Abstract
Voltage-gated ion channels (VGICs) comprise multiple structural units, the assembly of which is required for function1,2. Structural understanding of how VGIC subunits assemble and whether chaperone proteins are required is lacking. High-voltage-activated calcium channels (CaVs)3,4 are paradigmatic multisubunit VGICs whose function and trafficking are powerfully shaped by interactions between pore-forming CaV1 or CaV2 CaVα1 (ref. 3), and the auxiliary CaVβ5 and CaVα2δ subunits6,7. Here we present cryo-electron microscopy structures of human brain and cardiac CaV1.2 bound with CaVβ3 to a chaperone—the endoplasmic reticulum membrane protein complex (EMC)8,9—and of the assembled CaV1.2–CaVβ3–CaVα2δ-1 channel. These structures provide a view of an EMC–client complex and define EMC sites—the transmembrane (TM) and cytoplasmic (Cyto) docks; interaction between these sites and the client channel causes partial extraction of a pore subunit and splays open the CaVα2δ-interaction site. The structures identify the CaVα2δ-binding site for gabapentinoid anti-pain and anti-anxiety drugs6, show that EMC and CaVα2δ interactions with the channel are mutually exclusive, and indicate that EMC-to-CaVα2δ hand-off involves a divalent ion-dependent step and CaV1.2 element ordering. Disruption of the EMC–CaV complex compromises CaV function, suggesting that the EMC functions as a channel holdase that facilitates channel assembly. Together, the structures reveal a CaV assembly intermediate and EMC client-binding sites that could have wide-ranging implications for the biogenesis of VGICs and other membrane proteins.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Coordinates and maps of the EMC–CaV1.2(ΔC)–CaVβ3 complex (PDB: 8EOI; Electron Microscopy Data Bank (EMDB): EMD-28376, EMD-28578, EMD-28579,EMD-40559, EMD-40560)and CaV1.2(ΔC)–CaVβ3–CaVα2δ-1 complex (PDB: 8EOG; EMDB: EMD-28375, EMD-28561, EMD-28564, EMD-40561) have been deposited at the at the PDB and EMDB. MS data have been deposited at the Mass Spectrometry Interactive Virtual Environment (https://massive.ucsd.edu/) under identifier MSV000090434. Source data are provided with this paper.
References
Catterall, W. A., Wisedchaisri, G. & Zheng, N. The chemical basis for electrical signaling. Nat. Chem. Biol. 13, 455–463 (2017).
Isacoff, E. Y., Jan, L. Y. & Minor Jr. D. L. Conduits of life’s spark: a perspective on ion channel research since the birth of neuron. Neuron 80, 658–674 (2013).
Zamponi, G. W., Striessnig, J., Koschak, A. & Dolphin, A. C. The physiology, pathology, and pharmacology of voltage-gated calcium channels and their future therapeutic potential. Pharmacol. Rev. 67, 821–870 (2015).
Nanou, E. & Catterall, W. A. Calcium channels, synaptic plasticity, and neuropsychiatric disease. Neuron 98, 466–481 (2018).
Buraei, Z. & Yang, J. The β subunit of voltage-gated Ca2+ channels. Physiol. Rev. 90, 1461–1506 (2010).
Dooley, D. J., Taylor, C. P., Donevan, S. & Feltner, D. Ca2+ channel α2δ ligands: novel modulators of neurotransmission. Trends Pharmacol. Sci. 28, 75–82 (2007).
Dolphin, A. C. Voltage-gated calcium channels and their auxiliary subunits: physiology and pathophysiology and pharmacology. J. Physiol. 594, 5369–5390 (2016).
Jonikas, M. C. et al. Comprehensive characterization of genes required for protein folding in the endoplasmic reticulum. Science 323, 1693–1697 (2009).
Hegde, R. S. The function, structure, and origins of the ER membrane protein complex. Annu. Rev. Biochem. 91, 651–678 (2022).
Shistik, E., Ivanina, T., Puri, T., Hosey, M. & Dascal, N. Ca2+ current enhancement by α2/δ and β subunits in Xenopus oocytes: contribution of changes in channel gating and α1 protein level. J. Physiol. 489, 55–62 (1995).
Singer, D. et al. The roles of the subunits in the function of the calcium channel. Science 253, 1553–1557 (1991).
Gurnett, C. A., De Waard, M. & Campbell, K. P. Dual function of the voltage-dependent Ca2+ channel α2δ subunit in current stimulation and subunit interaction. Neuron 16, 431–440 (1996).
