. A commensal skin bacterium, Cu. avidum, producing avidumicin has anti-Cu. acnes activity. Therefore, avidumicin is a novel cyclic bacteriocin with a narrow antimicrobial spectrum. These findings suggest that Cu. avidum and avidumicin represent potential alternative agents in antimicrobial therapy for acne vulgaris.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Nakase K, Aoki S, Sei S, Fukumoto S, Horiuchi Y, Yasuda T, et al. Characterization of acne patients carrying clindamycin-resistant Cutibacterium acnes: A Japanese multicenter study. J Dermatol. 2020;47:863–9.
Hayashi N, Akamatsu H, Iwatsuki K, Shimada-Omori R, Kaminaka C, Kurokawa I, et al. Japanese dermatological association guidelines: Guidelines for the treatment of acne vulgaris 2017. J Dermatol. 2018;45:898–935.
Nakase K, Hayashi N, Akiyama Y, Aoki S, Noguchi N. Antimicrobial susceptibility and phylogenetic analysis of Propionibacterium acnes isolated from acne patients in Japan between 2013 and 2015. J Dermatol. 2017;44:1248–54.
Sardana K, Gupta T, Garg VK, Ghunawat S. Antibiotic resistance to Propionobacterium acnes: worldwide scenario, diagnosis and management. Expert Rev Anti Infect Ther. 2015;13:883–96.
Nakase K, Yoshida A, Saita H, Hayashi N, Nishijima S, Nakaminami H, et al. Relationship between quinolone use and resistance of Staphylococcus epidermidis in patients with acne vulgaris. J Dermatol. 2019;46:782–6.
Schommer NN, Gallo RL. Structure and function of the human skin microbiome. Trends Microbiol. 2013;21:660–8.
Chiller K, Selkin BA, Murakawa GJ. Skin microflora and bacterial infections of the skin. J Investig Dermatol Symp Proc. 2001;6:170–4.
O’Sullivan JN, Rea MC, O’Connor PM, Hill C, Ross RP. Human skin microbiota is a rich source of bacteriocin-producing staphylococci that kill human pathogens. FEMS Microbiol Ecol. 2019;95:fiy241.
Khalfallah G, Gartzen R, Moller M, Heine E, Lutticken R. A new approach to harness probiotics against common bacterial skin pathogens: Towards living antimicrobials. Probiot Antimicrob Proteins. 2021;13:1557–71.
Gallo RL, Murakami M, Ohtake T, Zaiou M. Biology and clinical relevance of naturally occurring antimicrobial peptides. J Allergy Clin Immunol. 2002;110:823–31.
Khelissa S, Chihib NE, Gharsallaoui A. Conditions of nisin production by Lactococcus lactis subsp. lactis and its main uses as a food preservative. Arch Microbiol. 2021;203:465–80.
Gotz F, Perconti S, Popella P, Werner R, Schlag M. Epidermin and gallidermin: Staphylococcal lantibiotics. Int J Med Microbiol. 2014;304:63–71.
Grice EA, Segre JA. The skin microbiome. Nat Rev Microbiol. 2011;9:244–53.
Rosenthal M, Goldberg D, Aiello A, Larson E, Foxman B. Skin microbiota: microbial community structure and its potential association with health and disease. Infect Genet Evol. 2011;11:839–48.
Nakase K, Nakaminami H, Takenaka Y, Hayashi N, Kawashima M, Noguchi N. Relationship between the severity of acne vulgaris and antimicrobial resistance of bacteria isolated from acne lesions in a hospital in Japan. J Med Microbiol. 2014;63:721–8.
Aoki S, Nakase K, Hayashi N, Noguchi N. Transconjugation of erm(X) conferring high-level resistance of clindamycin for Cutibacterium acnes. J Med Microbiol. 2019;68:26–30.
Nakase K, Okamoto Y, Aoki S, Noguchi N. Long-term administration of oral macrolides for acne treatment increases macrolide-resistant Propionibacterium acnes. J Dermatol. 2018;45:340–3.
Perez RH, Sugino H, Ishibashi N, Zendo T, Wilaipun P, Leelawatcharamas V, et al. Mutations near the cleavage site of enterocin NKR-5-3B prepeptide reveal new insights into its biosynthesis. Microbiology. 2017;163:431–41.
Grande Burgos MJ, Pulido RP, Del Carmen Lopez Aguayo M, Galvez A, Lucas R. The cyclic antibacterial peptide enterocin AS-48: Isolation, mode of action, and possible food applications. Int J Mol Sci. 2014;15:22706–27.
Wirawan RE, Swanson KM, Kleffmann T, Jack RW, Tagg JR. Uberolysin: a novel cyclic bacteriocin produced by Streptococcus uberis. Microbiology. 2007;153:1619–30.
