Abstract
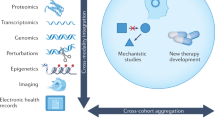
The application of bioinformatics has revolutionized the practice of medicine in the past 20 years. From early studies that uncovered subtypes of cancer to broad efforts spearheaded by the Cancer Genome Atlas initiative, the use of bioinformatics strategies to analyse high-dimensional data has provided unprecedented insights into the molecular basis of disease. In addition to the identification of disease subtypes — which enables risk stratification — informatics analysis has facilitated the identification of novel risk factors and drivers of disease, biomarkers of progression and treatment response, as well as possibilities for drug repurposing or repositioning; moreover, bioinformatics has guided research towards precision and personalized medicine. Implementation of specific computational approaches such as artificial intelligence, machine learning and molecular subtyping has yet to become widespread in urology clinical practice for reasons of cost, disruption of clinical workflow and need for prospective validation of informatics approaches in independent patient cohorts. Solving these challenges might accelerate routine integration of bioinformatics into clinical settings.
Key points
-
Retrospective classification of tumours using novel bioinformatics approaches has provided unprecedented insights into the molecular basis of urological cancers.
-
Molecular classifiers provide a useful adjunct to standard-of-care for the management of urological cancers, but prospective prediction of treatment response using molecular classifiers is not yet applied routinely.
-
Machine learning (ML) and artificial intelligence (AI) algorithms might circumvent the challenge of inter-observer variability in histopathology and could be incorporated into routine clinical practice.
-
Benign urology and functional urological disorders require improved patient phenotyping to fully realize the power of bioinformatics, as observed in oncology.
-
Incorporation of ML and AI approaches into routine clinical practice will require adherence to best practices including transparency in reporting results, and external validation in independent samples or patient cohorts before implementation.
Similar content being viewed by others
Introduction
Bioinformatics is an interdisciplinary field that encompasses a range of disciplines including biology, computer science, mathematics, information science and statistics, and enables the extraction and interpretation of meaning from high-dimensional datasets. Bioinformatics focuses on the analysis of biomolecules such as DNA, RNA and proteins, but bioinformatics principles have also been applied to the analysis of other high-content data types, such as physiological signals, histopathological and radiological images, and language. To harness the potential contained within these datasets, investigators are increasingly using artificial intelligence (AI) and machine learning (ML) algorithms to enable prognostication and prediction of outcomes and response to treatment, particularly in individuals with cancer, with the aim of obtaining potential clinical benefits.
Bioinformatics analysis is currently routinely used in both research and clinical care, which reflects the convergence of advances in methodology (including the development of massively parallel sequencing), analytical capabilities (including instrumentation), computation (such as enhanced processing power) and logistics (including cloud-based storage). Collectively, these improvements have enabled cost-effective data generation, data processing and high-dimensional analysis at a level that was unimaginable 20 years ago.
Bioinformatics has wide-ranging applications, including mapping mutations within a single tumour cell, identifying molecular subtypes within different types of cancer, predicting recurrence of kidney stones and detecting detrusor overactivity on urodynamics (UDS) tracings using ML. The knowledge obtained through bioinformatics analysis can be used, in principle, for surveillance, prognostication and treatment response prediction in a range of urological diseases with the promise of clinical benefit.
In this Review, we discuss the development and implementation of specific computational approaches, including ML algorithms, and the application of these approaches to discrete aspects of urology, both in clinical practice and in research. We consider the effect of bioinformatics analysis in genitourinary cancers, including the identification of molecular subtypes, prognostic and predictive biomarkers, and mechanisms of tumour evolution. We also discuss computational approaches to functional urological disorders. We conclude with consideration of best practices in the use of bioinformatics and ML applications in urology, and how these practices might be implemented to improve clinical care and research in our field. Owing to space constraints, this article mainly covers the use of bioinformatics associated with the molecular aspects of genitourinary diseases and only briefly discusses clinical applications of ML and AI in urology, which have been covered elsewhere1,2,3.
Primer on bioinformatics
The majority of bioinformatics studies focus on DNA and RNA expression data. However, a variety of high-content data types are amenable to bioinformatics analysis, including molecular data (DNA methylation profiles, mutational signatures, circulating tumour DNA, cell-free DNA, germline variants, proteomics, lipidomics and metabolomics), images (histological, radiological, MRI), physiological data (UDS profiles) and text. Preprocessing of data varies with input — which is not discussed in this article — but the analytical pipelines applied to relevant features that describe a particular type of data are similar.
Advances in the generation of high-content datasets (for example, using microarray technology or next-generation sequencing (NGS)) led to the necessity for appropriate methods to analyse these high-dimensional data. Bioinformatics is essential to all aspects of large-scale biological projects, from the processing of sequencing data to the generation of sophisticated predictive models. Traditional statistical methods are not suitable for large, complex datasets such as omics datasets, owing to the inherent variability and the computational burden of big data4, leading to the emergence of AI for analysis. AI is the simulation of human intelligence processes by computer systems. An AI algorithm is a process executed on a dataset to create a model, and an AI model might include multiple algorithms to solve a specific problem. In ML, a dataset is a collection of labelled or unlabelled data used to train a ML model. Datasets to be analysed include a variety of data points, described by a number of features, such as age, sex, tumour stage, nodal stage, tumour volume, time point, concentration of administered drug, gene expression level, protein intensity, metabolite abundance or urodynamic parameters. Features can be classified as categorical (for example, cancer versus no cancer, tumour grade, or true or false) or continuous (for example, fold change of gene expression or tumour volume). Datasets might also contain image-based features obtained from histological images, or text features derived from natural language processing (NLP) approaches. A dataset is typically split into two subsets: the training set and the test or testing set. Common problems that can occur in splitting the data into training and testing sets include imbalanced classes, small sample size, data leakage and non-representative sampling. Failure to appreciate these issues can result in learning rubbish (learning garbage) or biases in the data leading to a model that performs well on the training data but fails to generalize to new, unseen data.
An analysis that yields discrete categories or classes is termed a classification algorithm. Examples of classification algorithms include algorithms that predict whether a tumour is benign or malignant, or determine whether comments written by a patient convey a positive or negative sentiment5. Conversely, an analysis that yields a continuous value is termed a regression algorithm. Importantly, the use of the term ‘regression’ in AI is different from the use of this term in statistics. In AI, a regression algorithm is an algorithm used for prediction (for example, of an individual’s life expectancy or of the optimal dose of chemotherapy)6, whereas, in statistics, regression is often used to refer to both binary outcomes (such as logistic regression) and continuous outcomes (such as linear regression).
ML is a subset of AI, and is defined as the use of algorithms and statistical models to analyse data and infer patterns within existing information7. The simplest form of ML analysis enables the user to classify various entities based on biological features, and to group subjects into discrete clusters based on similarity. Depending on the type of algorithm applied, ML might be unsupervised, supervised or reinforced. Unsupervised learning algorithms are able to identify patterns in unlabelled data (for example, mRNA expression profiles used to define molecular subtypes) (Fig. 1). With the unsupervised algorithm (classifier), unknown data points (such as patients or cells) are not classified based on prior information, but based on properties inherent to the data themselves (such as histological data or gene expression). Unsupervised training algorithms are used primarily in pattern detection and descriptive modelling, and include hierarchical clustering, principal component analysis (PCA), k-means clustering, independent component analysis, singular value decomposition and association rules8,9. Conversely, supervised learning algorithms are applied to data in which labels have already been assigned (Fig. 1) (for example, gene expression profiles from tumours versus non-tumour specimens). With this kind of data, an unknown sample could be analysed using a supervised learning approach and assigned to one group or the other based on the gene expression profile. Supervised algorithms are used to build predictive models and include k-nearest neighbours, naive Bayes, decision trees, linear regression, support vector machine (SVM) and neural networks (NNs)10. Another type of supervised learning relevant to cancer is anomaly detection, which is an important application of ML in urology, and has the potential to improve patient outcomes, increase efficiency and accuracy of diagnoses, and reduce health-care costs. Anomalies are defined as data points that deviate substantially from the norm or expected pattern, and detecting these anomalies can help identify potential health issues or abnormalities11. In urology, anomaly detection can be used for various purposes, such as identifying kidney stones from CT scans or ultrasonography images of the kidneys, even when the stones are small or difficult to detect by manual inspection; detecting bladder cancer, with ML algorithms used to identify biomarkers or abnormal cells in urine or biopsy samples, which indicate the presence of bladder cancer; monitoring patient health, as ML algorithms can be used to analyse patient data, such as vital signs, laboratory results and medication usage, and detect any anomalies that might indicate a health issue or the need for further medical attention; and predicting disease progression or recurrence, which can help guide treatment decisions and follow-up schedules. Anomaly detection in urology can be performed using various ML techniques, such as SVM, decision trees and NNs. These algorithms can be trained on large datasets of patient data and validated using test datasets to ensure accuracy and reliability.
A, The difference between clustering, classification, regression and anomaly detection methods is shown. Aa, Clustering problems classify inputs in different groups in an unsupervised manner; for example, defining new prostate cancer subtypes by training an unsupervised algorithm on RNA sequencing data. Ab, In classification problems, models are constructed using classification algorithms, and these models are subsequently applied to unseen instances for prediction, considering factors such as feature selection, train and test procedures, and variables such as prostate specific antigen (PSA) value, age, estimated prostate volume and other clinical variables to determine whether a patient has prostate cancer (yes or no). Ac, Regression problems predict a continuous value based on the input; for example, predicting the chance of prostate cancer based on a PSA level of 4 ng/ml. Regression problems use linear regression and decision tree algorithms, but support vector machine can also be used. Ad, Anomaly detection consists of identifying the outliers within a dataset, for example detecting cancer by training an anomaly detection model on transcriptomic data from normal samples and using this model to differentiate cancer based on the deviation of cancer transcriptomes from normal. B, Example of an input table describing discriminative features for a urological problem to predict the survival of a patient based on patient profile using a supervised artificial intelligence (AI) approach. In this scenario, the goal is to predict the survival of patients based on their profile using discriminative features. Supervised learning requires labelled data with a specific target variable to train the AI model and make predictions. C, Visualization of underfitting, overfitting and optimal fitting. Ca, Representation of a model. Each dot is a data point. Blue dots are the training set and red dots are the testing set. Cb, Visualization of overfitting. The differences between a predicted value (length of dashed lines) and a true value (dotted line) is the residual error, and bias can be considered as mean of residual errors. In overfitting, the training model (blue dots) shows low bias but high variation, as the model tightly fits the training data but fails to generalize well to new instances, leading to inconsistent and unreliable predictions. However, when applied to the testing dataset, the model struggles to generalize and adapt to new instances, leading to a high bias. This effect occurs because the model is too specialized to the training data, failing to capture the underlying patterns in the broader context of the testing dataset. Cc, Visualization of underfitting. In underfitting, the model shows both high bias and low variation, indicating the inability of this model to accurately capture the underlying patterns and complexities of the data. Cd, Visualization of optimal fitting. The double-headed arrows indicating ‘variability’ in the figure refer to the discrepancy between the distance of the blue dots (training set) from the fitting line and the distance of the red dots (testing set) from the fitting line (dashed lines).
Sometimes, supervised and unsupervised approaches are combined in semi-supervised learning, in which small amounts of labelled data are combined with large amounts of unlabelled data. This approach can improve performance in situations in which labelled data are costly to obtain12; for example, annotation of histopathology images, particularly those of variable quality and/or complexity is laborious and costly, and requires specialized expertise. In this context, semi-supervised learning provides a practical compromise between the wide availability of unlabelled data from routine histopathology and the need for well-annotated data required to train a model using supervised learning13. Semi-supervised learning has been used successfully for grading of genitourinary malignancies including prostate and bladder cancer14,15.
Other categories of AI include artificial neural networks (ANNs), deep learning (DL) and NLP, a specific type of ML that focuses on the identification and extraction of clinically relevant information from medical records and reports16. Each of these analyses has been used in urology (Fig. 2A). ANNs and DL are distinguished from ML approaches by the ability to achieve high performance even with large amounts of data, along with reduced dependence on human guidance for both input data and computational costs. To solve a problem using ML, input features need to be extracted carefully, which often requires substantial domain expertise (for example, to identify tumour areas from digital histological images). Conversely, with ANNs, feature extraction is performed automatically. ML can be used as a single model or built as an ensemble of models to solve a problem. These models can then be incorporated into ANN layers to enable prediction. ANNs and DL also differ in the number of network layers, as a typical ANN includes two or three layers, whereas a DL network has more layers, and therefore can be used to handle large amounts of data. Traditional ML algorithms rely on manual feature engineering, and therefore struggle with complex and non-linear data relationships. When data are high-dimensional or unstructured (such as images, audio or text), DL shows superiority by automatically extracting intricate patterns. However, traditional ANNs might fail with highly complex problems, in which the deep architectures of DL excel. Conversely, DL models require a large amount of data for successful training and under-perform with limited labelled data, risking overfitting, in turn making traditional ML a suitable choice. The interpretable models of ML handle data scarcity while capturing complex patterns with manual feature engineering and statistical learning principles.
A, Artificial intelligence (AI) consists of programming systems to perform tasks that normally require a human level of intelligence. Machine learning (ML) consists of using and developing computer systems that are able to learn and adapt without following explicit instructions by using algorithms and statistical models to analyse and draw inferences from patterns in data. A neural network (NN) consists of a series of algorithms that endeavours to recognize underlying relationships in a set of data through a process that mimics the way the human brain operates. Deep learning (DL) refers to a neuronal network with more than three hidden layers. In the figure, the concentric structure shows the hierarchical relationship between different concepts in the field of AI. B, Schematic representation of a NN structure (part Ba), forward propagation (inference) (part Bb), and back-propagation (part Bc). A NN is a computational model inspired by the structure and function of biological NNs used for learning and making predictions from data. Inference refers to the process of using a trained model to make predictions or draw conclusions based on new, unseen data. Back-propagation is a method used to train NNs by calculating and adjusting the weights of the network based on the error between predicted and actual outputs (blue arrow). PSA, prostate-specific antigen.
ANNs are inspired by the human brain and are composed of a set of algorithms. An ANN is composed of processing elements termed neurons, nodes or perceptrons17,18, which are connected through links, each with a particular associated weight (Fig. 2B). Neurons that receive information (for example, quantifiable patient information such as prostate-specific antigen (PSA) level, gene copy number or mutation status) are termed input neurons, which process information and transfer this information to other neurons within the NN, generating an output (output neurons). The weight is the strength of the connection between neurons, with larger weight reflecting larger influence, and indicates how much influence the input will have on the output. The simplest ANN has three main components: an input layer, a hidden layer and an output layer (Fig. 2B). Layers are formed by groups of neurons in which calculations take place, with the calculations from one layer typically transferred to the next layer, although some networks enable the information to be transferred within a layer or to a previous layer. In a simple example, an input layer consisting of clinical variables such as patient age, PSA level and tumour volume could be processed through a hidden layer to generate an output decision of whether to perform radical prostatectomy or not (Fig. 2Ba). A situation in which information is processed through multiple hidden layers to arrive at a decision would be termed DL.
The number of hidden layers needed to discriminate between artificial ANNs and DL depends on the specific problem and the complexity of the dataset. Generally, DL models have more than one hidden layer, whereas ANN models have only one hidden layer. However, the number of hidden layers alone does not determine whether a model is classified as an ANN or DL. DL models typically use a larger number of hidden layers than ANNs, but also use more complex architectures and algorithms, such as convolutional NNs (CNNs) or recurrent NNs, that enable these models to learn hierarchical features and patterns in the data. In general, a DL model with more than one hidden layer is better suited for tasks that require learning complex features and patterns, such as image or speech recognition, whereas an ANN model with a single hidden layer might be sufficient for simple tasks, such as classification of numerical data.
Ultimately, the choice between ANN and DL models depends on the specific problem, the available data and the computational resources and expertise available. Investigators need to carefully evaluate the performance and interpretability of the models and choose the appropriate model for a specific problem. Most DL networks are feedforward, which means that the information is propagated forward from input to output (Fig. 2Bb). Conversely, models can be trained in the opposite direction — from output to input — in a process called backpropagation (Fig. 2Bc), in which mathematical methods are used to adjust how accurately a NN processes specific inputs. This process enables the predicted output to be compared with the actual output, and, in turn, the network can be refined.
Several general performance trends need to be considered to compare ML, NN and DL algorithms on datasets of varying size (Box 1). In general, the performance of ML, NN and DL algorithms depends on various factors, including the size and complexity of the dataset, the specific problem and the computational resources available. Choosing the appropriate algorithm for a specific problem and carefully evaluating the performance and interpretability of the model is important.
R and Python are two of the most popular programming languages for ML. The choice between R and Python for ML depends on factors such as the specific project requirements, the user’s skillset and experience, and the availability of resources and support (a working example of coding in R or Python to predict life expectancy in patients with bladder cancer is provided in the Supplementary Information). Additional aspects of AI, ML and DL approaches have been discussed in detail in other reviews19,20.
Bioinformatics tools in urology
Among the earliest clinical applications of bioinformatics analysis in urology was the ability to define discrete molecular subtypes through gene expression data from tumours that are histologically similar, and to link subtypes to important clinical characteristics. In early studies21,22,23,24,25,26, either cDNA or oligonucleotide microarrays — the technologies routinely available at the time — were used to identify differentially expressed genes between tumours and non-tumour tissues. A major contributor to these efforts was the development of Oncomine, a database of microarray datasets and a web browser enabling data mining27. This resource enabled interrogation of differentially expressed genes both across and within cancer types, along with gene ontology annotations to indicate function and subcellular localization, and, in turn, the potential targetability of a gene or set of genes. As investigators began to evaluate thousands or tens of thousands of genes simultaneously, as opposed to a handful of genes in previous studies, new approaches for statistical analysis were needed to account for multiple comparisons, and also to discern patterns inherent in the data.
With the development of NGS, expression profiling largely moved from the array technology. NGS involves direct sequencing of nucleic acid molecules and, differently from microarray analysis, does not require prior knowledge of the genes or transcripts to be interrogated. Thus, NGS enables the discovery of new genes, non-coding sequences and nucleotide variations such as mutations and splice variants. One notable example of the power of NGS is the development of The Cancer Genome Atlas (TCGA), an initiative that aims to gain insights into the genetic and genomic basis of cancer. The atlas includes molecular data on DNA, RNA, protein and epigenetic modifications, providing novel insights into copy number variations, gene mutations, fusions, transcriptional profiles and proteomic profiles across more than 30 types of cancer. Analysis of TCGA data has uncovered previously unanticipated tumour subtypes, improving accuracy of patient phenotyping and prognosis, as well as actionable molecular targets and pathways driving cancer progression. Through this initiative, standards for sample acquisition and processing (such as tumour cell enrichment and clinical data), analytical platforms (microarrays versus sequencing) and computational analysis were established to harmonize the information obtained from multiple different centres. Of relevance to urology, atlases for bladder urothelial carcinoma28, prostate adenocarcinoma29, testicular germ cell tumours (GCT) 30, kidney clear cell carcinoma31 and kidney papillary cell carcinoma32 have been published to date. NGS has been the mainstay for generation of cancer atlases and for discovery-based studies; however, the cost, analytical complexity and need for high-quality RNA particularly from archival specimens render NGS prohibitive for clinical use. The NanoString nCounter platform has emerged as a potential solution to enable direct (non-amplified) measurement of several hundred mRNA targets in a single sample concurrently, with high sensitivity and reproducibility33.