Davies, A. et al. Functional biology of the α2δ subunits of voltage-gated calcium channels. Trends Pharmacol. Sci. 28, 220–228 (2007).
Field, M. J. et al. Identification of the α2-δ-1 subunit of voltage-dependent calcium channels as a molecular target for pain mediating the analgesic actions of pregabalin. Proc. Natl Acad. Sci. USA 103, 17537–17542 (2006).
Dolphin, A. C. Voltage-gated calcium channel α2δ subunits: an assessment of proposed novel roles. F1000Res https://doi.org/10.12688/f1000research.16104.1 (2018).
Bauer, C. S. et al. The increased trafficking of the calcium channel subunit α2δ-1 to presynaptic terminals in neuropathic pain is inhibited by the α2δ ligand pregabalin. J. Neurosci. 29, 4076–4088 (2009).
Cassidy, J. S., Ferron, L., Kadurin, I., Pratt, W. S. & Dolphin, A. C. Functional exofacially tagged N-type calcium channels elucidate the interaction with auxiliary α2δ-1 subunits. Proc. Natl Acad. Sci. USA 111, 8979–8984 (2014).
Deutsch, C. The birth of a channel. Neuron 40, 265–276 (2003).
Christianson, J. C. et al. Defining human ERAD networks through an integrative mapping strategy. Nat. Cell Biol. 14, 93–105 (2011).
Guna, A., Volkmar, N., Christianson, J. C. & Hegde, R. S. The ER membrane protein complex is a transmembrane domain insertase. Science 359, 470–473 (2018).
Richard, M., Boulin, T., Robert, V. J., Richmond, J. E. & Bessereau, J. L. Biosynthesis of ionotropic acetylcholine receptors requires the evolutionarily conserved ER membrane complex. Proc. Natl Acad. Sci. USA 110, E1055–E1063 (2013).
Talbot, B. E., Vandorpe, D. H., Stotter, B. R., Alper, S. L. & Schlondorff, J. S. Transmembrane insertases and N-glycosylation critically determine synthesis, trafficking, and activity of the nonselective cation channel TRPC6. J. Biol. Chem. 294, 12655–12669 (2019).
Coelho, J. P. L. et al. A network of chaperones prevents and detects failures in membrane protein lipid bilayer integration. Nat. Commun. 10, 672 (2019).
Satoh, T., Ohba, A., Liu, Z., Inagaki, T. & Satoh, A. K. dPob/EMC is essential for biosynthesis of rhodopsin and other multi-pass membrane proteins in Drosophila photoreceptors. eLife 4, e06306 (2015).
Chitwood, P. J., Juszkiewicz, S., Guna, A., Shao, S. & Hegde, R. S. EMC is required to initiate accurate membrane protein topogenesis. Cell 175, 1507–1519 (2018).
Miller-Vedam, L. E. et al. Structural and mechanistic basis of the EMC-dependent biogenesis of distinct transmembrane clients. eLife 9, e62611 (2020).
Shurtleff, M. J. et al. The ER membrane protein complex interacts cotranslationally to enable biogenesis of multipass membrane proteins. eLife 7, e37018 (2018).
O’Donnell, J. P. et al. The architecture of EMC reveals a path for membrane protein insertion. eLife 9, e57887 (2020).
Pleiner, T. et al. Structural basis for membrane insertion by the human ER membrane protein complex. Science 369, 433–436 (2020).
Bagchi, P., Inoue, T. & Tsai, B. EMC1-dependent stabilization drives membrane penetration of a partially destabilized non-enveloped virus. eLife 5, e21470 (2016).
Hofmann, F., Flockerzi, V., Kahl, S. & Wegener, J. W. L-type CaV1.2 calcium channels: from in vitro findings to in vivo function. Physiol. Rev. 94, 303–326 (2014).
Hullin, R. et al. Cardiac L-type calcium channel β-subunits expressed in human heart have differential effects on single channel characteristics. J. Biol. Chem. 278, 21623–21630 (2003).
Zamponi, G. W. Targeting voltage-gated calcium channels in neurological and psychiatric diseases. Nat. Rev. Drug Discov. 15, 19–34 (2016).
Marcantoni, A., Calorio, C., Hidisoglu, E., Chiantia, G. & Carbone, E. Cav1.2 channelopathies causing autism: new hallmarks on Timothy syndrome. Pflugers Arch. 472, 775–789 (2020).