Masuda Y, Ono H, Kitagawa H, Ito H, Mu F, Sawa N, et al. Identification and characterization of leucocyclicin Q, a novel cyclic bacteriocin produced by Leuconostoc mesenteroides TK41401. Appl Environ Microbiol. 2011;77:8164–70.
Martin-Visscher LA, van Belkum MJ, Garneau-Tsodikova S, Whittal RM, Zheng J, McMullen LM, et al. Isolation and characterization of carnocyclin A, a novel circular bacteriocin produced by Carnobacterium maltaromaticum UAL307. Appl Environ Microbiol. 2008;74:4756–63.
Sawa N, Zendo T, Kiyofuji J, Fujita K, Himeno K, Nakayama J, et al. Identification and characterization of lactocyclicin Q, a novel cyclic bacteriocin produced by Lactococcus sp. strain QU 12. Appl Environ Microbiol. 2009;75:1552–8.
Kemperman R, Kuipers A, Karsens H, Nauta A, Kuipers O, Kok J. Identification and characterization of two novel clostridial bacteriocins, circularin A and closticin 574. Appl Environ Microbiol. 2003;69:1589–97.
Kalmokoff ML, Cyr TD, Hefford MA, Whitford MF, Teather RM. Butyrivibriocin AR10, a new cyclic bacteriocin produced by the ruminal anaerobe Butyrivibrio fibrisolvens AR10: characterization of the gene and peptide. Can J Microbiol. 2003;49:763–73.
Kawai Y, Kemperman R, Kok J, Saito T. The circular bacteriocins gassericin A and circularin A. Curr Protein Pept Sci. 2004;5:393–8.
Scholz R, Vater J, Budiharjo A, Wang Z, He Y, Dietel K, et al. Amylocyclicin, a novel circular bacteriocin produced by Bacillus amyloliquefaciens FZB42. J Bacteriol. 2014;196:1842–52.
Potter A, Ceotto H, Coelho MLV, Guimaraes AJ, Bastos M. The gene cluster of aureocyclicin 4185: the first cyclic bacteriocin of Staphylococcus aureus. Microbiology. 2014;160:917–28.
Golneshin A, Gor MC, Williamson N, Vezina B, Van TTH, May BK, et al. Discovery and characterisation of circular bacteriocin plantacyclin B21AG from Lactiplantibacillus plantarum B21. Heliyon. 2020;6:e04715.
Koizumi J, Nakase K, Hayashi N, Nasu Y, Hirai Y, Nakaminami H. Multidrug-resistant Cutibacterium avidum isolated from patients with acne vulgaris and other infections. J Glob Antimicrob Resist. 2022;28:151–7.
van de Guchte M, van der Vossen JM, Kok J, Venema G. Construction of a lactococcal expression vector: expression of hen egg white lysozyme in Lactococcus lactis subsp. lactis. Appl Environ Microbiol. 1989;55:224–8.
Morgan-Kiss RM, Cronan JE. The Lactococcus lactis FabF fatty acid synthetic enzyme can functionally replace both the FabB and FabF proteins of Escherichia coli and the FabH protein of Lactococcus lactis. Arch Microbiol. 2008;190:427–37.
Linares DM, Kok J, Poolman B. Genome sequences of Lactococcus lactis MG1363 (revised) and NZ9000 and comparative physiological studies. J Bacteriol. 2010;192:5806–12.
Lo-Ten-Foe JR, de Smet AM, Diederen BM, Kluytmans JA, van Keulen PH. Comparative evaluation of the VITEK 2, disk diffusion, etest, broth microdilution, and agar dilution susceptibility testing methods for colistin in clinical isolates, including heteroresistant Enterobacter cloacae and Acinetobacter baumannii strains. Antimicrob Agents Chemother. 2007;51:3726–30.
Nakase K, Nakaminami H, Toda Y, Noguchi N. Determination of the mutant prevention concentration and the mutant selection window of topical antimicrobial agents against Propionibacterium acnes. Chemotherapy. 2017;62:94–9.
Maqueda M, Sanchez-Hidalgo M, Fernandez M, Montalban-Lopez M, Valdivia E, Martinez-Bueno M. Genetic features of circular bacteriocins produced by Gram-positive bacteria. FEMS Microbiol Rev. 2008;32:2–22.
Barnard E, Shi B, Kang D, Craft N, Li H. The balance of metagenomic elements shapes the skin microbiome in acne and health. Sci Rep. 2016;6:39491.
Da F, Joo HS, Cheung GYC, Villaruz AE, Rohde H, Luo X, et al. Phenol-soluble modulin toxins of Staphylococcus haemolyticus. Front Cell Infect Microbiol. 2017;7:206.
Rozalska M, Derczynska A, Maszewska A. Synergistic hemolysins of coagulase-negative staphylococci (CoNS). Acta Biochim Pol. 2015;62:757–64.