Many tools and resources for interrogation of gene or protein expression with a prognostic value in cancer are available34 (Table 1). These resources include cBioPortal, for the evaluation of multiple genomic readouts in a specific tumour type, such as copy number alterations, mutations, methylation data, mRNA and microRNA (miRNA) expression, and protein data35; UALCAN, for the analysis of RNA sequencing (RNA-seq) and clinical data from TCGA36; and TCPAv3.0, for the analysis of reverse phase protein array (RPPA) data from tumours37.
Classification approaches to define molecular subtypes of tumours in genitourinary oncology have largely relied on RNA-seq or microarray analysis applied to bulk tissue, in which transcript levels within a tumour specimen are averaged. These approaches are associated with a loss of information at the level of individual cells. Moreover, tumours are heterogeneous, resulting from the coexistence of multiple cell lineages and differentiation stages, some of which are influenced by clonal evolution. Thus, tissue analysis pipelines and bioinformatics approaches that enable molecular profiling at single-cell resolution have been increasingly used.
Single-cell technologies
Single-cell RNA-seq (scRNA-seq) is by far the most widely used single cell approach to probe cellular heterogeneity38. In cancer biology, scRNA-seq has provided important insights into tumour heterogeneity, complexity of the tumour microenvironment39 and tumour evolution over time or with treatment40, and has led to the identification of previously unidentified cell types41. In benign urology, single-cell approaches have helped clarify the role of the immune microenvironment in interstitial cystitis (IC)–bladder pain syndrome (IC–BPS)42,43 and identify cell types not previously described in the prostate44, including cells implicated in prostate regeneration following androgen deprivation45. Clustering and dimensionality reduction of scRNA-seq data are performed using several approaches including PCA, t-distributed stochastic neighbour embedding, uniform manifold approximation and projection with Seurat46, pcaReduce47, SC3 (ref. 48) and SNN-cliq49 (Fig. 3 and Table 2). Annotation of clusters is then performed to infer specific cell types and biological states50.
The ability to interrogate tumours at single-cell resolution has enabled investigators to identify the cell of origin for various tumour types including renal cell carcinoma (RCC)51,52 and prostate cancer53,54, in turn providing novel insights into crucial events in cancer initiation. These analyses helped identify cell states associated with disease progression; additionally, transition analysis using tools such as Monocle55, diffusion pseudotime (DPT)56, Wishbone57 and Waddington-OT58 enabled investigators to estimate temporal evolution and infer cellular trajectories in cancer (Fig. 3). In one study, the heterogeneity of prostate cancer was assessed by analysing >30,000 single-cell transcriptomes from multiple prostate tumours, and several distinct transcriptional programmes associated with tumour progression were identified, including an activated endothelial cell type implicated in invasion and enriched in castration-resistant prostate cancer (CRPC)59. In bladder cancer, single-cell sequencing led to the identification of a type of inflammatory cancer-associated fibroblast implicated in tumour progression, and of tumour-promoting mechanisms mediated by these cells60.
With the emergence of immune checkpoint inhibitors, much emphasis has been placed on defining the tumour immune contexture at single-cell resolution61, particularly to understand immune mechanisms associated with poor responses to these therapeutics. In several studies, scRNA-seq was used to define the cellular landscape of normal and cancerous tissue to identify dysregulation of immune cells in the context of cancer62,63,64,65. In one of these studies, no substantial differences in the number of immune cells were observed between prostate cancer and normal tissue samples; however, transcriptional alterations in specific immune cell subsets were identified in prostate cancer compared with normal tissue samples62. For example, decreased expression of genes associated with antigen presentation and processing, as well as genes encoding co-activating receptors such as CD40, was observed in mononuclear phagocytes. Conversely, in a prostate-specific subset of macrophages termed MAC-MT, which are enriched for metallothionein gene expression, the expression of both co-activating receptors and of the B cell survival factor BAFF was increased in prostate cancer samples. Moreover, differently from other macrophage subsets, MAC-MT were enriched in IFNγ and TNF gene signatures. In agreement with activation of this pro-inflammatory transcriptional programme, prostate cancer biopsy samples enriched with MAC-MT were associated with improved disease-free survival. Thus, the use of scRNA-seq led to the identification of a novel macrophage subset enriched in prostate cancer that is associated with favourable prognosis and that provides new insights into prostate biology.
The latest extension of scRNA analysis is spatial transcriptomics (Fig. 3), in which transcriptomics is applied to discrete regions of tissues in situ. Briefly, RNA is isolated directly from tissue sections immobilized on specialized slides that enable RNA capture and reverse transcription to cDNA for subsequent sequencing. Gene expression profiles can then be ascribed to specific cells within the tissue section by registration with a conventional haematoxylin and eosin (H&E) image of the section66. Current platforms for spatial analysis include Slide-seq67, Visium from 10x Genomics and the GeoMx digital spatial profiling platform from Nanostring68. Spatial transcriptomics enables retention of valuable information on tissue organization and cell–cell interactions that is lost with the tissue dissociation that is required for conventional scRNA-seq analysis69. Analysis focuses on five steps70,71 including spatial clustering performed using methods such as SpaCell72 or SpaGCN73; spatially variable gene detection using Trendsceek74, SpatialDE75 or SPARK76; cell-type deconvolution using stereoscope77 or SPOTlight78; enhancement of gene expression resolution with RCTD79 or XFuse80; and inference of cell–cell communication with Giotto81 or SpaOTsc82.
Spatial transcriptomics in urology has been used, for example, to explore gene expression within multiple cancer foci within a single prostate83, highlighting extensive heterogeneity (including differences in gene expression between central and peripheral regions of the tumour) and providing important new insights into the interactions between tumour and microenvironment. The authors noted that pathways such as the citrate cycle, oxidative phosphorylation and pentose phosphate pathway were activated in the centre of one tumour focus, whereas pathways activated at the periphery were related to inflammation. Additionally, inflammation and reactive stroma were observed early in tumour development, indicating that these phenomena might precede detection of genetic changes within the tumour. Normal stroma showed enrichment for actin cytoskeleton and motility, whereas reactive stroma associated with the tumour was enriched for oxidative stress and integrin-linked kinase activity. Building on these metabolic alterations identified within specific regions of the primary tumour, a new informatics pipeline was used to identify metabolic genes and pathways that differed in discrete regions of the same tumour, and to gain new insights into how tumour cells adapt to the immediate microenvironment84. Moreover, different targets specific to tumour cells were identified, including the fatty acid desaturase SCD1, the prostaglandin transporter SLCO2A1, as well as several additional perturbations in carbon, lipid and amino acid metabolism that could be targeted using small-molecule inhibitors to kill tumour cells.
Bulk RNA sequencing deconvolution
scRNA-seq is a powerful tool, but cannot currently be used on a large scale owing to the high cost. Additionally, substantial technical expertise is required to perform the technique robustly, and dissociation protocols are known to affect gene expression, particularly for solid tissue samples such as tumours85. To circumvent these challenges, several methodologies have been developed during the past two decades to infer proportions of individual cell types from bulk transcriptomics data. Currently, >40 different bulk RNA-seq deconvolution methods have been developed86, among which CIBERSORTx has been widely used. CIBERSORTx imputes gene expression profiles and offers an estimation of the abundances of different cell types in a mixed cell population using gene expression data (from RNA-seq or microarray analysis) and the support vector regression ML algorithm87. An early version of this algorithm — CIBERSORT — was used to measure infiltration of immune cell subsets and identify potential novel diagnostic biomarkers for Hunner’s lesion IC (HIC)88. Application of CIBERSORT to publicly available microarray datasets from GEO89,90 showed an enrichment of T follicular helper cells and CD3+CD4+HLA-DR+ memory T cells in HIC compared with controls. Neither cell population had been observed previously in HIC, and the authors concluded that these cell subsets could have utility as diagnostic biomarkers. IC is a diagnosis of exclusion, and the underlying pathobiology is incompletely understood; thus, the identification of previously unanticipated cell types provided new insights on pathogenesis, and also highlights potential biomarkers for diagnosis. Validation of these findings in independent studies will be important to support this interesting hypothesis.
Deconvolution methods enabled researchers to use the wealth of existing bulk RNA-seq data such as that in TCGA to infer different cell types relevant to urological diseases. In one study, deconvolution was used in addition to transcriptomics and proteomics analyses to estimate the presence of immune cells in tumours from patients with non-muscle-invasive bladder cancer (NMIBC)91. In this study, high immune cell infiltration was observed in aggressive class 2b tumours, with an enrichment of cytotoxic T lymphocytes and T helper (TH) cells. Moreover, this high immune infiltration was associated with a reduced risk of tumour recurrence (P = 0.022). Thus, results obtained from deconvolution provided valuable insights into the biology associated with NMIBC tumour subtypes. Single-cell analyses in the context of clinical studies are still lacking owing to cost, inadequate sample collection and a paucity of clinical follow-up data. To address this challenge, in a study involving 412 patients with muscle-invasive bladder cancer (MIBC), scRNA-seq analysis of normal bladder tissue was combined with deconvolution of bulk RNA-seq data from TCGA to explore intratumoural heterogeneity and estimate cell types and epithelial lineages within bladder tumours92. scRNA-seq of normal bladder led to the identification of five cell clusters — basal, intermediate, umbrella, epithelial-to-mesenchymal transition (EMT)-like and TNNT1+ — based on the expression of marker genes, which were used subsequently to deconvolute bulk RNA-seq data from TCGA. Deconvolution analysis led to the identification of five epithelial cell states within bladder tumours, among which umbrella cells and EMT-like cells were the dominant types. Subsequently, the association between epithelial cell lineages inferred by deconvolution and clinical outcomes based on data from TCGA was assessed, and a significant reduction in overall survival (OS) was observed in patients with a predominant EMT-like lineage compared with patients without enrichment in the EMT-like lineage (P = 0.0009). Thus, valuable prognostic information can be derived through deconvolution of bulk RNA data. In another study63, scRNA-seq was carried out on tumour and normal tissue samples from patients with discrete subtypes of non-clear-cell RCC (nccRCC) including papillary RCC (pRCC), chromophobe RCC (chRCC), collecting duct carcinoma and sarcomatoid RCC. This analysis was complemented with deconvolution of bulk RNA-seq data from 274 patients with nccRCC using CIBERSORTx to investigate cellular heterogeneity among nccRCC subtypes and explore the tumour microenvironment. Results from this study showed that both exhaustion of CD8+ T cells and enrichment of tumour-associated macrophages were correlated with a poor prognosis across nccRCC subtypes. Results from this study highlighted the power of deconvolution to extract meaningful information regarding cellular composition of the tumour microenvironment from bulk RNA-seq data.
Bladder urothelial carcinoma
Bladder urothelial carcinoma is a heterogeneous disease classified historically as either NMIBC or MIBC. Until the early 2000s, bladder urothelial carcinoma prognosis was based on histopathological assessment of tumour grade and stage at presentation. Discrete molecular changes were known to be associated with NMIBC or MIBC, but the advent of expression profiling together with the development of appropriate analytical tools has led to the identification of molecular subtypes associated with prognosis and treatment response. Furthermore, novel DL algorithms have been applied in the context of diagnostic urine cytology. Together, these advances highlight the influence of bioinformatics on improved understanding of bladder cancer pathophysiology and management.
Bladder cancer diagnosis
Bladder cancer is most often revealed by haematuria, which typically prompts preliminary evaluation by urinary cytology and cystoscopy. Cystoscopy is performed using both conventional white light cystoscopy (WLC) and blue light cystoscopy93. Cystoscopy is largely effective, but a considerable proportion of tumours remain undetected, highlighting the need for improved detection strategies. Blue light cystoscopy has been shown to improve detection, but the widespread use of this approach is hampered by the need for specialized instrumentation. Thus, a number of groups have used ML approaches to enhance detection and diagnosis from WLC images. In one study, CNNs were used to develop CystoNet, a DL algorithm, to enhance detection of bladder cancer from white light video images94. CystoNet enabled the detection of 95% of papillary tumours and 100% of flat tumours analysed, showing robust sensitivity and specificity. A similar image-based approach using a CNN was applied to blue light images, and showed sensitivity and specificity of ~96% and ~88%, respectively, for tumour classification95. In another study, a Cystoscopy Artificial Intelligence Diagnostic System (CAIDS) was designed to interpret cystoscopy images based on standardized identification of relevant features96, and showed a diagnostic accuracy ranging from 97.8% to 99.1% in validation datasets, improving speed of image assessment and sensitivity compared with those obtained by expert urologists (speed: 12 s with CAIDS versus 35 min by expert urologists; sensitivity 95.4% versus 75.4% with CAIDS and expert urologists, respectively). Results from this and other studies97,98 showed a clear benefit of AI in diagnostic cystoscopy, but the performance of this approach will need to be assessed in prospective studies including diverse patient populations before routine integration into the clinical workflow99.
Urine cytology relies on pathological assessment of cellular features such as nuclear hyperchromaticity and the nuclear-to-cytoplasmic ratio to assess tumour grade. Several groups have developed AI algorithms to improve assessment of urine cytology, particularly to overcome the challenge of poor inter-observer agreement. In one study100, digital images from 1,615 patients undergoing urine cytology for detection or follow-up monitoring of urothelial carcinoma were analysed using six CNNs to extract both cellular features (such as nuclear-to-cytoplasmic ratio, irregularity of nuclear membrane and chromatin granularity) and slide-level features (such as urothelial cell number and atypical cell count) that were subsequently used to train a classifier to predict diagnosis. The classifier was validated on a further 790 images and achieved an optimal sensitivity of 79.5%, a specificity of 84.5% and an area under the curve (AUC) of 0.88 for high-grade urothelial carcinoma. This study constituted a considerable advance in terms of efficiency of screening of whole-slide images, as well as improved accuracy in diagnosis. In a similar analysis, a different DL approach, namely the 16-layer Visual Geometry Group CNN, was implemented to analyse urine cytology slides101, and achieved an AUC of 0.989 in discriminating benign from malignant specimens, with a sensitivity of 90.51% and a specificity of 96.82%. This study was based on a small sample size, but the results showed that the accuracy of diagnosis was improved over earlier studies by increasing the depth of the network. To explore the utility of AI to improve urine cytology in a prospective setting, a multicentre study (VISIOCYT1) was conducted using VisioCyt screening, which combines urine cytology image analysis with AI for accurate prediction of urothelial carcinoma. The results from the first phase of this study, including 598 patients (449 with confirmed bladder tumours and 149 without bladder tumours), showed a substantial improvement in sensitivity and specificity with VisioCyt (84.9% and 81.2%) compared with standard urine cytology (43% and 100%). Moreover, the enhanced sensitivity of VisioCyt was most notable for low-grade tumours, with a 77% sensitivity for VisioCyt versus 26.3% for standard pathological review102. The use of VisioCyt might reduce the need for repeating an invasive procedure such as cystoscopy, in turn maintaining patient adherence to routine monitoring for tumour recurrence.
Subtyping to improve classification and prognosis
Molecular profiling has provided new insights into the biology of bladder tumours, and has also enabled identification of molecular features of both prognostic and predictive clinical benefit. In early studies, gene expression patterns and genomic alterations were leveraged to classify tumours using hierarchical clustering to identify molecular subtypes, and the prognostic ability of this approach was compared with that of standard histopathological evaluation21,25,26,103,104,105. In 2012, the first molecular taxonomy for Ta and T1 bladder cancer based on hierarchical clustering was published, and included five major subtypes of urothelial carcinoma with different prognoses106: urobasal A; genomically unstable; urobasal B; squamous cell carcinoma (SCC)-like; and infiltrated. In similar studies, consensus clustering and supervised clustering of mRNA expression data from MIBCs were used to define discrete subtypes reminiscent of breast tumour subtypes107,108. Results from these studies showed that the basal subtype had the worst prognosis, whereas p53-like tumours were associated with resistance to neoadjuvant chemotherapy (NAC), highlighting the potential clinical relevance of subtype identification. In another study, a 47-gene predictor, BASE47, was developed to define the minimal gene set enabling accurate subtype classification; since this publication, BASE47 has undergone optimization as a potential clinical-grade diagnostic assay109.
Further molecular subtyping efforts by TCGA Research Network yielded four subtypes based on 131 MIBC samples28, whereas an updated multiplatform analysis by TCGA based on >400 MIBC samples led to the identification of five molecular subtypes that showed substantial overlap with clusters described previously and provided prognostic information110. In this study, a neuronal subtype was also defined, and was associated with the lowest survival; these results have been validated in independent cohorts111,112,113,114.
Similar clustering analyses have been performed in early-stage NMIBC91,115,116, which constitute the bulk of bladder cancer diagnoses. In these studies, discrete molecular subtypes with distinct biological behaviour, outcome and likelihood of treatment response were identified. In the first large-scale transcriptomics analysis of NMIBC, tumour samples (n = 460) were accrued by the UROMOL consortium115. Unsupervised clustering showed three molecular subtypes, with class 1 tumours enriched for early cell-cycle genes, class 2 tumours enriched for late cell-cycle genes, and class 3 tumours showing frequent expression of the stem cell marker CD44 and primitive cytokeratins. This study was under-powered to estimate progression to MIBC; however, class 2 tumours were enriched in samples from patients with an increased risk of progression compared with the other classes. In an independent analysis of NMIBC samples (n = 140), two genomic subclasses of Ta tumours characterized by different copy number profiles were identified, GS1 (no or few copy number alterations) and GS2 (frequent chromosome 9 deletion)116. Based on RNA expression profiles, samples in this study aligned closely with the previously described Urobasal A subtype106 and with UROMOL2016 class 2 samples115, although the extent of progression to MIBC differed. In another study, bulk RNA-seq data from patients with Ta, T1 or carcinoma in situ were used to perform consensus clustering of the 4,000 most varying genes and identified four classes — 1, 2a, 2b and 3 — which differed in their association with progression-free survival (PFS) (1 > 3 > 2b > 2a) and in the association with clinical parameters. In this analysis, class 2 was divided into 2a, which showed high genomic instability and tumour mutational burden (TMB), and class 2b, which was enriched for immune infiltration. In addition to expression of genes associated with cancer stem cells and EMT, class 2b tumours were shown to have a higher total immune infiltration score than all other classes, reflecting an enrichment of immune-related gene expression and immune cell infiltration as verified by spatial proteomics analysis91. The tumour immune microenvironment has been implicated as an important determinant of response to Bacillus Calmette–Guérin (BCG)117, a mainstay of treatment for NMIBC. Multiomics analysis of NMIBC specimens from patients treated with BCG was carried out to explore the molecular basis of BCG response or failure. In this study, T cell exhaustion following BCG treatment was observed in patients with high-grade recurrence118; moreover, gene expression signatures in pretreatment tissues from patients who experienced high-grade recurrence following BCG treatment were enriched in cell cycle and immune-related genes. Notably, tumours that were defined as class 2a and 2b before treatment based on the UROMOL2021 classification91 had a worse outcome following BCG treatment. Thus, both pretreatment subtype and tumour features could be used to identify patients at risk of high-grade recurrence in response to BCG. The refinements to the classification of NMIBC continue to provide insights on the molecular basis underlying different clinical outcomes; however, large studies, including clinical trials, are required to validate these observations.