Chen, Z., Mondal, A. & Minor Jr. D. L. Structural basis for CaVα2δ:gabapentin binding. Nat. Struct. Mol. Biol. https://doi.org/10.1038/s41594-023-00951-7 (2023).
Kozai, D. et al. Recognition mechanism of a novel gabapentinoid drug, mirogabalin, for recombinant human α2δ1, a voltage-gated calcium channel subunit. J. Mol. Biol. 435, 168049 (2023).
Dooley, D. J., Donovan, C. M., Meder, W. P. & Whetzel, S. Z. Preferential action of gabapentin and pregabalin at P/Q-type voltage-sensitive calcium channels: inhibition of K+-evoked [3H]-norepinephrine release from rat neocortical slices. Synapse 45, 171–190 (2002).
Infield, D. T. et al. Cation–π Interactions and their functional roles in membrane proteins. J. Mol. Biol. 433, 167035 (2021).
Chen, Y. H. et al. Structural basis of the α1-β subunit interaction of voltage-gated Ca2+ channels. Nature 429, 675–680 (2004).
Van Petegem, F., Clark, K. A., Chatelain, F. C. & Minor Jr. D. L. Structure of a complex between a voltage-gated calcium channel β-subunit and an α-subunit domain. Nature 429, 671–675 (2004).
Van Petegem, F., Duderstadt, K. E., Clark, K. A., Wang, M. & Minor Jr. D. L. Alanine-scanning mutagenesis defines a conserved energetic hotspot in the CaVα1 AID-CaVβ interaction site that is critical for channel modulation. Structure 16, 280–294 (2008).
Bai, L., You, Q., Feng, X., Kovach, A. & Li, H. Structure of the ER membrane complex, a transmembrane-domain insertase. Nature 584, 475–478 (2020).
Wu, J. et al. Structure of the voltage-gated calcium channel Cav1.1 at 3.6 Å resolution. Nature 537, 191–196 (2016).
Yao, X., Gao, S. & Yan, N. Structural basis for pore blockade of human voltage-gated calcium channel Cav1.3 by motion sickness drug cinnarizine. Cell Res. https://doi.org/10.1038/s41422-022-00663-5 (2022).
Gao, S., Yao, X. & Yan, N. Structure of human Cav2.2 channel blocked by the painkiller ziconotide. Nature 596, 143–147 (2021).
Zhao, Y. et al. Cryo-EM structures of apo and antagonist-bound human Cav3.1. Nature 576, 492–497 (2019).
Brown, J. P., Dissanayake, V. U., Briggs, A. R., Milic, M. R. & Gee, N. S. Isolation of the [3H]gabapentin-binding protein/α2δ Ca2+ channel subunit from porcine brain: development of a radioligand binding assay for α2δ subunits using [3H]leucine. Anal. Biochem. 255, 236–243 (1998).
Dolphin, A. C. Calcium channel auxiliary α2δ and beta subunits: trafficking and one step beyond. Nat. Rev. Neurosci. 13, 542–555 (2012).
Wu, J. et al. Structure of the voltage-gated calcium channel Cav1.1 complex. Science 350, aad2395 (2015).
Gumerov, V. M. et al. Amino acid sensor conserved from bacteria to humans. Proc. Natl Acad. Sci. USA 119, e2110415119 (2022).
Wang, M., Offord, J., Oxender, D. L. & Su, T. Z. Structural requirement of the calcium-channel subunit α2δ for gabapentin binding. Biochem. J. 342, 313–320 (1999).
Zhao, Y. et al. Molecular basis for ligand modulation of a mammalian voltage-gated Ca2+ Channel. Cell 177, 1495–1506 (2019).
He, L. et al. Structure, gating, and pharmacology of human CaV3.3 channel. Nat. Commun. 13, 2084 (2022).
Gao, S. & Yan, N. Structural basis of the modulation of the voltage-gated calcium ion channel Cav1.1 by dihydropyridine compounds. Angew. Chem. Int. Ed. Engl. 60, 3131–3137 (2021).
Tao, X., Lee, A., Limapichat, W., Dougherty, D. A. & MacKinnon, R. A gating charge transfer center in voltage sensors. Science 328, 67–73 (2010).
Arrigoni, C. et al. Quaternary structure independent folding of voltage-gated ion channel pore domain subunits. Nat. Struct. Mol. Biol. 29, 537–548 (2022).