Luty RS, Fadil AG, Najm JM, Abduljabbar HH, Kashmar SAA. Uropathogens antibiotic susceptibility as an indicator for the empirical therapy used for urinary tract infections: a retrospective observational study. Iran J Microbiol. 2020;12:395–403.
McGinley KJ, Webster GF, Leyden JJ. Regional variations of cutaneous propionibacteria. Appl Environ Microbiol. 1978;35:62–6.
Huang F, Teng K, Liu Y, Cao Y, Wang T, Ma C, et al. Bacteriocins: Potential for Human Health. Oxid Med Cell Longev. 2021;2021:5518825.
Zainal Baharin NH, Khairil Mokhtar NF, Mohd Desa MN, Gopalsamy B, Mohd Zaki NN, Yuswan MH, et al. The characteristics and roles of antimicrobial peptides as potential treatment for antibiotic-resistant pathogens: a review. PeerJ. 2021;9:e12193.
Perez RH, Zendo T, Sonomoto K. Novel bacteriocins from lactic acid bacteria (LAB): various structures and applications. Micro Cell Fact. 2014;13:S3.
Ben-Shushan G, Zakin V, Gollop N. Two different propionicins produced by Propionibacterium thoenii P-127. Peptides. 2003;24:1733–40.
Ben Braiek O, Smaoui S. Enterococci: between emerging pathogens and potential probiotics. Biomed Res. Int. 2019;2019:5938210.
Cleveland J, Montville TJ, Nes IF, Chikindas ML. Bacteriocins: safe, natural antimicrobials for food preservation. Int J Food Microbiol. 2001;71:1–20.
Kong W, Lu T. Cloning and optimization of a nisin biosynthesis pathway for bacteriocin harvest. ACS Synth Biol. 2014;3:439–45.
Renye JA Jr, Somkuti GA. Nisin-induced expression of pediocin in dairy lactic acid bacteria. J Appl Microbiol. 2010;108:2142–51.
Perez RH, Ishibashi N, Inoue T, Himeno K, Masuda Y, Sawa N, et al. Functional analysis of genes involved in the biosynthesis of enterocin NKR-5-3B, a novel circular bacteriocin. J Bacteriol. 2016;198:291–300.
Conlan BF, Gillon AD, Craik DJ, Anderson MA. Circular proteins and mechanisms of cyclization. Biopolymers. 2010;94:573–83.
Zheng S, Sonomoto K. Diversified transporters and pathways for bacteriocin secretion in gram-positive bacteria. Appl Microbiol Biotechnol. 2018;102:4243–53.
Martinez-Bueno M, Valdivia E, Galvez A, Coyette J, Maqueda M. Analysis of the gene cluster involved in production and immunity of the peptide antibiotic AS-48 in Enterococcus faecalis. Mol Microbiol. 1998;27:347–58.
Borrero J, Kelly E, O’Connor PM, Kelleher P, Scully C, Cotter PD, et al. Plantaricyclin A, a novel circular bacteriocin produced by Lactobacillus plantarum NI326: purification, characterization, and heterologous production. Appl Environ Microbiol. 2018;84:e01801–17.
Major D, Flanzbaum L, Lussier L, Davies C, Caldo KMP, Acedo JZ. Transporter protein-guided genome mining for head-to-tail cyclized bacteriocins. Molecules. 2021;26:7218.
Gabrielsen C, Brede DA, Nes IF, Diep DB. Circular bacteriocins: biosynthesis and mode of action. Appl Environ Microbiol. 2014;80:6854–62.
Flaherty RA, Freed SD, Lee SW. The wide world of ribosomally encoded bacterial peptides. PLoS Pathog. 2014;10:e1004221.
Abengozar MA, Cebrian R, Saugar JM, Garate T, Valdivia E, Martinez-Bueno M, et al. Enterocin AS-48 as evidence for the use of bacteriocins as new leishmanicidal agents. Antimicrob Agents Chemother. 2017;61:e02288–16.
Acknowledgements
We are very grateful to Dr Jan Kok and Dr Harma Karsens for kindly providing Lactococcus lactis MG1363 and NZ9000 and their shuttle vectors pMG36c and pMG36e. We would like to thank Dr Konno for suggesting and trying isolation of avidumicin. We are very appreciative of the kindness suggestion from Dr Takeshi Zendo. We would like to thank Editage (www.editage.com) for English language editing.
Funding
This work was supported by JST, SPRING Grant Number JPMJSP2134, Japan (JK).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Koizumi, J., Nakase, K., Noguchi, N. et al. Avidumicin, a novel cyclic bacteriocin, produced by Cutibacterium avidum shows anti-Cutibacterium acnes activity. J Antibiot 76, 511–521 (2023). https://doi.org/10.1038/s41429-023-00635-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-023-00635-w