Considering the multitude of subtyping efforts and the challenges that arise in comparing the different approaches, researchers have been motivated to harmonize classification schemes to improve clinical application and use of these schemes. A consensus molecular classification of MIBC was published in 2020 and included the definition of six subtypes based on >1,700 transcriptomic profiles119. Importantly, information from multiple classification systems has been incorporated into a commercially available test termed Decipher Bladder, in which an oligonucleotide array of >200 genes is used to classify a tumour as one of five subtypes: luminal; luminal infiltrated; basal; basal claudin-low; or neuroendocrine-like. This test is used with existing clinical parameters to inform treatment decisions in patients with urothelial carcinoma112,120,121.
To understand whether molecular subtypes are associated with clinical outcomes, a quantitative PCR with reverse transcription (RT-qPCR) panel of marker genes derived from published reports was used to interrogate several independent cohorts of MIBC samples and compared with published basal and luminal subtypes obtained with other classifications (such as TCGA and Lund) for the ability to predict metastasis and survival122. Molecular subtypes were associated with tumour grade, but clinical parameters, such as tumour stage, nodal status and lymphovascular invasion, outperformed molecular subtypes in predicting recurrence-free survival and OS. Furthermore, co-occurrence of markers of the basal and luminal subtypes were observed within the same tumour specimen, emphasizing intratumour heterogeneity. Based on this analysis, the authors concluded that molecular subtypes are reflective of tumour biology, but the prognostic utility of these subtypes requires further evaluation.
Starting from the observation that clinical parameters showed superior ability to predict outcomes in bladder cancer123, in a 2020 study, a DL approach was used to predict the molecular subtype based on whole-slide histological images of MIBCs123. A NN was used to classify a training set of H&E images from MIBC samples in TCGA database for which subtyping was also performed. Using class activation maps — a method to highlight which regions in an image are used by a NN for classification — the authors identified specific histological and morphological features that were most relevant to each subtype, such as pleomorphic nuclei for basal tumours or mesenchymal cell morphology for luminal p53-like tumours. The application of this approach to a validation cohort enabled the prediction of molecular subtypes based only on histological features. Subtype classification by the NN obtained from histology alone was superior in terms of accuracy to evaluation by pathologists, particularly when the pathologists were provided with only small tiles from the whole-slide images. However, when provided with whole-slide images as well as class activation maps, the overall accuracy of classification by the pathologists improved from ~38% to nearly 60%.
Beyond broad tumour subtypes, informatics in the form of AI has been used in the context of discrete molecular alterations. The fibroblast growth factor receptor (FGFR) inhibitor erdafitinib is the first targeted therapy approved for the treatment of advanced bladder cancer124. However, the use of this therapy is restricted to patients with FGFR3 or FGFR2 mutations, the identification of which requires specialized molecular assessment. To circumvent the need for costly molecular testing, AI was used to detect FGFR3 mutation status from H&E-stained tissues125. Using digital images from >300 patients with MIBC from TCGA, along with the FGFR3 mutation status determined by whole-exome sequencing, the authors trained a DL network to predict FGFR3 mutation status from histology. The algorithm was validated on an independent cohort of 182 tumours. The algorithm showed effective identification of tumours harbouring FGFR3 mutations, with AUCs of 0.701 and 0.725 in the training and validation cohorts, respectively, compared with evaluation by an expert pathologist (AUCs of 0.563 and 0.607 for the training and validation cohorts, respectively). The algorithm was also able to predict the presence of heterogeneity in mutation status within a specific tumour. These findings indicate that the use of a DL algorithm led to the identification of FGFR mutations in histological images that were not detectable by a pathologist. Thus, the authors concluded that the application of DL to patient selection is feasible and could be incorporated into clinical management pending prospective validation in additional studies.
Subtyping to predict treatment response
In addition to tumour classification, gene classifiers and/or molecular subtypes have also been explored as tools to predict response to treatment, particularly in patients receiving cisplatin-based NAC for muscle-invasive disease126. Considering the substantial toxicity associated with chemotherapy and the effect of delays on alternative treatments, gene expression analysis has been added to existing prognostic factors to identify patients who are likely to respond — or not — to cisplatin-based NAC127. Results from a retrospective, microarray-based gene expression profiling of tumour samples from patients receiving cisplatin-based NAC showed discrete gene sets that distinguished responders from non-responders128,129,130. Based on these observations, a small prospective study was conducted to determine whether a predicted response (to either methotrexate–vinblastine–doxorubicin–cisplatin (MVAC) or carboplatin–gemcitabine (CaG)) based on expression profiling corresponded to an actual response to NAC131. The expression of genes that had been previously identified as predictive of response to MVAC (14 genes) or CaG (12 genes)128,129 was assessed by RT-qPCR in biopsy samples obtained from patients (n = 33) before receiving NAC; a prediction score was calculated and was used to prospectively assign patients to receive MVAC or CaG. A total of 88% of patients showed tumour shrinkage following treatment with the appropriate drug regimen, as well as improved survival. The authors concluded that in principle, expression profiles could help identify patients who are most likely to respond to NAC, but large prospective trials are needed. Similarly, in another study, gene expression profiling was used to determine whether molecular subtypes could affect clinical outcomes in patients receiving cisplatin-based chemotherapy plus bevacizumab132. In this study, patients with basal subtype tumours showed increased 5-year OS (91%) compared with patients with luminal (73%) or p53-like (36%) tumour subtypes (P = 0.015, log-rank test). Bone metastases were evident only in patients with p53-like subtype tumours, and these tumours were chemoresistant. The authors recognized that the small number of specimens suitable for expression profiling was a limitation; however, these findings highlight the heterogeneity of urothelial carcinoma, as well as the potential utility of subtypes to inform treatment.
The emergence of multiple classification schemes and the lack of harmonization has been a challenge in obtaining a potential clinical benefit from tumour subtyping. To address this challenge, the concordance between four molecular subtyping classification schemes — consisting of three to five subtypes — was assessed. In this study, four schemes were assessed in predicting outcomes in patients treated with or without NAC: University of North Carolina (UNC; claudin-low, basal, luminal); MD Anderson Cancer Center (MDA; basal, p53-like, luminal); TCGA (clusters IV, III, II, I); and Lund (SCC-like, Uro B, infiltrated, Uro A, genomically unstable)133. First, the authors performed whole transcriptome profiling on tissues from 343 patients with MIBC obtained before NAC; samples were subsequently classified according to subtypes using all four schemes, and, lastly, the association with survival was determined. Patients with basal subtype tumours (basal, Uro B, SCC-like or cluster III, depending on the classification scheme), who had worse OS than patients with luminal subtype tumours before NAC treatment, showed a marked improvement in OS with NAC, whereas patients with claudin-low/cluster IV tumours or p53-like tumours showed no improvement in survival. These observations showed the potential clinical relevance of subtyping before treatment. Molecular subtyping schemes are valuable to compare relative gene expression patterns among tumours from groups of patients; however, these schemes are of limited utility to assign an individual patient sample to a subtype. To address this limitation, the authors trained a genomic subtyping classifier (GSC) to assign a single sample to one of four subtypes (basal, claudin-low, luminal or luminal–infiltrated) based on the UNC, MDA, TCGA and Lund classifications and on sensitivity to treatment. Patients with tumours defined as basal using the GSC showed 3-year OS of 49% in the absence of treatment, which increased to 78% with NAC (n = 68; P < 0.001). Patients with luminal tumours had the best OS irrespective of NAC. Collectively, these findings confirmed a change of outcome in patients with basal subtype tumours following NAC, and provided proof-of-concept for the potential utility of subtyping using the GSC to guide appropriate treatment. Bladder cancer is known to be heterogeneous, with histological variants often co-occurring with conventional urothelial carcinomas in a specific lesion134. Considering this concern, some authors have suggested that treatment decisions based on subtype classification approaches, especially using limited amounts of tissue, might not adequately capture the presence of histological variants135, and, therefore, caution should be taken when associating tumour subtypes with prognosis or treatment planning. Moreover, although much research is being conducted in this area, the current consensus is that molecular subtyping can assist in stratification of patients for NAC, but further prospective validation is needed to assess to what extent subtypes can inform treatment136.
Immune checkpoint blockade has emerged as a promising new therapeutic option for bladder cancer137. Thus, genomic classifiers have also been evaluated for their ability to predict response to immunotherapy. In a study in which a novel classifier derived from TCGA subtypes110 was applied to transcriptome data from the IMvigor210 trial (in which the PDL1 inhibitor atezolizumab was assessed)138, patients with the neuronal subtype showed a high objective response rate to immunotherapy, which was associated with the best OS among all subtypes (P = 0.012)139. In another study, several classifiers including the GSC133, the consensus classifier119, the TCGA subtypes and the bladder cancer-specific Immune190 signature (based on differentially expressed genes in PPARγ active tumours140) were applied to expression data from patients in the PURE-01 trial (in which neoadjuvant pembrolizumab was assessed in patients with MIBC)141. Regardless of the classifier used, basal subtype tumours were enriched for immune marker gene expression. Additionally, patients with basal subtype tumours showing high median Immune190 scores had increased PFS compared with patients with basal tumours with low Immune190 scores (P = 0.04). Thus, molecular subtyping based on immune signatures might help identify patients who are likely to benefit from immunotherapy.
Differently from tumour subtyping schemes, in several studies, gene expression markers identified from in vitro drug treatment assays have been used to understand drivers of chemosensitivity and chemoresistance, and to propose novel therapeutic strategies142,143. Analysis of the gene expression and drug response profile of the NCI-60 cell line panel to >100,000 chemical compounds, together with expression profiles from 40 bladder cancer cell lines, led to development of a gene expression biomarker-based approach named the co-expression extrapolation (COXEN) algorithm144. Briefly, this approach is used to define a gene expression signature that reflects sensitivity to a drug (for example, cisplatin) across multiple cell lines, and to determine the degree of co-expression between this signature and expression profiles from tumours treated with the same drug to develop a prediction model. This information can then be used to predict the likelihood of response to a specific chemotherapeutic regimen in patients. In a phase II study (SWOG S1314), the ability of COXEN scores to predict response to NAC was assessed in 167 patients with urothelial bladder cancer randomized to receive either of two cisplatin-based NAC regimens — dose-dense MVAC or gemcitabine–cisplatin145. COXEN scores for each treatment regimen did not predict response, but in a pooled analysis incorporating both treatment arms, the score for gemcitabine–cisplatin showed a significant association with pathological downstaging (P = 0.02). The authors concluded that these findings supported the clinical utility of predictive markers derived from in vitro drug responses, and that further studies of the potential of COXEN are warranted.
The usefulness of preclinical models and in silico approaches for predicting cancer drug responses is often limited by the validity of in vitro models and by the availability of data to train algorithms, respectively. To address these limitations, a protein–protein interaction (PPI) network data from the STRING database using pharmacogenomic data from 3D colorectal cancer (n = 19) and bladder cancer (n = 9) organoids was developed to build a ML framework to identify biomarkers that could predict drug responses in patients146. This approach is based on the idea that genes associated with similar phenotypic outcomes tend to be in close association in PPI networks147,148, and by extension, biomarkers of response to a specific drug might also be in close proximity in interaction networks. Biomarkers identified through the ML model built from this integrated approach were able to predict OS in patients with colorectal cancer (P = 0.014) or bladder cancer (P = 0.01) receiving 5-fluorouracil or cisplatin, respectively. Conversely, ML models that used only gene expression data did not predict survival as strongly as this integrated approach. The results from this study highlight the need for appropriate in vitro models for drug screening, the benefits of network analysis in identifying strong predictors of response, and the power of network-based ML models for efficient prediction of patients who are likely to respond, or not, to a specific drug treatment149.
Other groups have explored the use of DL approaches to characterize changes in bladder tumours from CT imaging with the aim of predicting response to NAC. In one study, a CNN was used to extract information from radiological images of patients with bladder cancer before and after chemotherapy to identify features that predict response to therapy150. In this study, the DL approach achieved comparable efficacy for prediction of complete response to that of two expert radiologists, but the authors concluded that prospective validation in large patient cohorts was necessary before clinical implementation.
Prostate cancer
Prostate cancer presents a number of challenges for clinicians including the absence of definitive symptoms, variable disease course, a lack of accurate risk assessment and a lack of curative treatments. Assessment of serum PSA level and the digital rectal examination are mainstays of diagnosis, whereas surgery, radiation and inhibition of androgen action have been the major therapeutic strategies for decades. Our understanding of multiple aspects of prostate cancer biology, progression and treatment have improved as a result of the incorporation of bioinformatics analysis into both preclinical and clinical evaluation.
Classification and prognosis
The use of bioinformatics to analyse global mutation and expression profiles of prostate cancer stretches back over 20 years. In one of the earliest studies, microarray analysis was used to identify differentially expressed genes between benign prostatic hyperplasia (BPH) tissue, prostate cancer and adjacent non-cancerous tissue. The findings were subsequently validated through immunohistochemistry and the expression profiles were found to be associated with clinical outcomes22. This analysis highlights the utility of combining large-scale gene expression and protein profiles with existing pathology and clinical data to identify potentially informative disease biomarkers. Gene expression profiles have also been used to explore cancer-causing genetic events. In an early application of Oncomine151, the investigators reasoned that chromosomal rearrangements resulting in overexpression of genes would be detectable in DNA microarray profiles with an analytical approach that captured deviation of expression profiles from the median. In this study, a bioinformatics method called cancer outlier profile analysis (COPA) — looking for outlier gene expression — was developed and applied to >10,000 microarray experiments from Oncomine, leading to the identification of robust outlier profiles for the ETS family transcription factors ERG or ETV1 in six separate prostate cancer studies. ERG and ETV1 overexpression was mutually exclusive across a number of prostate cancer datasets, consistently with oncogenic translocations in other cancer types. The absence of ERG and ETV1 amplification despite transcript overexpression led the investigators to explore the possibility of DNA rearrangements as a potential explanation for this outlier expression. Further characterization of 5′ transcripts for ERG and ETV1 uncovered fusion with the prostate-specific gene TMPRSS2. The presence of TMPRSS2–ERG and TMPRSS2–ETV1 gene fusions was validated in specimens from patients with both localized and metastatic prostate cancer. The ERG fusion is one of the earliest genetic alterations in prostate cancer and occurs in >50% of patients152,153. Thus, the discovery of this genomic event led to the identification of other genomic fusion events, with bioinformatics techniques having a major role, and to the use of fusion events as diagnostic and prognostic biomarkers for prostate cancer154.
In another study, unsupervised hierarchical clustering of copy number alterations was used to identify six subtypes of prostate cancer that showed significant differences in the likelihood of biochemical recurrence (BCR) between the minimally altered cluster 2 tumours and the highly altered cluster 5 tumours (P < 0.005)155. In this study, the majority of metastatic samples were found in clusters showing a high extent of copy number alterations (clusters 5 and 6); moreover, samples in cluster 5, in which copy number alterations were observed across the genome, were associated with a higher risk of BCR than samples in cluster 6 (P < 0.05), in which alterations were restricted to chromosomes 7 and 8.
Since the publication of TCGA for primary prostate cancer29, several classification schemes have been developed to assess prostate cancer prognosis, including the PCS156 and PAM50 (ref. 157). The PCS classifier is a 37-gene diagnostic panel obtained from transcriptome data from >2,000 prostate cancer samples that enabled identification of three distinct subtypes of prostate cancer — PCS1, PCS2 and PCS3 — which differ in pathway activation and prognosis, with PCS1 being associated with the poorest metastasis-free survival (MFS)156. PAM50, a 50-gene classifier, was developed originally in the context of breast cancer103 and was subsequently applied to prostate cancer samples, leading to classification of prostate cancer into three subtypes — luminal A, luminal B and basal — which differ in prognosis, with luminal B-type tumours showing the worst outcomes in terms of both MFS and disease-specific survival157. Although prostate cancer does not have a clear immunohistochemically recognizable basal subtype, expression of TP63 and cytokeratins (such as CK5 and CK14) was shown to be higher in the basal subtype than in luminal subtypes, enabling the identification of tumours with a basal nature156,158,159. A direct comparison between the PCS and PAM50 classifiers was carried out using tumour samples from ~10,000 patients, and some consensus in terms of clinical outcome was reported between PCS1 and luminal B subtypes, between PCS2 and luminal A subtypes, and between PCS3 and basal subtypes160. Notably, PCS provided a better classification of the expression of luminal and basal marker genes than PAM50.
Similarly to gene signatures, the cell cycle progression (CCP) score developed for breast tumours161 has also been incorporated into a RT-qPCR assay to determine prostate tumour cell proliferation and aggressiveness162,163. In one study including patients with either localized prostate cancer who received conservative treatment or patients who had undergone radical prostatectomy, the CCP score enabled prediction of clinical outcomes, including BCR following prostatectomy, and time to death in patients undergoing transurethral resection of the prostate162. In a follow-up study, the CCP score was the strongest independent predictor of death in patients diagnosed with prostate cancer by needle biopsy164. Based on this prognostic ability, this assay is now commercially available as the Prolaris test from Myriad Genetics. The Decipher is a genomic classifier originally developed to estimate the risk of distant metastasis and includes 22 transcriptomic biomarkers identified by random forest classification based on differential mRNA expression in prostatectomy specimens165. This classifier was validated in a patient cohort at risk of progression and was shown to be the major predictor of metastasis in a multivariate analysis166. In subsequent studies, the Decipher GC was shown to predict risk of metastasis following prostatectomy, based on the analysis of biopsy material167. Lastly, the OncotypeDx genomic prostate score (GPS) includes a 17-gene signature to predict aggressive prostate cancer at the time of diagnosis to inform treatment decisions (for example, to assist in the choice of definitive treatment versus active surveillance)168. The utility of the GPS to predict the risk of distant metastasis and death from prostate cancer following radical prostatectomy has also been explored169,170. In one study, GPS was calculated from archival diagnostic biopsy samples from 279 men with a median follow-up time of 9.8 years who underwent radical prostatectomy, and a strong association was observed between GPS and both time to metastasis and time to death from prostate cancer. None of the 31 patients with low-risk or intermediate-risk disease and a GPS of <20 developed metastases or died from prostate cancer during the follow-up period169. A similar outcome was observed in another study including 428 patients who received radical prostatectomy with a follow-up time of 20 years. In this study, a GPS of <20 was associated with a low risk of either distant metastases or death from prostate cancer, whereas the risk for either outcome was increased substantially in patients with a GPS of >40 (ref. 170). The authors concluded that the incorporation of GPS into existing models could improve the estimation risk of distant metastasis and prostate cancer-specific mortality at 20 years compared with using only clinical factors such as grade, stage and PSA level, but that prospective studies are necessary for validation. Taken together, results from these studies show that genome classifiers seem to have most clinical benefit in the setting of disease classified as low risk at the time of diagnosis, for which these tools can aid in the decision to undergo active surveillance versus definitive treatment, or might help identify men at risk of aggressive disease. Current limitations include the retrospective nature and the small sample sizes of studies in which genomic classifiers were assessed, as well as the high cost171,172.