Bourdin, B., Briot, J., Tetreault, M. P., Sauve, R. & Parent, L. Negatively charged residues in the first extracellular loop of the L-type CaV1.2 channel anchor the interaction with the CaVα2δ1 auxiliary subunit. J. Biol. Chem. 292, 17236–17249 (2017).
Dahimene, S. et al. The α2δ-like protein Cachd1 increases N-type calcium currents and cell surface expression and competes with α2δ-1. Cell Rep. 25, 1610–1621 (2018).
Canti, C. et al. The metal-ion-dependent adhesion site in the Von Willebrand factor-A domain of α2δ subunits is key to trafficking voltage-gated Ca2+ channels. Proc. Natl Acad. Sci. USA 102, 11230–11235 (2005).
Gurnett, C. A., Felix, R. & Campbell, K. P. Extracellular interaction of the voltage-dependent Ca2+ channel α2δ and α1 subunits. J. Biol. Chem. 272, 18508–18512 (1997).
Opatowsky, Y., Chomsky-Hecht, O., Kang, M. G., Campbell, K. P. & Hirsch, J. A. The voltage-dependent calcium channel β subunit contains two stable interacting domains. J. Biol. Chem. 278, 52323–52332 (2003).
Altier, C. et al. The Cavβ subunit prevents RFP2-mediated ubiquitination and proteasomal degradation of L-type channels. Nat. Neurosci. 14, 173–180 (2011).
Waithe, D., Ferron, L., Page, K. M., Chaggar, K. & Dolphin, A. C. β-Subunits promote the expression of CaV2.2 channels by reducing their proteasomal degradation. J. Biol. Chem. 286, 9598–9611 (2011).
Fang, K. & Colecraft, H. M. Mechanism of auxiliary β-subunit-mediated membrane targeting of L-type (CaV1.2) channels. J. Physiol. 589, 4437–4455 (2011).
Altier, C. et al. Trafficking of L-type calcium channels mediated by the postsynaptic scaffolding protein AKAP79. J. Biol. Chem. 277, 33598–33603 (2002).
Obermair, G. J. et al. Reciprocal interactions regulate targeting of calcium channel beta subunits and membrane expression of α1 subunits in cultured hippocampal neurons. J. Biol. Chem. 285, 5776–5791 (2010).
Leroy, J. et al. Interaction via a key tryptophan in the I-II linker of N-type calcium channels is required for β1 but not for palmitoylated β2, implicating an additional binding site in the regulation of channel voltage-dependent properties. J. Neurosci. 25, 6984–6996 (2005).
Pleiner, T. et al. A selectivity filter in the ER membrane protein complex limits protein misinsertion at the ER. J. Cell Biol. 222, e202212007 (2023).
Opatowsky, Y., Chen, C. C., Campbell, K. P. & Hirsch, J. A. Structural analysis of the voltage-dependent calcium channel β subunit functional core and its complex with the α1 interaction domain. Neuron 42, 387–399 (2004).
Dolphin, A. C. & Lee, A. Presynaptic calcium channels: specialized control of synaptic neurotransmitter release. Nat. Rev. Neurosci. 21, 213–229 (2020).
Felix, R., Gurnett, C. A., De Waard, M. & Campbell, K. P. Dissection of functional domains of the voltage-dependent Ca2+ channel α2δ subunit. J. Neurosci. 17, 6884–6891 (1997).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996).
Pravda, L. et al. MOLEonline: a web-based tool for analyzing channels, tunnels and pores (2018 update). Nucleic Acids Res. 46, W368–W373 (2018).
Schmidt, T. G. & Skerra, A. The Strep-tag system for one-step purification and high-affinity detection or capturing of proteins. Nat. Protoc. 2, 1528–1535 (2007).
Liao, M., Cao, E., Julius, D. & Cheng, Y. Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 504, 107–112 (2013).
Elegheert, J. et al. Lentiviral transduction of mammalian cells for fast, scalable and high-level production of soluble and membrane proteins. Nat. Protoc. 13, 2991–3017 (2018).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2585 (2014).
Lee, H., Lolicato, M., Arrigoni, C. & Minor, D. L. Jr. Production of K2P2.1 (TREK-1) for structural studies. Methods Enzymol. 653, 151–188 (2021).
Shaya, D. et al. Voltage-gated sodium channel (NaV) protein dissection creates a set of functional pore-only proteins. Proc. Natl Acad. Sci. USA 108, 12313–12318 (2011).