Prediction of treatment response
In addition to tumour classification, gene classifiers have also been assessed for their ability to inform treatment decisions in prostate cancer156. In one study the PCS classifier was used to profile circulating tumour cells (CTCs) from patients with CRPC by analysing scRNA-seq data from 77 CTCs from 13 patients. Two groups of CTCs were identified based on low versus high expression of genes enriched in the PCS1 subtype (the subtype associated with the poorest MFS). Moreover, the patients who underwent disease progression on enzalutamide treatment showed greater enrichment of PCS1-like genes than patients who did not show disease progression. The authors concluded that PCS has the potential to subtype individual tumours using both tissue and liquid biopsy (CTC-based) approaches. Published classifiers or signatures developed in-house were also used in different studies to predict response to androgen deprivation therapy (ADT)157,173,174. Patients with the luminal B subtype (identified as having the worst prognosis according to the PAM50 classifier) showed a better response to postoperative ADT than patients with non-luminal B subtype tumours157. In another study173, genomic profiling was used to identify 49 genes differentially expressed in prostate tumours between patients who did or did not receive ADT, which provided a robust signature of response to treatment. In this study, a high ADT response signature predicted response to adjuvant ADT, identifying patients who were likely to respond or not to therapy. The 49 genes profiled in this study were distinct from those in the PAM50 classifier, but the authors noted that tumours of luminal B subtype tended to have high ADT response signature values, in agreement with the observation that luminal B subtype tumours have an improved response to ADT157. The Decipher GC classifier was shown to be predictive of response to ADT in combination with the non-steroid anti-androgen apalutamide174. In the SPARTAN trial, gene expression data were obtained from primary tumour specimens from 233 patients with non-metastatic CRPC (nmCRPC) and used to define Decipher GC score and assign basal or luminal subtype score175. Patients were considered at high risk (GC >0.6) or low risk (GC ≤0.6) of developing metastases based on GC scores. Among men with nmCRPC receiving ADT plus placebo, patients with high GC scores had significantly shorter MFS than patients with low GC scores (P = 0.01). However, patients with high and low GC scores showed comparable MFS when treated with ADT plus apalutamide (P = 0.75). Notably, patients with high-risk GC scores had significantly longer MFS (P < 0.001) and OS (P = 0.03) after treatment with ADT plus apalutamide than patients with high GC scores receiving ADT plus placebo. Patients with low-risk GC scores also showed increased MFS following ADT plus apalutamide (P = 0.04), but the treatment effect in patients with high GC scores was larger, highlighting the potential benefit of apalutamide plus ADT in this patient subgroup. Lastly, patients with luminal subtype tumours had more favourable long-term outcomes following treatment with ADT plus apalutamide than patients with basal subtype tumours. These findings show the benefit of molecular subtyping using both the Decipher GC and basal–luminal subtype scores in identifying patients who are likely to benefit most from apalutamide treatment.
Gene expression profiles have shown the ability to define discrete molecular subtypes in retrospective analyses of prostate cancer. However, identifying specific molecular attributes that enable prediction of lethal disease remains challenging. To address this challenge, a biologically informed NN of genes, pathways and processes relevant to prostate cancer — P-NET — was developed to facilitate clinical predictions and improve translational benefit176. Using P-NET, molecular information such as copy number data and mutation status for an individual patient is analysed in the context of existing biological information relevant to prostate cancer extrapolated from datasets in the Reactome pathway knowledgebase177. P-NET was trained and tested using data from >1,000 prostate cancer samples, and the utility of this network was confirmed by the fact that known genes implicated in CRPC such as PTEN, TP53 and AR were highly predictive. P-NET was shown to be superior to existing ML models in predicting metastatic versus primary CRPC176. P-NET also enabled the identification of novel genes associated with disease progression such as MDM4, which was found to be amplified in tumour samples and associated with resistance to the anti-androgen enzalutamide in in vitro analyses. Sensitivity of prostate cancer cells to the MDM4 inhibitor Ro-5963 was also shown, highlighting the ability of P-NET to identify vulnerabilities for which inhibitors already exist, and emphasizing the translational potential of this strategy.
Machine learning in prostate cancer
Classification and molecular subtyping of tumours has been a major focus in genitourinary oncology for the past decade, but ML and AI approaches have been used in urology for considerably longer. In one of the earliest studies178, an ANN was built using 14 variables including multiple parameters related to PSA level, data from digital rectal examinations, and ultrasound measurement of the prostate to predict the result of a prostate biopsy. The ANN enabled the identification (preoperatively) of patients who were likely to experience recurrence with 90% accuracy, indicating that NN might improve decision-making in clinical management of prostate cancer.
The evaluation of tumour grade and stage is central to cancer diagnosis and prognosis; thus, histopathological images have been increasingly incorporated into ML and AI algorithms. Advances in digital pathology and computer-assisted prostate cancer diagnosis have facilitated the development of tools for prostate cancer detection and grading, such as computer-aided diagnosis systems for quantitative image analysis enabling automated accurate and quantitative detection of prostate lesions179. The resulting availability of extensive digital input data has enabled development of ML algorithms to enhance prostate cancer diagnosis, and multiple original papers and review articles describing the application of AI and ML approaches to prostate cancer diagnosis and prognosis have emerged. In this section, just a selection of studies showing the evolution of informatics tools and approaches as applied to crucial questions in the field are discussed. In-depth discussion on clinical applications of AI and ML in prostate cancer, as well as radiomics and MRI for prostate cancer detection is outside the scope of this Review, and has been covered elsewhere1,180,181,182,183.
In 2012, a boosted Bayesian multi-resolution system was developed, and enabled the deconstruction of a whole-slide image into a series of images at different resolution levels184. Areas defined as cancer by the classifier at low resolution were then flagged for further investigation with an increased resolution. Based on this approach, the authors described acceptable classification at different resolution levels, with AUCs of 0.76–0.84 (comparing highest versus lowest resolution), as well as a substantial decrease in computational time required for the analysis compared with analysis obtained only at the highest resolution. The authors concluded that this approach provided an efficient and automated tool to identify areas of cancer in prostate core biopsies as a precursor to determination of Gleason grade.
Gleason grading of prostate tumours suffers from substantial inter-observer variability potentially resulting in either over-treatment or under-treatment185. In one study, this issue was addressed by incorporating histopathological annotations from multiple experts to train and validate a computer-aided diagnosis system186. In several other studies, multiple CNNs were used to address the challenge of accurate Gleason grading14,187,188,189, all of which showed excellent performance in the discrimination of benign from cancer tissue, as well as Gleason grading accuracy in external validation (AUC 0.85–0.99). For example, in one of these studies, the performance of the DL system was better than that of relatively inexperienced pathologists (<15 years of experience; two-sided permutation test, P = 0.036) and was comparable to that of pathologists with >15 years of experience (two-sided permutation test, P = 0.96)14. Moreover, results from several studies supported the possibility of incorporating AI algorithms (such as Galen Prostate, which serves as a second read quality control system, and Paige Prostate) into routine clinical practice to support clinical decision-making,188,190,191,192.
BCR after radical prostatectomy is associated with increased risk of metastasis and disease-specific mortality193, highlighting the need for efficient identification of patients who are at high risk of recurrence. Various nomograms for prediction of BCR have been developed, but many limitations exist, including the number of variables that can be assessed and the length of follow-up monitoring. Thus, ML approaches have been assessed to address these limitations194,195,196. In one study in which the performance of supervised ML algorithms in predicting BCR following radical prostatectomy was compared with that of conventional nomograms196, BCR at 5 years using ML algorithms yielded AUC values of 0.894 (naive Bayes), 0.888 (random forest) and 0.855 (SVM), showing a better performance than that of standard nomograms (AUC 0.749–0.799). Interestingly, the authors observed that ML approaches were comparable to standard regression analysis in predicting BCR, but argued that ML approaches enable the incorporation of additional variables such as genomics or MRI data that could not be added to a standard regression model.
Dissemination of tumour to lymph nodes is associated with increased disease severity and an increased risk of cancer-specific mortality197. DL approaches have been used to predict local metastasis based on primary prostate tumour histology. Existing methods to predict the likelihood of lymph node metastasis typically rely on clinical information such as PSA level, biopsy findings such as percentage of positive cores and Gleason grading, and MRI metrics198,199. In 2021, the ability of CNN to improve risk prediction of lymph node metastasis over existing models was reported200. H&E-stained histological sections from primary prostate tumours (n = 218) obtained at radical prostatectomy for which lymph nodes were also available were used to train a CNN to detect morphological patterns that predict metastasis. The CNN-based algorithm was able to predict lymph node metastasis with an improved AUC compared with the established Memorial Sloan Kettering Cancer Center nomogram (AUC 0.68 versus 0.63, respectively)200. Together with lymphovascular invasion, the CNN-based prediction probability was also an independent predictor of lymph node metastasis in multivariate analysis. Based on these observations, the authors concluded that implementation of a CNN has potential for assessment of lymph node metastasis risk in prostate cancer, but that external validation is required before clinical use.
Renal cell carcinoma
Analysis of discrete histological subtypes of RCC by TCGA research network — including clear cell RCC (ccRCC) (n = 446), pRCC (n = 161) and chRCC (n = 66) — showed that specific molecular subtypes are associated with clinical outcomes, which has implications for treatment31,32,201. For example, the hypermethylated CpG island methylator phenotype (CIMP) subtype of pRCC showed early onset and was associated with poor OS. Subsequent integrated analysis of CNV, mRNA, miRNA and long non-coding RNA profiles confirmed findings obtained from the original genomic analysis of discrete histological subtypes, and also enabled comparison across all RCC samples in TCGA, including those excluded from initial characterization owing to incorrect histological classification but that were subsequently reclassified for inclusion in the analysis (n = 843)202. This comprehensive characterization showed features associated with reduced survival in all RCC subtypes, such as the presence of a TH2 cell immune signature and DNA hypermethylation, as well as features unique to each subtype. CIMP pRCC, previously shown to be associated with the worst survival among pRCC subtypes, showed the poorest survival also among all RCC subtypes. Within chRCC, a subset of tumours were considered metabolic outliers, with diminished expression of Krebs cycle, ETC and AMPK genes, but increased expression of ribose metabolism genes; these so-called metabolically divergent chRCC were high-stage tumours showing DNA hypermethylation, and were associated with much worse OS than other chRCC. Gene signatures characteristic of immune cell infiltration, particularly TH2 cells, were elevated in ccRCC compared with most pRCC and chRCC samples, and were associated with decreased survival, although CIMP pRCC and metabolically divergent chRCC also showed immune cell signature enrichment. Conversely, the TH17 cell signature was positively associated with increased survival in ccRCC and chRCC. Taken together, this extensive analysis uncovered both shared and unique characteristics to inform prognosis and subtype-tailored management of patients with RCC203.
Associations between transcriptomic and/or genomic features in RCC and response to treatment have been explored in several studies204,205,206,207,208. In one study, a 16-gene prognostic assay and associated recurrence score was developed to predict recurrence of ccRCC following surgery205. The panel consisted of cancer-specific genes relevant to ccRCC biology including genes associated with vascular function, cell division, immune response and inflammation. The recurrence score was a strong predictor of recurrence and survival, and could identify subgroups of patients with divergent recurrence risk, from very low to high. This recurrence score assay was subsequently validated in patients from the S-TRAC trial, in which sunitinib was assessed as adjuvant treatment in patients at high risk of recurrence of RCC following nephrectomy, and improved disease-free survival was observed in patients with high-risk RCC209. In this study205, high recurrence scores were significantly associated with time to recurrence and disease-free survival in patients receiving placebo (P < 0.001), providing independent prognostic information in addition to conventional clinical metrics of tumour stage, node involvement and metastasis.
ccRCC is characterized by angiogenesis and high immune cell infiltration, and treatment with anti-VEGF pathway inhibitors and immune checkpoint blockade targeting PDL1 have both led to improved outcomes in some patients with RCC. To gain insights into the heterogeneity in therapy response to these agents, a multiomics approach was applied to patient samples from the IMmotion151 trial (in which treatment with atezolizumab plus bevacizumab was compared with sunitinib in patients with metastatic RCC210). Analysis of RNA-seq data, genomics data, PDL1 staining, variant histology, and clinical data on >800 RCC tumour samples from the IMmotion151 trial yielded seven molecular subtypes206: angiogenic–stromal; angiogenic; complement–ω-oxidation; T-effector–proliferative; proliferative; stromal–proliferative; and small nucleolar RNA (snoRNA). Subtypes showed distinct transcriptomic profiles, genomic alterations, PDL1 status and the presence of sarcomatoid features, and were associated with differing clinical outcomes. The angiogenic–stromal and angiogenic clusters were associated with increased PFS regardless of treatment, whereas the stromal–proliferative cluster was associated with reduced PFS in both treatment groups. Atezolizumab plus bevacizumab was associated with improved objective response rate in patients in the T-effector–proliferative, proliferative, and snoRNA clusters compared with treatment with sunitinib. Collectively, these analyses highlight crucial features of each cluster that could explain the response — or not — to treatment, and provide insights into new targets for therapeutic intervention.
Testicular germ cell tumours
Molecular characterization of testicular cancer by a group within TCGA network focused on testicular GCT including pure seminoma and non-seminomatous GCT (NSGCT). Tumour tissue samples (n = 137) were analysed using exome sequencing, single-nucleotide polymorphism analysis, RNA-seq, DNA methylation and RPPA analysis30. In agreement with observations in multiple other tumour types, discrete histological GCT subtypes were associated with different molecular characteristics. Driver mutations were rare and restricted to seminomatous tissue, with mutations in KIT being among the most frequent. Widespread lymphocyte infiltration and a lack of global DNA methylation had been observed in prior analyses of seminomas, but this integrated analysis showed that these features are more evident in KIT-mutant tumours than in other subtypes. Features enriched in NSGCT included several miRNAs such as miR-371 and miR-375. In principle, these features could be used in future studies either alone or together to identify patients who could avoid chemotherapy or aggressive surgical intervention, in turn minimizing treatment-related morbidity.
Penile and urethral cancer
Rare genitourinary malignancies such as penile and urethral cancers are not included in TCGA, but have also benefited from genomics and bioinformatics analysis carried out in several studies211,212,213,214,215. The incidence of penile cancer has been increasing as a result of exposure to human papillomavirus, but treatment options for this cancer are limited owing to an insufficient understanding of molecular drivers of the disease. To address this limitation, the first targeted genomic profiling of 60 penile cancer specimens was performed211 using the Oncomine Comprehensive Panel, a focused assay that enables the interrogation of somatic variants relevant to solid tumours and is based on rigorous bioinformatics analysis of >700,000 samples216. This study showed that clinical stage, lack of p16 expression and amplification of CCND1 and MYC are associated with reduced PFS or disease-specific survival. Additionally, amplification of EGFR was observed in several specimens, but the expression of the EGFR protein was discordant, which might have important implications for EGFR-targeted treatment. In another study, NanoString analysis of ~700 cancer-relevant genes was used to define a prognostic gene expression signature in patients with advanced penile SCC (n = 25) receiving cisplatin-based chemotherapy212. With this approach, MAML2, KITLG and JAK1 were shown to be associated with poor outcomes, with MAML2 significantly associated with a poor OS (P = 0.0003) in multivariate analysis. In a study in which whole-exome sequencing of 34 penile SCCs was carried out, the findings were integrated with TCGA data on SCCs from other organs including bladder, cervix, head and neck, oesophagus and lung to identify convergent pathways214; in this study, two mutational patterns were identified, characterized by APOBEC activity (MP1) and defective DNA mismatch repair (MP2), respectively. Enrichment of MP1 was associated with increased TMB and worse survival than MP2 (P = 0.0039). Additionally, enrichment of genes of the Notch pathway was observed in the majority of samples, in agreement with findings reported for head and neck SCCs217.
With regard to urethral cancer, a comprehensive genomic profiling of 127 metastatic urethral carcinoma specimens including urothelial, squamous, adenocarcinoma and clear cell subtypes was performed to characterize genomic alterations, TMB and microsatellite instability status215. Important findings from this analysis included frequent occurrence of genomic alterations in PIK3CA across all tumour subtypes and increased TMB and PDL1 protein staining levels in urothelial and squamous tumour subtypes. These findings show the ability of genomic profiling to highlight novel actionable targets in urethral cancer such as PIK3CA and ERBB2, for which targeted therapies exist. Elevated TMB and positive staining for PDL1 in a subset of patients also predicted responsiveness to immune checkpoint inhibitors.
Taken together, these studies provide strong rationale for the use of previously untested therapeutic interventions in penile and urethral cancers based on genomic alterations and mutational patterns identified through tumour genomic profiling.
Bioinformatics in benign urological disorders
In addition to a wide application in genitourinary oncology, bioinformatics approaches, including AI and ML, have also been implemented in benign conditions affecting the urinary tract, including functional disorders2.
Molecular classification of disease
The application of molecular classifiers to benign urological disorders is relatively infrequent compared with the use of these tools in urological oncology, owing in part to the lack of genomic alterations underlying the majority of benign conditions. However, transcriptional and other molecular signatures associated with discrete disease states are starting to be investigated in some studies218. Hierarchical clustering was used to classify patients with non-Hunner’s lesion IC–BPS on the basis of mRNA and miRNA sequencing of bladder biopsies219. In this study, mRNAs and miRNAs differentially expressed between patients with non-Hunner’s lesion IC–BPS versus individuals without BPS suggested enrichment of signalling pathways associated with smooth muscle proliferation and contraction and peripheral nervous system re-organization in patients with IC–BPS, but limited enrichment of immune-related pathways (neutrophil chemotaxis and IFNγ-mediated signalling) in these patients. Conversely, the analysis of transcriptional profiles in HIC–BPS versus individuals without BPS showed enrichment in genes associated with immune cell infiltration220, suggesting that molecular classification can be applied to benign disorders and might have diagnostic utility.
The power of single-cell analysis in benign urology has been shown in several studies. In 2018, the first cellular atlas of the normal human prostate at single-cell resolution was provided44. Using scRNA-seq analysis, the presence of basal, luminal and neuroendocrine epithelial cells within the prostate was confirmed, but two cell types not previously described in this tissue — club and hillock — were also identified. This study provided an important resource for studies aimed at understand the cellular basis of prostate disease. In another study by the same group, stromal cell populations within the prostate were analysed using scRNA-seq, which led to the identification of two novel fibroblast subtypes with distinct anatomical localization within the prostate, namely interstitial fibroblasts and peri-epithelial fibroblasts221. Interstitial fibroblasts were present in interstitial spaces between glands, whereas peri-epithelial fibroblasts were found adjacent to the epithelium of the urethra, glands and ejaculatory ducts. Together, results from these studies show the potential for single-cell analysis to discover previously unidentified cell types.
Benign prostatic hyperplasia
BPH occurs frequently in ageing men, and is associated with the emergence of obstructive lower urinary tract symptoms (LUTS). The molecular understanding of BPH pathogenesis has lagged behind that of prostate cancer, but results from two studies have provided insights on the genomic and transcriptomic underpinnings of this disease222,223. In one of these studies, BPH samples (n = 37, 35 of which were <100 cm3), normal prostate samples (n = 19) and BPH stromal nodules (n = 9) were analysed using RNA-seq222. Unsupervised clustering of transcriptome data enabled to distinguish among the three groups of samples and to identify features characteristic of secretory epithelium, stroma and immune cells that differed among the groups. Stromal markers were enriched in both BPH and stromal nodules compared with normal prostate samples, whereas immune cell markers were enriched only in BPH stromal nodules compared with BPH or normal prostate samples. In this study, a 65-gene stromal signature was identified, increased expression of which showed significant positive correlation with the IPSS bother score (P = 0.02), a validated measure of patient urinary symptoms. Conversely, high expression of an androgen receptor and secretory epithelium signature was not associated with the IPSS bother score, suggesting that stromal features are associated with functional consequences.
In a separate study, genomic, epigenomic and transcriptomic characterization of BPH (n = 18) and matched control tissue samples was performed223, but this analysis focused on prostates >100 cm3. Control tissues were from men undergoing radical prostatectomy for prostate cancer who did not have BPH. In this study, integration of transcriptomic and epigenomic profiles led to the identification of two distinct subtypes enriched for signatures of stromal elements (BPH-A) or dysregulated metabolism (BPH-B). Further analysis of transcriptome data using Connectivity Map enabled the identification of therapeutic compounds associated with mTOR signalling inhibition in the BPH-A subgroup. This evidence prompted assessment of the effect of mTOR inhibition on prostate size in patients receiving mTOR inhibitors for conditions unrelated to BPH or prostate cancer. This analysis was retrospective in nature, but some patients did show a reduction in prostate size with mTOR inhibitor treatment, leading the authors to conclude that a subset of BPH is influenced by mTOR signalling.
Urolithiasis
The application of ML strategies to urolithiasis was one of the earliest applications of AI in non-cancer urology224. In one study, a NN was trained using multiple clinical parameters including prior stone occurrence, treatment and metabolic status, and was used to identify risk factors that could predict stone recurrence225. This model showed sensitivity and specificity of 91% and 92%, respectively, with an AUC of 0.964, which exceeded the performance of conventional analysis. In many subsequent studies, AI algorithms have been applied to different aspects of urolithiasis pathogenesis including stone composition226, prediction of postoperative outcomes227,228, and improvement in the efficacy of extracorporeal shock wave lithotripsy229. An ANN was developed to predict postoperative outcomes following percutaneous nephrolithotomy (PCNL), including the stone-free rate, as well as complications such as the need for blood transfusion or additional procedures227. The algorithm showed accuracies ranging from 81% to over 98% depending on the postoperative parameter measured. In a subsequent study from the same group, SVM was compared with existing systems, including Guy’s stone score and the CROES nomogram230, and the ML algorithm showed an accuracy of 80–95% in predicting outcomes of PCNL. Furthermore, the AUC for prediction of stone-free status was 0.915 using the ML algorithm compared with 0.615 for Guy’s stone score and 0.621 for the CROES nomogram. Thus, the SVM model outperformed Guy’s stone score and the CROES nomogram, showing the potential for AI to enhance clinical management of stone disease.
Urodynamic analyses
UDS is part of the standard clinical work-up for the diagnosis of functional urological disorders. Interpretation of UDS is not standardized, leading to a lack of consensus among providers regarding specific findings231,232. In two studies, mathematical modelling and ML were implemented for objective detection of detrusor overactivity in UDS233,234. In one of these studies, manifold learning and dynamic time warping pattern-matching algorithms were applied to patterns of detrusor overactivity measured in paediatric patients undergoing UDS from 799 studies233. In this study, the sensitivity and specificity of detrusor overactivity detection were ~77% and 81%, respectively, suggesting that a ML approach has the potential to standardize the interpretation of discrete findings in UDS. In another study, the inclusion of additional parameters such as abdominal pressure slightly improved accuracy and specificity in the detection of detrusor overactivity234, but the two approaches showed comparable performance.
Urinary tract infections
Urinary tract infections (UTIs) are the most common infection in women, affecting >50% of women during the lifetime, but are also a major health challenge in both adults and children235,236. Thus, efficient diagnosis and treatment are crucial to minimize the health-care burden of these infections. Uncomplicated UTIs arise in the absence of comorbidities or urological abnormalities, whereas complicated UTIs occur in patients with a history of UTI or urological conditions such as stones, neurogenic bladder or diabetes237,238,239. AI approaches have been implemented to facilitate rapid diagnosis of UTI, to identify causative agents, and to predict the potential for development of resistance.
In one study, six ML algorithms to predict UTI based on medical history and other clinical variables were developed and applied to >80,00ient encounters for which urinalysis and symptoms of UTI were known240. AUC was 0.826–0.904 among all models, with the extreme gradient boosting (XGBoost) algorithm showing the best performance. XGBoost was more sensitive and specific in terms of UTI diagnosis than provider assessment. This study was retrospective in nature, but showed the potential for AI to rapidly provide or exclude UTI diagnosis and, in turn, to improve treatment decisions.
In another study, four ML algorithms — decision tree, SVM, random forest and ANN — were applied to a dataset including symptoms, clinical variables and laboratory results from patients presenting with UTI241. In this study, the classification accuracy ranged from 93% to 98%, with sensitivity and specificity of ~95–98% and ~86–100%, respectively. The ANN showed the most robust performance among all the classifiers in terms of both positive predictive value and negative predictive value, and supported the potential of AI in the context of UTI diagnosis.
In addition to diagnosis, identification of specific microbial species in UTI is important to guide treatment with antimicrobial agents. Current culture-based methods of urinalysis for suspected UTI are relatively slow and are based on the ability to culture bacteria242,243. In one study244, three ML classifiers — naive Bayes, BayesNet and Hoeffding tree — were applied to mass spectra from both urine specimens and bacterial cultures to define a signature including 82 peptides that could be used to identify microbial species in clinical samples with high specificity and sensitivity. This peptide signature enabled researchers to identify the dominant bacterial species (among the 15 species found most commonly in the urine of patients with UTI) in 4 hours and without the need for bacterial culture.
In children presenting with a febrile UTI, ML has been explored as a strategy to predict the risk of recurrent UTI and vesicoureteral reflux (VUR)245, which, together, are associated with an elevated risk of renal scarring. Current treatment guidelines for this condition are controversial and would benefit from tools that enable objective risk assessment in children with UTI. In this study245, an optimal classification tree was applied to data from the RIVUR and CUTIE clinical trials, including children with VUR presenting with a first UTI over a 2-year period. ML enabled the probability that a child would have recurrent UTI-associated VUR to be estimated, facilitating appropriate treatment decisions at the individual level.
Urinary tract obstruction
AI was used >20 years ago to predict outcomes in children with ureteropelvic junction obstruction (UPJO)246. In this study, an ANN was constructed using data from 100 patients who underwent pyeloplasty and subsequent imaging to assess outcomes. The ANN showed 100% sensitivity and specificity with correct prediction of outcomes in all patients including the test set compared with conventional linear regression, which yielded sensitivity and specificity of 52–94%. Considering the small sample size, caution might be necessary in the interpretation of these findings. In another study247, a commercially available NN software was applied to data including demographics, renal features, urinalysis and UTI status of patients with UPJO diagnosed before birth, and showed a 75% sensitivity in predicting outcomes of pyeloplasty. In current practice, multiple variables are considered in assessing the need for re-intervention for UPJO following pyeloplasty, but prediction of patients who will require re-operation remains challenging, owing to the inherent variability in postoperative resolution of hydronephrosis. In a 2022 study, a model was developed to enable efficient prediction of the likelihood of re-operation for UPJO following pyeloplasty, including time to the subsequent procedure248. Application of this model showed that postoperative anteroposterior pelvic diameter (APD) was associated with the likelihood of cure and with time to re-intervention, with an increased APD at the second follow-up visit associated with reduced time to re-operation. Differently from other studies, the ML approach used in this study led to the identification of risk factors for re-operation, as well as of interactions between risk factors that might enable a personalized approach to predict success of pyeloplasty.
A challenge in functional urology is the co-occurrence of LUTS in a single patient, leading to a lack of clarity of treatment pathways. Pressure-flow studies are considered the gold standard for discriminating between LUTS consequent to obstruction or LUTS consequent to detrusor dysfunction, but are invasive, time-consuming and expensive for routine use249. The need for pressure-flow studies could be circumvented by predicting the outcome of pressure-flow studies from non-invasive test data, such as prostate volume measurements by imaging, uroflowmetry and symptom score questionnaires. In one study, the use of an ANN to predict bladder outlet obstruction in men with LUTS was compared with conventional regression models250. A variety of diagnostic data obtained non-invasively from 1,900 patients were analysed, and the ANN showed a sensitivity and specificity of 71% and 69%, respectively; linear regression provided equivalent results. Thus, the application of the chosen NN did not improve prediction of obstruction compared with conventional linear regression.
In several studies, ML was used to improve prediction of outcomes in boys with outlet obstruction induced by posterior urethral valves (PUVs)251,252,253. Bladder outlet obstruction secondary to PUVs increases the risk of renal damage, and, therefore, timely diagnosis of obstruction in children is of high importance. However, existing diagnostic procedures are often invasive and subject to substantial inter-observer variability, highlighting the need for improved testing with increased accuracy. An ANN was used to analyse data from non-invasive clinical work-up in boys with LUTS, a subset of whom were subsequently found to have PUVs251. In this study, the ANN showed an accuracy of ~93% in predicting late-presenting PUVs, with an AUC of 0.98, suggesting that this approach fulfils the criteria for accurate and timely diagnosis and, therefore, can guide appropriate treatment. In another study, data from ultrasound imaging in children with congenital anomalies of the kidney and urinary tract were used to build a classifier that would distinguish patients with PUVs from patients with clinically insignificant mild hydronephrosis252. The innovation in this study was in the use of multiple images in different planes to build a multi-instance classifier that showed increased performance compared with a classifier built on single images in a specific plane (accuracy 92.5%, sensitivity 87.3% and specificity 98.6% with the multi-view model incorporating both sagittal and transverse images, versus accuracy 90.4–91.2%, sensitivity 86.8–87.3% and specificity 94.5–96.0% with models incorporating either sagittal or transverse images). In another study, the potential of ML to facilitate personalized management of patients with PUVs based on prediction of progressive deterioration in renal function was highlighted253. In this study, data including demographics, estimated glomerular filtration rate, serum creatinine, imaging and the need for clean intermittent catheterization (CIC) were used to train a random survival forest model with the same variables used for standard Cox proportional hazards regression analysis. For each of the three clinically relevant end points considered — progression to chronic kidney disease, renal replacement therapy, and CIC — the ML model outperformed Cox proportional hazards regression analysis. The authors concluded that ML approaches fulfilled an important role in risk stratification and personalizing clinical management in this patient population.
Electronic health record or electronic medical record informatics in urology
Beyond the evaluation of cellular and molecular features of disease, AI and ML approaches have been at the forefront of advances in health informatics and attempts to harness data captured within electronic health records (EHRs) or electronic medical records (EMRs). Extraction of clinically meaningful information from medical records has required the development of AI tools in the form of NLP. For example, an NLP programme was implemented to extract relevant variables from pathology reports of patients undergoing prostate biopsy, and correctly identified >99% of patients with prostate cancer following biopsy254. In another study, researchers aimed to develop an NLP system to extract prostate pathology details from postoperative pathology reports and to compare the accuracy of this system with that of manual abstraction (used as a gold standard). The results showed that NLP and clinician-entered structured data elements (SDEs) had >90% accuracy (defined as percentage agreement with manual abstraction) for Gleason scores, margin status, extracapsular extension, seminal vesicle invasion, stage, and lymph node status. NLP and SDEs were also highly concordant (Cohen’s κ coefficient of at least 0.92) for all data elements extracted, and moderately-to-highly concordant with manual abstraction (Cohen’s κ coefficient of at least 0.84 for NLP versus manual abstraction, and at least 0.79 for SDEs versus manual abstraction)255. An NLP engine was developed to evaluate bladder cancer pathology reports within the Veterans Health Administration and applied to over 30,000 reports including >10,000 patients256. Successful retrieval of crucial pathological parameters was reported for 99% of patients using the NLP engine, with excellent accuracy reported for a majority of parameters compared with the gold standard of manual information abstraction by experts. In another study, NLP algorithms were applied to pathology reports of transurethral resection of bladder tumour. In one of these studies, an NLP algorithm enabled highly reliable automated extraction of data on tumour grade and stage, including the extent of involvement of muscularis propria257, compared with manual review of reports. To maximize the utility of EHRs and EMRs for both clinical care and research across urology, areas for improvement include the use of standardized language across health-care institutions, ongoing education of medical providers to ensure structured data entry into electronic records, and the investment of adequate resources to ensure minimal disruption to clinical workflow while ensuring the richness of data records.
Challenges, solutions and future opportunities — best practices to use bioinformatics and ML
The use of bioinformatics and AI in urology has exploded in the past few years, particularly with the wealth of data emerging from omics and other high-throughput analyses. Many prediction models have emerged, a majority of which have been shown to be more accurate and provide less variable estimates of risk than predictions made subjectively; however, few if any of these methods are being used clinically or even cited in clinical guidelines, raising some concerns about the utility of these tools258. The reasons for low citation rates and low real-world use of these approaches are multifactorial, but some crucial issues have emerged. The first issues to consider are the type and amount of biospecimen available and the analyses for which these specimens are suitable; for example, biopsy specimens are appropriate for application of a multiplex PCR-based assay, but might be inappropriate for spatial proteomics or other image-based analysis. Another issue is biospecimen heterogeneity. Moreover, the time and cost of developing point-of-care tests or models (from discovery efforts to the development of Clinical Laboratory Improvement Amendments-approved products or pipelines), together with uncertain value over existing metrics as well as lack of reimbursement, are considerable disincentives to the use of bioinformatics in a real-world setting. Incorporation of necessary bioinformatics expertise and training within both research and clinical teams is also essential for effective clinical translation. Lastly, prospective evaluation of assays developed in retrospective patient cohorts, as well as validation of results in independent patient cohorts, are needed before clinical implementation. Estimates indicate that only ~10% of clinical prediction models across medicine have been validated externally259. To address this point, international challenges such as the Grand Challenge for medical image analysis have been launched by consortia to provide an opportunity for validation and verification of AI models and, in turn, to enhance reproducibility260. Several challenges in urology include PROSTATEx, to predict the clinical significance of prostate lesions from MRI data; PI-CAI, focused on the ability of AI to detect and diagnose clinically significant prostate cancer using MRI; PAIP, to develop an algorithm for automatic detection of perineural invasion in different tumour types; and Prostate cANcer graDe Assessment (PANDA), focused on prostate cancer grade assessment. PANDA is a histopathology competition based on >10,000 digitized prostate biopsies, which aims to promote the development of reproducible algorithms for Gleason grading that could be applied essentially worldwide261. These challenges can promote advances in the field, but some concerns have been expressed regarding the reporting of findings from challenges, including highly variable challenge design, lack of cross-comparison of results between different challenges, uncertain reproducibility, and insufficient reporting of data262; thus, caution is warranted in the interpretation of results from these challenges.
In general, results from systematic reviews show that the methodological conduct of ML-based prediction models in clinical medicine and clinical trials is variable263,264 and that increasing rigor is required before these models are incorporated into routine use.
Reporting standards is another issue that hampers the routine application of bioinformatics into clinical practice. Across studies in which models were developed, the availability of code was limited to <20% of models, the version control system (such as Github, Gitlab, etc.) was not implemented in a majority of papers, and in most cases, the definitive model was not completely presented. The lack of a version control system in AI can lead to inability to track changes, difficulty in reproducing results, loss of important information and security risks. To avoid these issues, a version control system for AI projects needs to be implemented. This implementation can include using tools such as Git or Subversion to track changes to code and data, as well as creating documentation and backups of important information. By implementing a version control system, AI teams can ensure the accuracy, reproducibility and security of models and datasets. These points emphasize the need for establishing minimum standards for use and reporting of bioinformatics and AI approaches, not just in urology but across all disciplines20. These standards include robust description of raw data such as input for model development, training and testing; justification of the model chosen; justification of performance metrics; transparency of code used; and external validation. In a 2021 study, a protocol for the development of reporting guidelines for AI-based and ML-based prediction models was described including crucial methodological details required to evaluate quality and validity, and to enable model users to assess the potential for bias265.
Bioinformatics research in most benign urological diseases is associated with distinctive challenges owing to idiosyncrasies of many of these conditions, such as the timescale over which benign disease develops. For example, bladder outlet obstruction secondary to prostatic enlargement might develop over decades. Thus, benign urological phenotypes can be defined as a continuous and complex spectrum of evolution from health to disease. Symptoms that are characteristic of overactive bladder such as urgency and frequency might change over time, as a condition might coexist with or transform into another form of dysfunction such as underactive bladder; additionally, these conditions could or could not lead to a debilitating clinical episode such as total organ failure. Moreover, in functional disorders, particularly those with a neurological component, whether any tissue should be biopsied for omics analyses is unclear. The fact that changes leading to benign urological diseases occur on a timescale of decades is associated with informatics challenges per se. This situation is further complicated by the fact that benign urological diseases commonly present as comorbid with other disease phenotypes such as obesity, diabetes, dyslipidaemia, hormonal imbalance, hypertension and metabolic syndrome, among others266. Collecting and modelling data from such a long temporal progression that is also affected by numerous environmental and lifestyle perturbations are exceptionally challenging in computational terms and will require innovative data integration methodologies for risk stratification and surveillance. One approach to address these challenges involves an increasingly rigorous phenotyping of individuals presenting with urological symptoms to improve clinical ‘phenomapping’, which is the stratification of different lower urinary tract dysfunction into aetiologically distinct subtypes, enabling bioinformatics to define a disease during a temporal trajectory.