Abderemane-Ali, F., Findeisen, F., Rossen, N. D. & Minor Jr. D. L. A selectivity filter gate controls voltage-gated calcium channel calcium-dependent inactivation. Neuron 101, 1134–1149 (2019).
Findeisen, F. & Minor Jr. D. L. Disruption of the IS6-AID linker affects voltage-gated calcium channel inactivation and facilitation. J. Gen. Physiol. 133, 327–343 (2009).
Hamill, O. P., Marty, A., Neher, E., Sakmann, B. & Sigworth, F. J. Improved patch-clamp techniques for high-resolution current recording from cells and cell-free membrane patches. Pflugers Arch. 391, 85–100 (1981).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Moriarty, N. W., Grosse-Kunstleve, R. W. & Adams, P. D. Electronic Ligand Builder and Optimization Workbench (eLBOW): a tool for ligand coordinate and restraint generation. Acta Crystallogr. D 65, 1074–1080 (2009).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Wallace, A. C., Laskowski, R. A. & Thornton, J. M. LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Protein Eng. 8, 127–134 (1995).
Laskowski, R. A. & Swindells, M. B. LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J. Chem. Inf. Model. 51, 2778–2786 (2011).
Acknowledgements
We thank T.-J. Yen, D. Bulkley, Y. Liu and H. Khant for technical help; T. Pleiner and R. Vorhees for the HEK293FT cells and EMC5-knockout and EMC6-knockout lines; and K. Brejc, H. M. Colecraft, D. Julius and G. Thiel for comments on the manuscript. This work was supported by grants NIH R01 HL080050 and NIH R01 DC007664 to D.L.M.; the Beckman Foundation to B.W.Z.; the National Science Foundation GRFP DGE-2034836 to J.L.M.; and an American Heart Association postdoctoral fellowship to F.A.-A. B.W.Z. is a Beckman Young Investigator. Some of this work was performed at the Stanford-SLAC Cryo-EM Center (S2C2), which is supported by the National Institutes of Health Common Fund Transformative High-Resolution Cryo-Electron Microscopy program (U24 GM129541). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Z.C. and D.L.M. conceived the study and designed the experiments. Z.C. and S.N. expressed and characterized the samples. Z.C., A.M. and F.A.-A. collected and analysed cryo-EM data. Z.C. and A.M. built and refined the atomic models. Z.C. collected and analysed the two-electrode voltage clamp data. F.A.-A. and S.J. collected and analysed the whole-cell electrophysiology data. J.L.M. and B.W.Z. collected and analysed the MS data. B.W.Z. and D.L.M. analysed data and provided guidance and support. Z.C., A.M., F.A.-A., J.L.M., B.W.Z. and D.L.M. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 EMC:CaV1.2(ΔC)/CaVβ3 and CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 Cryo-EM analysis.
a-d, Exemplars of purified CaV1.2(ΔC)/CaVβ3: a, SEC (Superose 6 Increase 10/300 GL). b, peak fraction SDS-PAGE. c, electron micrograph (~105,000x magnification), and d, 2D class averages. e–h Exemplars of purified CaV1.2(ΔC)/CaVβ3/CaVα2δ-1: e, SEC (Superose 6 Increase 10/300 GL). f, peak fraction SDS-PAGE. Magenta bars in ‘a’ and ‘e’ mark peak fraction. g, electron micrographs (~105,000x magnification), and h, 2D class averages. i, Workflow for electron microscopy data processing for the CaV1.2(ΔC)/CaVβ3 and CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 samples. Initial cryoSPARC-3.2 Ab initio reconstruction identified a population of particles containing the EMC:CaV1.2(ΔC)/CaVβ3 complex in the CaV1.2(ΔC)/CaVβ3 sample and populations of particles containing either the EMC:CaV1.2(ΔC)/CaVβ3 complex or CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 complex in the CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 sample. Red arrows indicate the three classes that were re-extracted, subjected to multiple rounds of 3D heterogeneous classification, and exported from cryoSPARC-3.2 for further 3D refinement in RELION-3.1. This resulted in two maps for the EMC:CaV1.2(ΔC)/CaVβ3 complex (ECAB Maps 1 and 2) and one for the CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 complex (CABAD Map 1). Multibody refinement was performed in RELION-3.1 to improve the features of flexible regions of the three maps. This resulted in the final map for the CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 complex (CABAD Map 2). ECAB Maps 1-2 with improved flexible features were merged (cross correlation = 0.9836) to obtain the final map for the EMC:CaV1.2(ΔC)/CaVβ3 complex (ECAB Map 3). Red boxes indicate the final maps used for model building. For ‘b-c’, N = 3. For ‘f-g’, N = 2.