Conclusions
Bioinformatics has enabled major advances in the understanding of urological disease, particularly in terms of molecular classification, prognostication, stratification and prediction of response to treatment for urological cancers and — to a lesser extent — benign urology. However, across the field of urology as a whole, adherence to best practices in ML, including transparency in reporting of algorithms and models, availability of code to ensure reproducibility by other investigators and external validation against independent datasets will be crucial to ensure that the urology field can truly benefit from the application of bioinformatics methodologies.
References
Goldenberg, S. L., Nir, G. & Salcudean, S. E. A new era: artificial intelligence and machine learning in prostate cancer. Nat. Rev. Urol. 16, 391–403 (2019).
Bentellis, I., Guerin, S., Khene, Z. E., Khavari, R. & Peyronnet, B. Artificial intelligence in functional urology: how it may shape the future. Curr. Opin. Urol. 31, 385–390 (2021).
Brodie, A. et al. Artificial intelligence in urological oncology: an update and future applications. Urol. Oncol. 39, 379–399 (2021).
Bzdok, D., Altman, N. & Krzywinski, M. Statistics versus machine learning. Nat. Methods 15, 233–234 (2018).
Esteva, A. et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542, 115–118 (2017).
Sidey-Gibbons, J. A. M. & Sidey-Gibbons, C. J. Machine learning in medicine: a practical introduction. BMC Med. Res. Methodol. 19, 64 (2019).
Lo Vercio, L. et al. Supervised machine learning tools: a tutorial for clinicians. J. Neural Eng. 17, 062001 (2020).
Eisen, M. B., Spellman, P. T., Brown, P. O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl Acad. Sci. USA 95, 14863–14868 (1998).
Jolliffe, I. T. & Cadima, J. Principal component analysis: a review and recent developments. Philos. Trans. A Math. Phys. Eng. Sci. 374, 20150202 (2016).
Rashidi, H. H., Tran, N. K., Betts, E. V., Howell, L. P. & Green, R. Artificial intelligence and machine learning in pathology: the present landscape of supervised methods. Acad. Pathol. 6, 2374289519873088 (2019).
Quinn, T. P., Nguyen, T., Lee, S. C. & Venkatesh, S. Cancer as a tissue anomaly: classifying tumor transcriptomes based only on healthy data. Front. Genet. 10, 599 (2019).
Yakimovich, A., Beaugnon, A., Huang, Y. & Ozkirimli, E. Labels in a haystack: approaches beyond supervised learning in biomedical applications. Patterns 2, 100383 (2021).
Eckardt, J. N., Bornhauser, M., Wendt, K. & Middeke, J. M. Semi-supervised learning in cancer diagnostics. Front. Oncol. 12, 960984 (2022).
Bulten, W. et al. Automated deep-learning system for Gleason grading of prostate cancer using biopsies: a diagnostic study. Lancet Oncol. 21, 233–241 (2020).
Marini, N., Otalora, S., Muller, H. & Atzori, M. Semi-supervised training of deep convolutional neural networks with heterogeneous data and few local annotations: an experiment on prostate histopathology image classification. Med. Image Anal. 73, 102165 (2021).
Doan, S., Conway, M., Phuong, T. M. & Ohno-Machado, L. Natural language processing in biomedicine: a unified system architecture overview. Methods Mol. Biol. 1168, 275–294 (2014).
Finne, P. et al. Predicting the outcome of prostate biopsy in screen-positive men by a multilayer perceptron network. Urology 56, 418–422 (2000).
Remzi, M. et al. An artificial neural network to predict the outcome of repeat prostate biopsies. Urology 62, 456–460 (2003).
Greener, J. G., Kandathil, S. M., Moffat, L. & Jones, D. T. A guide to machine learning for biologists. Nat. Rev. Mol. Cell Biol. 23, 40–55 (2022).
Chen, A. B. et al. Artificial intelligence applications in urology: reporting standards to achieve fluency for urologists. Urol. Clin. North. Am. 49, 65–117 (2022).
Thykjaer, T. et al. Identification of gene expression patterns in superficial and invasive human bladder cancer. Cancer Res. 61, 2492–2499 (2001).
Dhanasekaran, S. M. et al. Delineation of prognostic biomarkers in prostate cancer. Nature 412, 822–826 (2001).
Luo, J. et al. Human prostate cancer and benign prostatic hyperplasia: molecular dissection by gene expression profiling. Cancer Res. 61, 4683–4688 (2001).
Singh, D. et al. Gene expression correlates of clinical prostate cancer behavior. Cancer Cell 1, 203–209 (2002).
Dyrskjot, L. et al. Identifying distinct classes of bladder carcinoma using microarrays. Nat. Genet. 33, 90–96 (2003).
Blaveri, E. et al. Bladder cancer outcome and subtype classification by gene expression. Clin. Cancer Res. 11, 4044–4055 (2005).
Rhodes, D. R. et al. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia 6, 1–6 (2004).
The Cancer Genome Atlas Research Network Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507, 315–322 (2014).
The Cancer Genome Atlas Research Network The molecular taxonomy of primary prostate cancer. Cell 163, 1011–1025 (2015).
Shen, H. et al. Integrated molecular characterization of testicular germ cell tumors. Cell Rep. 23, 3392–3406 (2018).
The Cancer Genome Atlas Research Network. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499, 43–49 (2013).
The Cancer Genome Atlas Research Network. Comprehensive molecular characterization of papillary renal-cell carcinoma. N. Engl. J. Med. 374, 135–145 (2016).
Veldman-Jones, M. H. et al. Evaluating robustness and sensitivity of the NanoString Technologies nCounter platform to enable multiplexed gene expression analysis of clinical samples. Cancer Res. 75, 2587–2593 (2015).
Zheng, H. et al. Comprehensive review of web servers and bioinformatics tools for cancer prognosis analysis. Front. Oncol. 10, 68 (2020).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Chandrashekar, D. S. et al. UALCAN: an update to the integrated cancer data analysis platform. Neoplasia 25, 18–27 (2022).
Chen, M. M. et al. TCPA v3.0: an integrative platform to explore the pan-cancer analysis of functional proteomic data. Mol. Cell Proteom. 18, S15–S25 (2019).
Navin, N. E. The first five years of single-cell cancer genomics and beyond. Genome Res. 25, 1499–1507 (2015).
Azizi, E. et al. Single-cell map of diverse immune phenotypes in the breast tumor microenvironment. Cell 174, 1293–1308.e36 (2018).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Shaffer, S. M. et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature 546, 431–435 (2017).
Su, F. et al. Multimodal single-cell analyses outline the immune microenvironment and therapeutic effectors of interstitial cystitis/bladder pain syndrome. Adv. Sci. 9, e2106063 (2022).
Peng, L. et al. Integrating single-cell RNA sequencing with spatial transcriptomics reveals immune landscape for interstitial cystitis. Signal Transduct. Target. Ther. 7, 161 (2022).
Henry, G. H. et al. A cellular anatomy of the normal adult human prostate and prostatic urethra. Cell Rep. 25, 3530–3542.e5 (2018).
Karthaus, W. R. et al. Regenerative potential of prostate luminal cells revealed by single-cell analysis. Science 368, 497–505 (2020).
Luecken, M. D. & Theis, F. J. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol. Syst. Biol. 15, e8746 (2019).
Zurauskiene, J. & Yau, C. pcaReduce: hierarchical clustering of single cell transcriptional profiles. BMC Bioinforma. 17, 140 (2016).
Kiselev, V. Y. et al. SC3: consensus clustering of single-cell RNA-seq data. Nat. Methods 14, 483–486 (2017).
Xu, C. & Su, Z. Identification of cell types from single-cell transcriptomes using a novel clustering method. Bioinformatics 31, 1974–1980 (2015).
Clarke, Z. A. et al. Tutorial: guidelines for annotating single-cell transcriptomic maps using automated and manual methods. Nat. Protoc. 16, 2749–2764 (2021).
Young, M. D. et al. Single-cell transcriptomes from human kidneys reveal the cellular identity of renal tumors. Science 361, 594–599 (2018).
Zhang, Y. et al. Single-cell analyses of renal cell cancers reveal insights into tumor microenvironment, cell of origin, and therapy response. Proc. Natl Acad. Sci. USA 118, e2103240118 (2021).
Dong, B. et al. Single-cell analysis supports a luminal-neuroendocrine transdifferentiation in human prostate cancer. Commun. Biol. 3, 778 (2020).
Song, H. et al. Single-cell analysis of human primary prostate cancer reveals the heterogeneity of tumor-associated epithelial cell states. Nat. Commun. 13, 141 (2022).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Haghverdi, L., Buttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Setty, M. et al. Wishbone identifies bifurcating developmental trajectories from single-cell data. Nat. Biotechnol. 34, 637–645 (2016).
Schiebinger, G. et al. Optimal-transport analysis of single-cell gene expression identifies developmental trajectories in reprogramming. Cell 176, 928–943.e2 (2019).
Chen, S. et al. Single-cell analysis reveals transcriptomic remodellings in distinct cell types that contribute to human prostate cancer progression. Nat. Cell Biol. 23, 87–98 (2021).
Chen, Z. et al. Single-cell RNA sequencing highlights the role of inflammatory cancer-associated fibroblasts in bladder urothelial carcinoma. Nat. Commun. 11, 5077 (2020).
Wu, T., Wu, X., Wang, H. Y. & Chen, L. Immune contexture defined by single cell technology for prognosis prediction and immunotherapy guidance in cancer. Cancer Commun. 39, 21 (2019).
Tuong, Z. K. et al. Resolving the immune landscape of human prostate at a single-cell level in health and cancer. Cell Rep. 37, 110132 (2021).
Chen, W. J. et al. Heterogeneity of tumor microenvironment is associated with clinical prognosis of non-clear cell renal cell carcinoma: a single-cell genomics study. Cell Death Dis. 13, 50 (2022).
Bi, K. et al. Tumor and immune reprogramming during immunotherapy in advanced renal cell carcinoma. Cancer Cell 39, 649–661.e5 (2021).
Wang, L. et al. Myeloid cell-associated resistance to PD-1/PD-L1 blockade in urothelial cancer revealed through bulk and single-cell RNA sequencing. Clin. Cancer Res. 27, 4287–4300 (2021).
Moses, L. & Pachter, L. Museum of spatial transcriptomics. Nat. Methods 19, 534–546 (2022).
Rodriques, S. G. et al. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Williams, C. G., Lee, H. J., Asatsuma, T., Vento-Tormo, R. & Haque, A. An introduction to spatial transcriptomics for biomedical research. Genome Med. 14, 68 (2022).
Longo, S. K., Guo, M. G., Ji, A. L. & Khavari, P. A. Integrating single-cell and spatial transcriptomics to elucidate intercellular tissue dynamics. Nat. Rev. Genet. 22, 627–644 (2021).
Rao, A., Barkley, D., Franca, G. S. & Yanai, I. Exploring tissue architecture using spatial transcriptomics. Nature 596, 211–220 (2021).
Dries, R. et al. Advances in spatial transcriptomic data analysis. Genome Res. 31, 1706–1718 (2021).
Tan, X., Su, A., Tran, M. & Nguyen, Q. SpaCell: integrating tissue morphology and spatial gene expression to predict disease cells. Bioinformatics 36, 2293–2294 (2020).
Hu, J. et al. SpaGCN: integrating gene expression, spatial location and histology to identify spatial domains and spatially variable genes by graph convolutional network. Nat. Methods 18, 1342–1351 (2021).
Edsgard, D., Johnsson, P. & Sandberg, R. Identification of spatial expression trends in single-cell gene expression data. Nat. Methods 15, 339–342 (2018).
Svensson, V., Teichmann, S. A. & Stegle, O. SpatialDE: identification of spatially variable genes. Nat. Methods 15, 343–346 (2018).
Sun, S., Zhu, J. & Zhou, X. Statistical analysis of spatial expression patterns for spatially resolved transcriptomic studies. Nat. Methods 17, 193–200 (2020).
Andersson, A. et al. Single-cell and spatial transcriptomics enables probabilistic inference of cell type topography. Commun. Biol. 3, 565 (2020).
Elosua-Bayes, M., Nieto, P., Mereu, E., Gut, I. & Heyn, H. SPOTlight: seeded NMF regression to deconvolute spatial transcriptomics spots with single-cell transcriptomes. Nucleic Acids Res. 49, e50 (2021).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. 40, 517–526 (2021).
Bergenstrahle, L. et al. Super-resolved spatial transcriptomics by deep data fusion. Nat. Biotechnol. 40, 476–479 (2022).
Dries, R. et al. Giotto: a toolbox for integrative analysis and visualization of spatial expression data. Genome Biol. 22, 78 (2021).
Cang, Z. & Nie, Q. Inferring spatial and signaling relationships between cells from single cell transcriptomic data. Nat. Commun. 11, 2084 (2020).
Berglund, E. et al. Spatial maps of prostate cancer transcriptomes reveal an unexplored landscape of heterogeneity. Nat. Commun. 9, 2419 (2018).
Wang, Y., Ma, S. & Ruzzo, W. L. Spatial modeling of prostate cancer metabolic gene expression reveals extensive heterogeneity and selective vulnerabilities. Sci. Rep. 10, 3490 (2020).
Denisenko, E. et al. Systematic assessment of tissue dissociation and storage biases in single-cell and single-nucleus RNA-seq workflows. Genome Biol. 21, 130 (2020).
Wang, X., Park, J., Susztak, K., Zhang, N. R. & Li, M. Bulk tissue cell type deconvolution with multi-subject single-cell expression reference. Nat. Commun. 10, 380 (2019).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782 (2019).
Lu, K. et al. Identification of novel biomarkers in Hunner’s interstitial cystitis using the CIBERSORT, an algorithm based on machine learning. BMC Urol. 21, 109 (2021).
Gamper, M. et al. Gene expression profile of bladder tissue of patients with ulcerative interstitial cystitis. BMC Genom. 10, 199 (2009).
Colaco, M. et al. Correlation of gene expression with bladder capacity in interstitial cystitis/bladder pain syndrome. J. Urol. 192, 1123–1129 (2014).
Lindskrog, S. V. et al. An integrated multi-omics analysis identifies prognostic molecular subtypes of non-muscle-invasive bladder cancer. Nat. Commun. 12, 2301 (2021).
Yu, L. et al. Prognostic significance of lineage diversity in bladder cancer revealed by single-cell sequencing. Front. Genet. 13, 862634 (2022).
Lopez, A. & Liao, J. C. Emerging endoscopic imaging technologies for bladder cancer detection. Curr. Urol. Rep. 15, 406 (2014).
Shkolyar, E. et al. Augmented bladder tumor detection using deep learning. Eur. Urol. 76, 714–718 (2019).
Ali, N. et al. Deep learning-based classification of blue light cystoscopy imaging during transurethral resection of bladder tumors. Sci. Rep. 11, 11629 (2021).
Wu, S. et al. An artificial intelligence system for the detection of bladder cancer via cystoscopy: a multicenter diagnostic study. J. Natl Cancer Inst. 114, 220–227 (2022).
Negassi, M., Suarez-Ibarrola, R., Hein, S., Miernik, A. & Reiterer, A. Application of artificial neural networks for automated analysis of cystoscopic images: a review of the current status and future prospects. World J. Urol. 38, 2349–2358 (2020).
Chan, E. O., Pradere, B. & Teoh, J. Y., European Association of Urology - Young Academic Urologists Urothelial Carcinoma Working Group The use of artificial intelligence for the diagnosis of bladder cancer: a review and perspectives. Curr. Opin. Urol. 31, 397–403 (2021).
Lenis, A. T. & Litwin, M. S. Does artificial intelligence meaningfully enhance cystoscopy. J. Natl Cancer Inst. 114, 174–175 (2022).
Sanghvi, A. B., Allen, E. Z., Callenberg, K. M. & Pantanowitz, L. Performance of an artificial intelligence algorithm for reporting urine cytopathology. Cancer Cytopathol. 127, 658–666 (2019).
Nojima, S. et al. A deep learning system to diagnose the malignant potential of urothelial carcinoma cells in cytology specimens. Cancer Cytopathol. 129, 984–995 (2021).
Lebret, T. et al. Artificial intelligence to improve cytology performances in bladder carcinoma detection: results of the VisioCyt test. BJU Int. 129, 356–363 (2022).
Sanchez-Carbayo, M., Socci, N. D., Lozano, J., Saint, F. & Cordon-Cardo, C. Defining molecular profiles of poor outcome in patients with invasive bladder cancer using oligonucleotide microarrays. J. Clin. Oncol. 24, 778–789 (2006).
Kim, W. J. et al. Predictive value of progression-related gene classifier in primary non-muscle invasive bladder cancer. Mol. Cancer 9, 3 (2010).
Lindgren, D. et al. Combined gene expression and genomic profiling define two intrinsic molecular subtypes of urothelial carcinoma and gene signatures for molecular grading and outcome. Cancer Res. 70, 3463–3472 (2010).
Sjodahl, G. et al. A molecular taxonomy for urothelial carcinoma. Clin. Cancer Res. 18, 3377–3386 (2012).
Damrauer, J. S. et al. Intrinsic subtypes of high-grade bladder cancer reflect the hallmarks of breast cancer biology. Proc. Natl Acad. Sci. USA 111, 3110–3115 (2014).
Choi, W. et al. Identification of distinct basal and luminal subtypes of muscle-invasive bladder cancer with different sensitivities to frontline chemotherapy. Cancer Cell 25, 152–165 (2014).
Kardos, J. et al. Development and validation of a NanoString BASE47 bladder cancer gene classifier. PLoS ONE 15, e0243935 (2020).
Robertson, A. G. et al. Comprehensive molecular characterization of muscle-invasive bladder cancer. Cell 171, 540–556.e25 (2017).
Sjodahl, G., Eriksson, P., Liedberg, F. & Hoglund, M. Molecular classification of urothelial carcinoma: global mRNA classification versus tumour-cell phenotype classification. J. Pathol. 242, 113–125 (2017).
Batista da Costa, J. et al. Molecular characterization of neuroendocrine-like bladder cancer. Clin. Cancer Res. 25, 3908–3920 (2019).
de Jong, J. J. et al. Long non-coding RNAs identify a subset of luminal muscle-invasive bladder cancer patients with favorable prognosis. Genome Med. 11, 60 (2019).
Grivas, P. et al. Validation of a neuroendocrine-like classifier confirms poor outcomes in patients with bladder cancer treated with cisplatin-based neoadjuvant chemotherapy. Urol. Oncol. 38, 262–268 (2020).
Hedegaard, J. et al. Comprehensive transcriptional analysis of early-stage urothelial carcinoma. Cancer Cell 30, 27–42 (2016).
Hurst, C. D. et al. Genomic subtypes of non-invasive bladder cancer with distinct metabolic profile and female gender bias in KDM6A mutation frequency. Cancer Cell 32, 701–715.e7 (2017).
Kates, M. et al. Adaptive immune resistance to intravesical BCG in non-muscle invasive bladder cancer: implications for prospective BCG-unresponsive trials. Clin. Cancer Res. 26, 882–891 (2020).
Strandgaard, T. et al. Elevated T-cell exhaustion and urinary tumor DNA levels are associated with bacillus Calmette-Guérin failure in patients with non-muscle-invasive bladder cancer. Eur. Urol. 82, 646–656 (2022).
Kamoun, A. et al. A consensus molecular classification of muscle-invasive bladder cancer. Eur. Urol. 77, 420–433 (2020).
de Jong, J. J. et al. Gene expression profiling of muscle-invasive bladder cancer with secondary variant histology. Am. J. Clin. Pathol. 156, 895–905 (2021).
Lotan, Y. et al. Patients with muscle-invasive bladder cancer with nonluminal subtype derive greatest benefit from platinum based neoadjuvant chemotherapy. J. Urol. 207, 541–550 (2022).
Morera, D. S. et al. Clinical parameters outperform molecular subtypes for predicting outcome in bladder cancer: results from multiple cohorts, including TCGA. J. Urol. 203, 62–72 (2020).
Woerl, A. C. et al. Deep learning predicts molecular subtype of muscle-invasive bladder cancer from conventional histopathological slides. Eur. Urol. 78, 256–264 (2020).
Roubal, K., Myint, Z. W. & Kolesar, J. M. Erdafitinib: a novel therapy for FGFR-mutated urothelial cancer. Am. J. Health Syst. Pharm. 77, 346–351 (2020).
Loeffler, C. M. L. et al. Artificial intelligence-based detection of FGFR3 mutational status directly from routine histology in bladder cancer: a possible preselection for molecular testing? Eur. Urol. Focus. 8, 472–479 (2021).
McConkey, D. J. et al. Therapeutic opportunities in the intrinsic subtypes of muscle-invasive bladder cancer. Hematol. Oncol. Clin. North. Am. 29, 377–394 (2015).
Motterle, G., Andrews, J. R., Morlacco, A. & Karnes, R. J. Predicting response to neoadjuvant chemotherapy in bladder cancer. Eur. Urol. Focus. 6, 642–649 (2020).
Takata, R. et al. Predicting response to methotrexate, vinblastine, doxorubicin, and cisplatin neoadjuvant chemotherapy for bladder cancers through genome-wide gene expression profiling. Clin. Cancer Res. 11, 2625–2636 (2005).
Kato, Y. et al. Predicting response of bladder cancers to gemcitabine and carboplatin neoadjuvant chemotherapy through genome-wide gene expression profiling. Exp. Ther. Med. 2, 47–56 (2011).
Als, A. B. et al. Emmprin and survivin predict response and survival following cisplatin-containing chemotherapy in patients with advanced bladder cancer. Clin. Cancer Res. 13, 4407–4414 (2007).
Kato, Y. et al. A prospective study to examine the accuracies and efficacies of prediction systems for response to neoadjuvant chemotherapy for muscle invasive bladder cancer. Oncol. Lett. 16, 5775–5784 (2018).
McConkey, D. J. et al. A prognostic gene expression signature in the molecular classification of chemotherapy-naive urothelial cancer is predictive of clinical outcomes from neoadjuvant chemotherapy: a phase 2 trial of dose-dense methotrexate, vinblastine, doxorubicin, and cisplatin with bevacizumab in urothelial cancer. Eur. Urol. 69, 855–862 (2016).
Seiler, R. et al. Impact of molecular subtypes in muscle-invasive bladder cancer on predicting response and survival after neoadjuvant chemotherapy. Eur. Urol. 72, 544–554 (2017).
Moschini, M. et al. Characteristics and clinical significance of histological variants of bladder cancer. Nat. Rev. Urol. 14, 651–668 (2017).
Warrick, J. I. et al. Intratumoral heterogeneity of bladder cancer by molecular subtypes and histologic variants. Eur. Urol. 75, 18–22 (2019).
Sjodahl, G. et al. Molecular subtypes as a basis for stratified use of neoadjuvant chemotherapy for muscle-invasive bladder cancer – a narrative review. Cancers 14, 1692 (2022).
Bellmunt, J., Powles, T. & Vogelzang, N. J. A review on the evolution of PD-1/PD-L1 immunotherapy for bladder cancer: the future is now. Cancer Treat. Rev. 54, 58–67 (2017).
Rosenberg, J. E. et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387, 1909–1920 (2016).
Kim, J. et al. The Cancer Genome Atlas expression subtypes stratify response to checkpoint inhibition in advanced urothelial cancer and identify a subset of patients with high survival probability. Eur. Urol. 75, 961–964 (2019).
Korpal, M. et al. Evasion of immunosurveillance by genomic alterations of PPARγ/RXRα in bladder cancer. Nat. Commun. 8, 103 (2017).
Necchi, A. et al. Impact of molecular subtyping and immune infiltration on pathological response and outcome following neoadjuvant pembrolizumab in muscle-invasive bladder cancer. Eur. Urol. 77, 701–710 (2020).
Havaleshko, D. M. et al. Prediction of drug combination chemosensitivity in human bladder cancer. Mol. Cancer Ther. 6, 578–586 (2007).
Lee, J. K. et al. A strategy for predicting the chemosensitivity of human cancers and its application to drug discovery. Proc. Natl Acad. Sci. USA 104, 13086–13091 (2007).
Smith, S. C., Baras, A. S., Lee, J. K. & Theodorescu, D. The COXEN principle: translating signatures of in vitro chemosensitivity into tools for clinical outcome prediction and drug discovery in cancer. Cancer Res. 70, 1753–1758 (2010).
Flaig, T. W. et al. A randomized phase II study of coexpression extrapolation (COXEN) with neoadjuvant chemotherapy for bladder cancer (SWOG S1314; NCT02177695). Clin. Cancer Res. 27, 2435–2441 (2021).
Kong, J. et al. Network-based machine learning in colorectal and bladder organoid models predicts anti-cancer drug efficacy in patients. Nat. Commun. 11, 5485 (2020).
Barabasi, A. L., Gulbahce, N. & Loscalzo, J. Network medicine: a network-based approach to human disease. Nat. Rev. Genet. 12, 56–68 (2011).
Menche, J. et al. Disease networks. Uncovering disease–disease relationships through the incomplete interactome. Science 347, 1257601 (2015).
Szklarczyk, D. et al. STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
Cha, K. H. et al. Bladder cancer treatment response assessment in CT using radiomics with deep-learning. Sci. Rep. 7, 8738 (2017).
Tomlins, S. A. et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 310, 644–648 (2005).
Pettersson, A. et al. The TMPRSS2:ERG rearrangement, ERG expression, and prostate cancer outcomes: a cohort study and meta-analysis. Cancer Epidemiol. Biomark. Prev. 21, 1497–1509 (2012).
Adamo, P. & Ladomery, M. R. The oncogene ERG: a key factor in prostate cancer. Oncogene 35, 403–414 (2016).
Rosen, P. et al. Clinical potential of the ERG oncoprotein in prostate cancer. Nat. Rev. Urol. 9, 131–137 (2012).
Taylor, B. S. et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 18, 11–22 (2010).
You, S. et al. Integrated classification of prostate cancer reveals a novel luminal subtype with poor outcome. Cancer Res. 76, 4948–4958 (2016).
Zhao, S. G. et al. Associations of luminal and basal subtyping of prostate cancer with prognosis and response to androgen deprivation therapy. JAMA Oncol. 3, 1663–1672 (2017).
Signoretti, S. et al. p63 is a prostate basal cell marker and is required for prostate development. Am. J. Pathol. 157, 1769–1775 (2000).
Stoyanova, T. et al. Prostate cancer originating in basal cells progresses to adenocarcinoma propagated by luminal-like cells. Proc. Natl Acad. Sci. USA 110, 20111–20116 (2013).
Yoon, J. et al. A comparative study of PCS and PAM50 prostate cancer classification schemes. Prostate Cancer Prostatic Dis. 24, 733–742 (2021).
Mosley, J. D. & Keri, R. A. Cell cycle correlated genes dictate the prognostic power of breast cancer gene lists. BMC Med. Genom. 1, 11 (2008).
Cuzick, J. et al. Prognostic value of an RNA expression signature derived from cell cycle proliferation genes in patients with prostate cancer: a retrospective study. Lancet Oncol. 12, 245–255 (2011).
Whitfield, M. L. et al. Identification of genes periodically expressed in the human cell cycle and their expression in tumors. Mol. Biol. Cell 13, 1977–2000 (2002).
Cuzick, J. et al. Prognostic value of a cell cycle progression signature for prostate cancer death in a conservatively managed needle biopsy cohort. Br. J. Cancer 106, 1095–1099 (2012).
Erho, N. et al. Discovery and validation of a prostate cancer genomic classifier that predicts early metastasis following radical prostatectomy. PLoS ONE 8, e66855 (2013).
Karnes, R. J. et al. Validation of a genomic classifier that predicts metastasis following radical prostatectomy in an at risk patient population. J. Urol. 190, 2047–2053 (2013).
Klein, E. A. et al. Decipher genomic classifier measured on prostate biopsy predicts metastasis risk. Urology 90, 148–152 (2016).
Klein, E. A. et al. A 17-gene assay to predict prostate cancer aggressiveness in the context of Gleason grade heterogeneity, tumor multifocality, and biopsy undersampling. Eur. Urol. 66, 550–560 (2014).
Van Den Eeden, S. K. et al. A biopsy-based 17-gene genomic prostate score as a predictor of metastases and prostate cancer death in surgically treated men with clinically localized disease. Eur. Urol. 73, 129–138 (2018).
Brooks, M. A. et al. GPS assay association with long-term cancer outcomes: twenty-year risk of distant metastasis and prostate cancer-specific mortality. JCO Precis. Oncol. 5, 325 (2021).
Hu, J. C. et al. Clinical utility of gene expression classifiers in men with newly diagnosed prostate cancer. JCO Precis. Oncol. 2, 163 (2018).
Fine, N. D., LaPolla, F., Epstein, M., Loeb, S. & Dani, H. Genomic classifiers for treatment selection in newly diagnosed prostate cancer. BJU Int. 124, 578–586 (2019).
Karnes, R. J. et al. Development and validation of a prostate cancer genomic signature that predicts early ADT treatment response following radical prostatectomy. Clin. Cancer Res. 24, 3908–3916 (2018).
Feng, F. Y. et al. Association of molecular subtypes with differential outcome to apalutamide treatment in nonmetastatic castration-resistant prostate cancer. JAMA Oncol. 7, 1005–1014 (2021).
Smith, M. R. et al. Apalutamide treatment and metastasis-free survival in prostate cancer. N. Engl. J. Med. 378, 1408–1418 (2018).
Elmarakeby, H. A. et al. Biologically informed deep neural network for prostate cancer discovery. Nature 598, 348–352 (2021).
Fabregat, A. et al. The reactome pathway knowledgebase. Nucleic Acids Res. 46, D649–D655 (2018).
Snow, P. B., Smith, D. S. & Catalona, W. J. Artificial neural networks in the diagnosis and prognosis of prostate cancer: a pilot study. J. Urol. 152, 1923–1926 (1994).
Mosquera-Lopez, C., Agaian, S., Velez-Hoyos, A. & Thompson, I. Computer-aided prostate cancer diagnosis from digitized histopathology: a review on texture-based systems. IEEE Rev. Biomed. Eng. 8, 98–113 (2015).
Turkbey, B. & Haider, M. A. Deep learning-based artificial intelligence applications in prostate MRI: brief summary. Br. J. Radiol. 95, 20210563 (2022).
Suarez-Ibarrola, R. et al. Artificial intelligence in magnetic resonance imaging-based prostate cancer diagnosis: where do we stand in 2021? Eur. Urol. Focus. 8, 409–417 (2022).
Ferro, M. et al. Radiomics in prostate cancer: an up-to-date review. Ther. Adv. Urol. 14, 17562872221109020 (2022).
Baydoun, A. et al. Artificial intelligence applications in prostate cancer. Prostate Cancer Prostatic Dis. https://doi.org/10.1038/s41391-023-00684-0 (2023).
Doyle, S., Feldman, M., Tomaszewski, J. & Madabhushi, A. A boosted Bayesian multiresolution classifier for prostate cancer detection from digitized needle biopsies. IEEE Trans. Biomed. Eng. 59, 1205–1218 (2012).
Berney, D. M. et al. The reasons behind variation in Gleason grading of prostatic biopsies: areas of agreement and misconception among 266 European pathologists. Histopathology 64, 405–411 (2014).
Nir, G. et al. Automatic grading of prostate cancer in digitized histopathology images: learning from multiple experts. Med. Image Anal. 50, 167–180 (2018).
Strom, P. et al. Artificial intelligence for diagnosis and grading of prostate cancer in biopsies: a population-based, diagnostic study. Lancet Oncol. 21, 222–232 (2020).
Pantanowitz, L. et al. An artificial intelligence algorithm for prostate cancer diagnosis in whole slide images of core needle biopsies: a blinded clinical validation and deployment study. Lancet Digit. Health 2, e407–e416 (2020).
Nagpal, K. et al. Development and validation of a deep learning algorithm for Gleason grading of prostate cancer from biopsy specimens. JAMA Oncol. 6, 1372–1380 (2020).
Campanella, G. et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 25, 1301–1309 (2019).
Huang, W. et al. Development and validation of an artificial intelligence-powered platform for prostate cancer grading and quantification. JAMA Netw. Open. 4, e2132554 (2021).
da Silva, L. M. et al. Independent real-world application of a clinical-grade automated prostate cancer detection system. J. Pathol. 254, 147–158 (2021).
Pound, C. R. et al. Natural history of progression after PSA elevation following radical prostatectomy. JAMA 281, 1591–1597 (1999).
Wong, N. C., Lam, C., Patterson, L. & Shayegan, B. Use of machine learning to predict early biochemical recurrence after robot-assisted prostatectomy. BJU Int. 123, 51–57 (2019).
Eksi, M. et al. Machine learning algorithms can more efficiently predict biochemical recurrence after robot-assisted radical prostatectomy. Prostate 81, 913–920 (2021).
Tan, Y. G. et al. Incorporating artificial intelligence in urology: supervised machine learning algorithms demonstrate comparative advantage over nomograms in predicting biochemical recurrence after prostatectomy. Prostate 82, 298–305 (2022).
Cheng, L. et al. Risk of prostate carcinoma death in patients with lymph node metastasis. Cancer 91, 66–73 (2001).
Gandaglia, G. et al. A novel nomogram to identify candidates for extended pelvic lymph node dissection among patients with clinically localized prostate cancer diagnosed with magnetic resonance imaging-targeted and systematic biopsies. Eur. Urol. 75, 506–514 (2019).
Luzzago, S. et al. A novel nomogram to identify candidates for active surveillance amongst patients with International Society of Urological Pathology (ISUP) grade group (GG) 1 or ISUP GG2 prostate cancer, according to multiparametric magnetic resonance imaging findings. BJU Int. 126, 104–113 (2020).
Wessels, F. et al. Deep learning approach to predict lymph node metastasis directly from primary tumour histology in prostate cancer. BJU Int. 128, 352–360 (2021).
Davis, C. F. et al. The somatic genomic landscape of chromophobe renal cell carcinoma. Cancer Cell 26, 319–330 (2014).
Ricketts, C. J. et al. The Cancer Genome Atlas comprehensive molecular characterization of renal cell carcinoma. Cell Rep. 23, 313–326.e5 (2018).
Linehan, W. M. & Ricketts, C. J. The Cancer Genome Atlas of renal cell carcinoma: findings and clinical implications. Nat. Rev. Urol. 16, 539–552 (2019).
Beuselinck, B. et al. Molecular subtypes of clear cell renal cell carcinoma are associated with sunitinib response in the metastatic setting. Clin. Cancer Res. 21, 1329–1339 (2015).
Rini, B. et al. A 16-gene assay to predict recurrence after surgery in localised renal cell carcinoma: development and validation studies. Lancet Oncol. 16, 676–685 (2015).
Motzer, R. J. et al. Molecular subsets in renal cancer determine outcome to checkpoint and angiogenesis blockade. Cancer Cell 38, 803–817.e4 (2020).
Buttner, F. A. et al. A novel molecular signature identifies mixed subtypes in renal cell carcinoma with poor prognosis and independent response to immunotherapy. Genome Med. 14, 105 (2022).
Motzer, R. J. et al. Molecular characterization of renal cell carcinoma tumors from a phase III anti-angiogenic adjuvant therapy trial. Nat. Commun. 13, 5959 (2022).
Rini, B. I. et al. Validation of the 16-gene recurrence score in patients with locoregional, high-risk renal cell carcinoma from a phase III trial of adjuvant sunitinib. Clin. Cancer Res. 24, 4407–4415 (2018).
Rini, B. I. et al. Atezolizumab plus bevacizumab versus sunitinib in patients with previously untreated metastatic renal cell carcinoma (IMmotion151): a multicentre, open-label, phase 3, randomised controlled trial. Lancet 393, 2404–2415 (2019).
McDaniel, A. S. et al. Genomic profiling of penile squamous cell carcinoma reveals new opportunities for targeted therapy. Cancer Res. 75, 5219–5227 (2015).
Necchi, A. et al. Gene expression profiling of advanced penile squamous cell carcinoma receiving cisplatin-based chemotherapy improves prognostication and identifies potential therapeutic targets. Eur. Urol. Focus. 4, 733–736 (2018).
Macedo, J. et al. Genomic profiling reveals the pivotal role of hrHPV driving copy number and gene expression alterations, including mRNA downregulation of TP53 and RB1 in penile cancer. Mol. Carcinog. 59, 604–617 (2020).
Chahoud, J. et al. Whole-exome sequencing in penile squamous cell carcinoma uncovers novel prognostic categorization and drug targets similar to head and neck squamous cell carcinoma. Clin. Cancer Res. 27, 2560–2570 (2021).
Jacob, J. et al. Comprehensive genomic profiling of histologic subtypes of urethral carcinomas. Urol. Oncol. 39, 731.e1–731.e15 (2021).
Hovelson, D. H. et al. Development and validation of a scalable next-generation sequencing system for assessing relevant somatic variants in solid tumors. Neoplasia 17, 385–399 (2015).
Sambandam, V. et al. PDK1 mediates NOTCH1-mutated head and neck squamous carcinoma vulnerability to therapeutic PI3K/mTOR inhibition. Clin. Cancer Res. 25, 3329–3340 (2019).
Hashemi Gheinani, A., Bigger-Allen, A., Wacker, A. & Adam, R. M. Systems analysis of benign bladder disorders: insights from omics analysis. Am. J. Physiol. Ren. Physiol. 318, F901–F910 (2020).
Gheinani, A. H. et al. Integrated mRNA-miRNA transcriptome analysis of bladder biopsies from patients with bladder pain syndrome identifies signaling alterations contributing to the disease pathogenesis. BMC Urol. 21, 172 (2021).