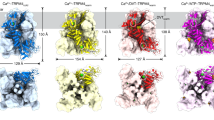
Extended Data Fig. 2 Cryo-EM maps of EMC:CaV1.2(ΔC)/CaVβ3 and CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 complexes.
EMC:CaV1.2(ΔC)/CaVβ3 complex a, side views and b, lumenal (left) and cytoplasmic (right) views. Subunits are coloured as: EMC1 (light blue), EMC2 (aquamarine), EMC3 (light magenta), EMC4 (Forest), EMC5 (light pink), EMC6 (white), EMC7 (marine), EMC8 (orange), EMC10 (smudge), CaV1.2 (bright orange), and CaVβ3 (lavender). CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 c, side views and d, extracellular (left) and cytoplasmic (right) views. Subunits are coloured as: CaV1.2 (slate), CaVβ3 (violet), and CaVα2δ-1 (greencyan). Detergent micelle is clear.
Extended Data Fig. 3 EMC:CaV1.2(ΔC)/CaVβ3 binding sites.
a, View of the TM dock interaction. EMC1 TM1 (slate) and CaV1.2 VSD I (yellow orange) interface. Interface buries 1051 Å2. Select elements and residues are indicated. EMC1 residues are in italics. b, LigPLOT92 diagram of EMC1 TM:CaV1.2 VSD I interactions showing ionic interactions (dashed lines) and van der Waals contacts ≤ 5Å. c, Sequence comparison of the indicated VSDI sequences for human CaV1.2 (HsCaV1.2 (109–182)) (Uniprot Q13936-20) with rabbit CaV1.1 (OcCaV1.1 (36–109)) (NCBI: NP_001095190.1), and human L-type (HsCaV1.1 (36–109), HsCaV1.3 (111–184), and HsCaV1.4 (77–150)) (NCBI: NP_000060.2, NP_000711.1, and NP_005174.2) and non-L-Type (HsCaV2.1 (83–156), HsCaV2.2 (80–153), and HsCaV2.3 (74–147)) (NCBI: NP_000059.3, NP_000709.1, and NP_001192222.1) channels. Red asterisks indicate residues involved in the cation-π pocket (120 and 123) and salt bridge (161). Red band highlights the residue that coordinates the Ca2+ ion in the CaVα2δ VWA domain. d, CaVβ3:EMC8 interaction. Callouts show the details of the indicated parts of the CaVβ3 NK loop interaction with EMC8. e, LigPLOT92 diagram of CaVβ3:EMC8 interactions showing ionic interactions (dashed lines) and van der Waals contacts ≤ 5Å. CaVβ:EMC8 hydrogen bond and salt bridge pairs are: Asp220:His208, Ser222:His208, Arg226:Glu166, Lys234:Thr8, Arg240:Asp56/Tyr87, Ser242:Lys204, and Gln279:His208. f, Sequence conservation for the indicated CaVβ elements from the EMC8 (top) and EMC2 (bottom) interaction sites. OcCaVβ3 (Uniprot P54286; 218–243, 277–282; 300–322); HsCaVβ1 (Uniprot Q02641.3; 270–295, 329–334; 352–374); HsCaVβ2 (Uniprot Q08289; 322–347, 381–386; 404–426); RnCaVβ2 (Uniprot Q8VGC3; 318–343, 377–382; 400–422); HsCaVβ3 (Uniprot P54284; 218–243, 277–282; 300–322); HsCaVβ4 (Uniprot O00305; 260–285, 319–324; 342–364). g, Superposition of rat CaVβ3 alone (violet, PDB:1VYU, chain B39) and CaVβ3 from the EMC complex. Boundaries of the disordered part of the T218-A243 loop in CaVβ3, and Q301, and ABP, are indicated, (RMSDCα = 1.39Å). h, LigPLOT92 diagram of CaVβ3:EMC2 interactions showing van der Waals contacts ≤ 5Å.
Extended Data Fig. 4 CaV1.2 structural details.
a–d, Structures of the indicated VSDs from the CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 complex. Gating charge residues, anionic counter charges (An1 and An2) and aromatic site of the charge transfer centre43,45,55 are shown. e, LigPLOT92 diagram of the CaVα2δ-1 leucine binding site showing hydrogen bonds and ionic interactions (dashed lines) and van der Waals contacts ≤ 5Å. f, LigPLOT92 diagram of blocking lipid: CaV1.2 showing van der Waals contacts ≤ 5Å. Domain I (yellow orange), Domain II (dark red), and Domain III (green) residues are indicated.
Extended Data Fig. 5 Conformational changes between EMC-bound and CaVα2δ-bound CaV1.2(ΔC)/CaVβ3.
Superposition of CaV1.2 from the EMC:CaV1.2(ΔC)/CaVβ3 and CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 complexes showing a, VSD conformational changes. b, PD conformational changes showing the superposition from ‘a’. Insets show each PD. Elements CaV1.2 from the EMC complex are: VSD I/PD I (yellow orange), VSD II/PD II (firebrick), VSD III/PD III (lime), VSD IV/PD IV (marine). CaV1.2 (slate) and CaVβ3 (violet) from CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 and CaVβ3 from the EMC (light teal) are semi-transparent. c, Superposition of CaVβ3 and the CaV1.2 AID helix from the EMC:CaV1.2(ΔC)/CaVβ3 and CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 complexes. Location of EMC8 is indicated by the orange oval. Red arrows in ‘a-c’ indicate conformational changes between CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 and EMC:CaV1.2(ΔC)/CaVβ3. d, Comparison of IIS0 and surrounding regions in the EMC complex (VSDII, firebrick; AID (yellow orange), and CaVβ3 (light teal)) their corresponding elements in CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 (slate). e, Interactions between VSD II:PD III in the CaVβ:AID:VSD II:PD III subcomplex from the EMC:CaV1.2(ΔC)/CaVβ3 structure.
Extended Data Fig. 6 CaV1.2 pore domain superposition for EMC-bound and CaVα2δ-bound CaV1.2(ΔC)/CaVβ3.
Pore domains from the EMC-bound complex are: PD I (yellow orange), PD II (firebrick), PD III (lime), and PD IV (marine). Pore domains from CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 are slate. Calcium ions are from CaV1.2(ΔC)/CaVβ3/CaVα2δ-1. Selectivity filter (SF), hydrophobic cavity, and inner gate regions and select residues are indicated.
Extended Data Fig. 7 Mutually exclusive interactions of the EMC holdase and CaVα2δ with the core CaV1.2/CaVβ3 complex and ordering of the CaV1.2 pore and VSDs by CaVα2δ.
a, Superposition of CaVα2δ-1 (semi-transparent, aquamarine) from the CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 structure with the EMC:CaV1.2/CaVβ3 complex. EMC1 surface is shown. Red oval highlights clash regions. Colours of the EMC:CaV1.2/CaVβ3 complex are as in Fig. 1a. Grey bars denote the membrane. b, Close up view of clash between EMC1 (light blue) and CaVα2δ-1 (aquamarine). EMC1, EMC3 (magenta), EMC5 (pink), EMC6 (white), VSD I (yellow orange), PD II (firebrick), and PD III (lime) from the EMC:CaV1.2/CaVβ3 complex are shown. Red oval highlights clash regions. CaVα2δ-1 domains are indicated. Divalent staple is indicated. SF calcium ions are from CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 and mark the location of the pore in the CaVα2δ-assembled channel. c, Superposition of VSD I (yellow orange), PD II (firebrick), and PD III (lime) from the EMC complex (semi-transparent) and their corresponding parts from CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 (slate). CaVα2δ-1 (aquamarine) is shown as a semi-transparent surface. Calcium ions are from CaV1.2(ΔC)/CaVβ3/CaVα2δ-1. Left inset shows the coordination of the divalent staple by the VWA domain MIDAS and D151 in CaV1.2(ΔC)/CaVβ3/CaVα2δ-1. Red distance shows the position of CaV1.2 D151 in the EMC complex relative to the calcium ion in the CaVα2δ complex. Asterisks mark positions where coordinated alanine mutation impair the ability of CaVα2δ to enhance CaV currents and surface expression59. Right inset shows the extensive contacts between PD II and PD III loops with CaVα2δ in the CaV1.2(ΔC)/CaVβ3/CaVα2δ-1 complex. The PD III loops are disordered in the EMC:CaV1.2/CaVβ3 complex. CaVα2δ-1 residues are in italics. d, Schematic showing of the conformational changes and interaction sites in the exchange between the EMC:CaV/CaVβ holdase complex and assembled CaV/CaVβ/CaVα2δ channel. Black ovals indicate key interaction sites in each complex.