Cheng, X. F. et al. Integrated analysis of microarray studies to identify novel diagnostic markers in bladder pain syndrome/interstitial cystitis with Hunner lesion. Int. J. Gen. Med. 15, 3143–3154 (2022).
Joseph, D. B. et al. Single-cell analysis of mouse and human prostate reveals novel fibroblasts with specialized distribution and microenvironment interactions. J. Pathol. 255, 141–154 (2021).
Middleton, L. W. et al. Genomic analysis of benign prostatic hyperplasia implicates cellular re-landscaping in disease pathogenesis. JCI Insight 5, e129749 (2019).
Liu, D. et al. Integrative multiplatform molecular profiling of benign prostatic hyperplasia identifies distinct subtypes. Nat. Commun. 11, 1987 (2020).
Yang, B., Veneziano, D. & Somani, B. K. Artificial intelligence in the diagnosis, treatment and prevention of urinary stones. Curr. Opin. Urol. 30, 782–787 (2020).
Michaels, E. K. et al. Use of a neural network to predict stone growth after shock wave lithotripsy. Urology 51, 335–338 (1998).
Black, K. M., Law, H., Aldoukhi, A., Deng, J. & Ghani, K. R. Deep learning computer vision algorithm for detecting kidney stone composition. BJU Int. 125, 920–924 (2020).
Aminsharifi, A. et al. Artificial neural network system to predict the postoperative outcome of percutaneous nephrolithotomy. J. Endourol. 31, 461–467 (2017).
Ganesan, V. & Pearle, M. S. Artificial intelligence in stone disease. Curr. Opin. Urol. 31, 391–396 (2021).
Muller, S. et al. Can a dinosaur think? Implementation of artificial intelligence in extracorporeal shock wave lithotripsy. Eur. Urol. Open. Sci. 27, 33–42 (2021).
Aminsharifi, A. et al. Predicting the postoperative outcome of percutaneous nephrolithotomy with machine learning system: software validation and comparative analysis with Guy’s stone score and the CROES nomogram. J. Endourol. 34, 692–699 (2020).
Venhola, M., Reunanen, M., Taskinen, S., Lahdes-Vasama, T. & Uhari, M. Interobserver and intra-observer agreement in interpreting urodynamic measurements in children. J. Urol. 169, 2344–2346 (2003).
Dudley, A. G. et al. Interrater reliability in pediatric urodynamic tracings: a pilot study. J. Urol. 197, 865–870 (2017).
Wang, H. S. et al. Pattern recognition algorithm to identify detrusor overactivity on urodynamics. Neurourol. Urodyn. 40, 428–434 (2021).
Hobbs, K. T. et al. Machine learning for urodynamic detection of detrusor overactivity. Urology 159, 247–254 (2022).
Doern, C. D. & Richardson, S. E. Diagnosis of urinary tract infections in children. J. Clin. Microbiol. 54, 2233–2242 (2016).
Medina, M. & Castillo-Pino, E. An introduction to the epidemiology and burden of urinary tract infections. Ther. Adv. Urol. 11, 1756287219832172 (2019).
Nitzan, O., Elias, M., Chazan, B. & Saliba, W. Urinary tract infections in patients with type 2 diabetes mellitus: review of prevalence, diagnosis, and management. Diabetes Metab. Syndr. Obes. 8, 129–136 (2015).
Pannek, J. & Wollner, J. Management of urinary tract infections in patients with neurogenic bladder: challenges and solutions. Res. Rep. Urol. 9, 121–127 (2017).
Ripa, F. et al. Association of kidney stones and recurrent UTIs: the chicken and egg situation. A systematic review of literature. Curr. Urol. Rep. 23, 165–174 (2022).
Taylor, R. A., Moore, C. L., Cheung, K. H. & Brandt, C. Predicting urinary tract infections in the emergency department with machine learning. PLoS ONE 13, e0194085 (2018).
Ozkan, I. A., Koklu, M. & Sert, I. U. Diagnosis of urinary tract infection based on artificial intelligence methods. Comput. Meth Prog. Bio 166, 51–59 (2018).
Price, T. K. et al. The clinical urine culture: enhanced techniques improve detection of clinically relevant microorganisms. J. Clin. Microbiol. 54, 1216–1222 (2016).
Szlachta-McGinn, A. et al. Molecular diagnostic methods versus conventional urine culture for diagnosis and treatment of urinary tract infection: a systematic review and meta-analysis. Eur. Urol. Open. Sci. 44, 113–124 (2022).
Roux-Dalvai, F. et al. Fast and accurate bacterial species identification in urine specimens using LC-MS/MS mass spectrometry and machine learning. Mol. Cell Proteom. 18, 2492–2505 (2019).
Advanced Analytics Group of Pediatric Urology and ORC Personalized Medicine Group Targeted workup after initial febrile urinary tract infection: using a novel machine learning model to identify children most likely to benefit from voiding cystourethrogram. J. Urol. 202, 144–152 (2019).
Bagli, D. J. et al. Artificial neural networks in pediatric urology: prediction of sonographic outcome following pyeloplasty. J. Urol. 160, 980–983 (1998).
Seckiner, I., Seckiner, S. U., Bayrak, O. & Erturhan, S. Use of artificial neural networks in the management of antenatally diagnosed ureteropelvic junction obstruction. Can. Urol. Assoc. J. 5, E152–E155 (2011).
Drysdale, E. et al. Personalized application of machine learning algorithms to identify pediatric patients at risk for recurrent ureteropelvic junction obstruction after dismembered pyeloplasty. World J. Urol. 40, 593–599 (2022).
Rademakers, K. et al. Male bladder outlet obstruction: time to re-evaluate the definition and reconsider our diagnostic pathway? ICI-RS 2015. Neurourol. Urodyn. 36, 894–901 (2017).
Sonke, G. S., Heskes, T., Verbeek, A. L., de la Rosette, J. J. & Kiemeney, L. A. Prediction of bladder outlet obstruction in men with lower urinary tract symptoms using artificial neural networks. J. Urol. 163, 300–305 (2000).
Abdovic, S. et al. Predicting posterior urethral obstruction in boys with lower urinary tract symptoms using deep artificial neural network. World J. Urol. 37, 1973–1979 (2019).
Yin, S. et al. Multi-instance deep learning of ultrasound imaging data for pattern classification of congenital abnormalities of the kidney and urinary tract in children. Urology 142, 183–189 (2020).
Kwong, J. C. et al. Posterior urethral valves outcomes prediction (PUVOP): a machine learning tool to predict clinically relevant outcomes in boys with posterior urethral valves. Pediatr. Nephrol. 37, 1067–1084 (2021).
Thomas, A. A. et al. Extracting data from electronic medical records: validation of a natural language processing program to assess prostate biopsy results. World J. Urol. 32, 99–103 (2014).
Odisho, A. Y. et al. Automating the capture of structured pathology data for prostate cancer clinical care and research. JCO Clin. Cancer Inf. 3, 1–8 (2019).
Schroeck, F. R. et al. Development of a natural language processing engine to generate bladder cancer pathology data for health services research. Urology 110, 84–91 (2017).
Glaser, A. P. et al. Automated extraction of grade, stage, and quality information from transurethral resection of bladder tumor pathology reports using natural language processing. JCO Clin. Cancer Inf. 2, 1–8 (2018).
Bashashati, A. & Goldenberg, S. L. AI for prostate cancer diagnosis – hype or today’s reality? Nat. Rev. Urol. 19, 261–262 (2022).
Yang, C. et al. Trends in the conduct and reporting of clinical prediction model development and validation: a systematic review. J. Am. Med. Inf. Assoc. 29, 983–989 (2022).
Reinke, A., Tizabi, M. D., Eisenmann, M. & Maier-Hein, L. Common pitfalls and recommendations for grand challenges in medical artificial intelligence. Eur. Urol. Focus. 7, 710–712 (2021).
Bulten, W. et al. Artificial intelligence for diagnosis and Gleason grading of prostate cancer: the PANDA challenge. Nat. Med. 28, 154–163 (2022).
Maier-Hein, L. et al. Why rankings of biomedical image analysis competitions should be interpreted with care. Nat. Commun. 9, 5217 (2018).
Zhou, Q., Chen, Z. H., Cao, Y. H. & Peng, S. Clinical impact and quality of randomized controlled trials involving interventions evaluating artificial intelligence prediction tools: a systematic review. NPJ Digit. Med. 4, 154 (2021).
Dhiman, P. et al. Methodological conduct of prognostic prediction models developed using machine learning in oncology: a systematic review. BMC Med. Res. Methodol. 22, 101 (2022).
Collins, G. S. et al. Protocol for development of a reporting guideline (TRIPOD-AI) and risk of bias tool (PROBAST-AI) for diagnostic and prognostic prediction model studies based on artificial intelligence. BMJ Open 11, e048008 (2021).
Calogero, A. E., Burgio, G., Condorelli, R. A., Cannarella, R. & La Vignera, S. Epidemiology and risk factors of lower urinary tract symptoms/benign prostatic hyperplasia and erectile dysfunction. Aging Male 22, 12–19 (2019).
Grossman, R. L. et al. Toward a shared vision for cancer genomic data. N. Engl. J. Med. 375, 1109–1112 (2016).
Regev, A. et al. The Human Cell Atlas. Elife 6, e27041 (2017).
Goldman, M. J. et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat. Biotechnol. 38, 675–678 (2020).
Papatheodorou, I. et al. Expression Atlas: gene and protein expression across multiple studies and organisms. Nucleic Acids Res. 46, D246–D251 (2018).
Uhlen, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Hu, X. et al. TumorFusions: an integrative resource for cancer-associated transcript fusions. Nucleic Acids Res. 46, D1144–D1149 (2018).
Omar, M. I. et al. Introducing PIONEER: a project to harness big data in prostate cancer research. Nat. Rev. Urol. 17, 351–362 (2020).
Dunning, M. J. et al. Mining human prostate cancer datasets: the “camcAPP” shiny app. EBioMedicine 17, 5–6 (2017).
Hu, Z. et al. Genomic characterization of genes encoding histone acetylation modulator proteins identifies therapeutic targets for cancer treatment. Nat. Commun. 10, 733 (2019).
Ghoshdastider, U. et al. Pan-cancer analysis of ligand–receptor cross-talk in the tumor microenvironment. Cancer Res. 81, 1802–1812 (2021).
Rohatgi, N., Ghoshdastider, U., Baruah, P., Kulshrestha, T. & Skanderup, A. J. A pan-cancer metabolic atlas of the tumor microenvironment. Cell Rep. 39, 110800 (2022).
Stewart, B. J. et al. Spatiotemporal immune zonation of the human kidney. Science 365, 1461–1466 (2019).
McMahon, A. P. et al. GUDMAP: the genitourinary developmental molecular anatomy project. J. Am. Soc. Nephrol. 19, 667–671 (2008).
Rigden, D. J. & Fernandez, X. M. The 2018 Nucleic Acids Research database issue and the online molecular biology database collection. Nucleic Acids Res. 46, D1–D7 (2018).
van der Wijst, M. et al. Science Forum: the single-cell eQTLGen consortium. Elife 9, e52155 (2020).
Abugessaisa, I. et al. SCPortalen: human and mouse single-cell centric database. Nucleic Acids Res. 46, D781–D787 (2018).
Ner-Gaon, H., Melchior, A., Golan, N., Ben-Haim, Y. & Shay, T. JingleBells: a repository of immune-related single-cell RNA-sequencing datasets. J. Immunol. 198, 3375–3379 (2017).
Cao, Y., Zhu, J., Jia, P. & Zhao, Z. scRNASeqDB: a database for RNA-seq based gene expression profiles in human single cells. Genes 8, 368 (2017).
Cao, Z. J., Wei, L., Lu, S., Yang, D. C. & Gao, G. Searching large-scale scRNA-seq databases via unbiased cell embedding with Cell BLAST. Nat. Commun. 11, 3458 (2020).
Soneson, C. & Robinson, M. D. Bias, robustness and scalability in single-cell differential expression analysis. Nat. Methods 15, 255–261 (2018).
Svensson, V., da Veiga Beltrame, E. & Pachter, L. A curated database reveals trends in single-cell transcriptomics. Database 2020, baaa073 (2020).
Li, M. et al. DISCO: a database of Deeply Integrated human Single-Cell Omics data. Nucleic Acids Res. 50, D596–D602 (2022).
Author information
Authors and Affiliations
Contributions
All authors researched data for the article. R.M.A., A.H.G. and S.Y. contributed substantially to discussion of the content. All authors wrote the article. R.M.A., A.H.G. and S.Y. reviewed and/or edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Urology thanks Roland Arnold, Andrew Hung and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Grand Challenge for medical image analysis: www.grand-challenge.org
PAIP: https://paip2021.grand-challenge.org/
PANDA: https://panda.grand-challenge.org/
PI-CAI: https://pi-cai.grand-challenge.org/
PROSTATEx: https://prostatex.grand-challenge.org/
Supplementary information
Glossary
- Area under the curve
-
(AUC). The AUC of a receiver operating characteristic curve is used to measure the accuracy of a model. An AUC of 0.5 represents a model that does not perform any better than random chance.
- Artificial neural networks
-
Computational models inspired by the structure and functioning of biological neural networks, used in machine learning to process complex data and make predictions or classifications.
- Biases
-
Biases are systematic errors or prejudices that can exist in artificial intelligence (AI) systems, data, algorithms or decision-making processes. Biases can arise owing to various factors, such as biased data collection, biased algorithm design or biased human decisions that influence the training process. For example, bias can be introduced if a training dataset used to develop an AI model for bladder cancer classification majorly consists of patient material from a specific demographic group or sex, or if certain genomic markers are prioritized.
- Cohen’s κ coefficient
-
A statistical parameter used to measure reliability between raters that can have values from −1 to +1, in which 0 is the extent of agreement expected by random chance.
- Data leakage
-
Data leakage occurs when information from the test set leaks into the training set, or vice versa. This phenomenon can happen when the data are pre-processed or cleaned before splitting, or when the test set is used to inform feature selection or model tuning. Data leakage can result in overly optimistic performance estimates, and the model might perform poorly on new, unseen data.
- Deep learning
-
A subfield of machine learning that uses artificial neural networks with multiple layers, enabling the artificial intelligence system to learn hierarchical representations and extract intricate patterns from large datasets.
- Dropout
-
A technique used in machine learning to prevent overfitting, in which certain information is temporarily ignored during training to ensure that the model does not overly rely on specific features, improving the ability of the model to generalize and make accurate predictions. By incorporating dropout, the ability of the model to generalize is improved and placing excessive importance on individual genes is avoided, ensuring a robust and reliable analysis.
- Early stopping
-
A strategy used during machine learning training to avoid overfitting, in which the training process is stopped before completion based on a specific measure (such as validation performance) to prevent the model from becoming too specialized to the training data, ensuring good ability to generalize to new, unseen data.
- Imbalanced classes
-
Classes are imbalanced when the proportion of observations in one class is much higher or lower than in the other. This phenomenon can lead to biased performance estimates, as the model might be accurate on the dominant class but perform poorly on the minority class. This problem can be addressed by using techniques such as stratified sampling, oversampling or undersampling.
- Learning rubbish (learning garbage)
-
This refers to the process of training an artificial intelligence system using low-quality or inaccurate data. For example, if in a study, the training dataset contains gene expression profiles from unrelated cancer types or includes samples with unreliable annotations, the AI system might learn from this rubbish data and produce misleading associations.
- Microarray
-
Microarrays are nucleic acid sequences corresponding to defined genes or transcripts arrayed on a solid phase support for hybridization with cDNA prepared from samples under investigation. Microarrays enable measurement of transcript abundance on a genome-wide scale.
- Natural language processing
-
A branch of artificial intelligence focused on the interaction between computers and human language, enabling machines to understand, interpret and generate human language text or speech.
- Negative predictive value
-
The proportion of individuals with a negative test result who do not have the disease.
- Non-representative sampling
-
Sampling is non-representative when the training or test set is not representative of the population from which the test was sampled. This phenomenon can happen when the data are collected from a biased or limited source, or when hidden confounding factors influence the outcome variable. Non-representative samples can lead to poor generalization and low predictive accuracy.
- Optimal fitting
-
Optimal fitting occurs when a model is sufficiently complex to capture underlying patterns in the data and generalizes well to new data. This fitting requires a balance between model complexity and the amount and quality of data available for training the model.
- Overfitting
-
Overfitting occurs when a model is too complex and fits the training data too well, resulting in poor performance on new data. This phenomenon can happen when a model is trained on a limited set of data and learns the noise in the data instead of the underlying patterns. For example, in a situation in which a decision tree model is used to predict the outcome of a therapy based on the expression of a large number of genes in a tumour (features), but many of these genes are irrelevant or noisy, the model might overfit the data.
- Positive predictive value
-
The proportion of individuals with a positive test result who actually have the disease.
- Semi-supervised learning
-
A machine learning technique that uses a combination of labelled and unlabelled data to train an artificial intelligence system, leveraging the available labelled data and the patterns inferred from the unlabelled data.
- Small sample size
-
The sample size is small when too few observations are available in the training or test set to build or evaluate a robust model. In this case, the model either memorizes the training data or fails to capture the underlying patterns, leading to overfitting or underfitting. Small sample size can be addressed by increasing the size of the dataset, using data augmentation techniques or using models with increased robustness.
- Supervised learning
-
A machine learning technique in which the artificial intelligence system is trained using labelled data, where the input and corresponding output pairs are provided to guide the learning process.
- Test or testing set
-
A subset of a dataset used to evaluate the performance of a machine learning model. The purpose of the test dataset is to measure how well the model performs on new, unseen data. The test dataset is used to estimate the accuracy of the model’s predictions on new data.
- Training set
-
A subset of a dataset used to train a machine learning model. The purpose of the training dataset is to build the model by learning the relationships between the input variables (features) and the output variable (target variable). The model uses the training dataset to determine how to make predictions.
- Underfitting
-
Underfitting occurs when a model is too simple and fails to capture the complexity of the data, resulting in poor performance on both the training data and new data. For example, if a linear regression model is used to predict the outcome of a therapy based on the size and number of tumours but the relationship between these variables is more complex, the model might underfit the data.
- Unsupervised learning
-
A machine learning technique in which the artificial intelligence system discovers patterns or structures in data without being explicitly guided or labelled.
- Validation
-
Validation is the process of assessing the accuracy of a model on data that have not yet been seen by the model. Cross-validation is a technique in which a dataset is divided into multiple training and test sets, and the model is trained and tested on each set, to evaluate the performance of the model on the dataset. Common problems with separating the training and test sets include feature selection performed on the entire set, which can lead to overfitting and poor performance on new data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hashemi Gheinani, A., Kim, J., You, S. et al. Bioinformatics in urology — molecular characterization of pathophysiology and response to treatment. Nat Rev Urol 21, 214–242 (2024). https://doi.org/10.1038/s41585-023-00805-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41585-023-00805-3