Extended Data Fig. 8 Exemplar purification of CaV subunit combinations.
a-c Superose 6 Increase 10/300 GL chromatogram and peak fraction SDS-PAGE for: a, CaV1.2(ΔC)/CaVβ3 and b, CaV1.2(ΔC). c, Relative detection by mass spectrometry from ‘a’ and ‘b’ of EMC proteins with respect to CaV1.2(ΔC) across 3 replicates. d, Superose 6 Increase 10/300 GL chromatogram and peak fraction SDS-PAGE for CaVβ3. CaVβ3-NK and CaVβ3-SH3 are CaVβ3 proteolytic fragments. e, Absolute detection of EMC proteins and CaVβ3 by mass spectrometry following expression and purification of CaVβ3. f, Superose 6 Increase 10/300 GL chromatogram and peak fraction SDS-PAGE for CaV1.2/CaVβ3. Magenta bars in ‘a’ ‘b’, ‘d’, and ‘f’ mark peak fraction. g, Absolute detection of CaV1.2, CaV1.2(ΔC), CaVβ3, and EMC proteins from ‘a’ and ‘f’. Error bars are calculated as SEM. ND denotes not detected.
Extended Data Fig. 9 Functional and biochemical characterization of CaV mutants.
a, Voltage dependent activation (left) and I-V relationships (right) for the indicated channels. Parentheses indicate ‘n’ independent cells examined over >2 independent experiments. b, Superose 6 Increase 10/300 GL chromatogram and peak fraction SDS-PAGE for CaV1.2(ΔC)/CaVβ3(11A). Magenta bar marks peak fraction. c, Relative detection by mass spectrometry from ‘b’ of EMC proteins with respect to CaV1.2(ΔC) across 3 replicates. Data for CaV1.2(ΔC)/CaVβ3 are from Extended Data Fig. 8c. ‘ND’ indicates ‘not detected. Data in ‘a’ and ‘c’ are presented as mean ± SEM.
Supplementary information
Supplementary Information
Supplementary Figs. 1–9, the legends for Supplementary Videos 1–12, Supplementary Table 2 and Supplementary References.
Supplementary Table 1
Proteins identified by MS. Data for CaV1.2(ΔC)–CaVβ3, CaV1.2(ΔC) alone and CaVβ3 alone (a), CaV1.2–CaVβ3 (b) and CaV1.2(ΔC)–CaVβ3(11A) (c). Filters were applied as described in the Methods.
Supplementary Video 1
Overview of the EMC–CaV1.2–CaVβ3 complex.
Supplementary Video 2
Lumenal view of CaV1.2 conformational changes after EMC binding.
Supplementary Video 3
Side view of CaV1.2 conformational changes after EMC binding.
Supplementary Video 4
CaV1.2 VSD I conformational changes after EMC binding.
Supplementary Video 5
CaV1.2 VSD II conformational changes after EMC binding.
Supplementary Video 6
CaV1.2 VSD IV conformational changes after EMC binding.
Supplementary Video 7
CaV1.2 PD III conformational changes after EMC binding.
Supplementary Video 8
CaV1.2 PD II conformational changes after EMC binding.
Supplementary Video 9
CaV1.2 PD IV conformational changes after EMC binding.
Supplementary Video 10
EMC lumenal domain movement induced by client binding.
Supplementary Video 11
EMC lumenal domain movement induced by client binding.
Supplementary Video 12
EMC transmembrane domain movement induced by client binding.
Supplementary Data
Source data for Supplementary Figs. 2 and 3 and Supplementary Table 1
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, Z., Mondal, A., Abderemane-Ali, F. et al. EMC chaperone–CaV structure reveals an ion channel assembly intermediate. Nature 619, 410–419 (2023). https://doi.org/10.1038/s41586-023-06175-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06175-5
This article is cited by
-
A tale of two calcium channels: structural pharmacology of Cav2.1 and Cav3.2
Cell Research (2024)
-
Structural bases of inhibitory mechanism of CaV1.2 channel inhibitors
Nature Communications (2024)
-
EMC rectifies the topology of multipass membrane proteins
Nature Structural & Molecular Biology (2024)
-
Fighting pain: the structure of gabapentin and its binding site in the Cavα2δ subunit
Nature Structural & Molecular Biology (2023)
-
Expulsion mechanism of the substrate-translocating subunit in ECF transporters
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.