Abstract
Host factors that mediate Leishmania genetic exchange are not well defined. Here we demonstrate that natural IgM (IgMn)1,2,3,4 antibodies mediate parasite genetic exchange by inducing the transient formation of a spherical parasite clump that promotes parasite fusion and hybrid formation. We establish that IgMn from Leishmania-free animals binds to the surface of Leishmania parasites to induce significant changes in the expression of parasite transcripts and proteins. Leishmania binding to IgMn is partially lost after glycosidase treatment, although parasite surface phosphoglycans, including lipophosphoglycan, are not required for IgMn-induced parasite clumping. Notably, the transient formation of parasite clumps is essential for Leishmania hybridization in vitro. In vivo, we observed a 12-fold increase in hybrid formation in sand flies provided a second blood meal containing IgMn compared with controls. Furthermore, the generation of recombinant progeny from mating hybrids and parental lines were only observed in sand flies provided with IgMn. Both in vitro and in vivo IgM-induced Leishmania crosses resulted in full genome hybrids that show equal patterns of biparental contribution. Leishmania co-option of a host natural antibody to facilitate mating in the insect vector establishes a new paradigm of parasite–host–vector interdependence that contributes to parasite diversity and fitness by promoting genetic exchange.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data are present in the main text or the supplementary materials. Whole-genome and transcriptome sequencing raw reads files have been deposited into the NCBI BioProject database with the identifier PRJNA988832. Proteome sequencing raw files have been deposited into the PRIDE database with the project identifier PXD038652. Source data are provided with this paper.
Code availability
Codes generated on this work used for copy number bioinformatics analysis are available at https://zenodo.org/record/8231973 and https://github.com/jlac/Leishmania_allelic_copy_number. Codes generated for this work used for statistical analysis are available at https://github.com/joedoehl/IgM-promotes-genetic-exchange-of-Leishmania.
References
Rapaka, R. R. et al. Conserved natural IgM antibodies mediate innate and adaptive immunity against the opportunistic fungus Pneumocystis murina. J. Exp. Med. 207, 2907–2919 (2010).
Boehm, T., Iwanami, N. & Hess, I. Evolution of the immune system in the lower vertebrates. Annu. Rev. Genomics Hum. Genet. 13, 127–149 (2012).
Panda, S. & Ding, J. L. Natural antibodies bridge innate and adaptive immunity. J. Immunol. 194, 13–20 (2015).
Mashoof, S. & Criscitiello, M. F. Fish immunoglobulins. Biology 5, 45 (2016).
Serafim, T. D. et al. Leishmaniasis: the act of transmission. Trends Parasitol. 37, 976–987 (2021).
Burza, S., Croft, S. L. & Boelaert, M. Leishmaniasis. Lancet 392, 951–970 (2018).
Alvar, J. et al. Leishmaniasis worldwide and global estimates of its incidence. PLoS ONE 7, e35671 (2012).
Tibayrenc, M., Kjellberg, F. & Ayala, F. J. A clonal theory of parasitic protozoa: the population structures of Entamoeba, Giardia, Leishmania, Naegleria, Plasmodium, Trichomonas, and Trypanosoma and their medical and taxonomical consequences. Proc. Natl Acad. Sci. USA 87, 2414–2418 (1990).
Tibayrenc, M. & Ayala, F. J. The clonal theory of parasitic protozoa: 12 years on. Trends Parasitol. 18, 405–410 (2002).
Tibayrenc, M. & Ayala, F. J. Is predominant clonal evolution a common evolutionary adaptation to parasitism in pathogenic parasitic protozoa, fungi, bacteria, and viruses? Adv. Parasitol. 97, 243–325 (2017).
Akopyants, N. S. et al. Demonstration of genetic exchange during cyclical development of Leishmania in the sand fly vector. Science 324, 265–268 (2009).
Rogers, M. B. et al. Genomic confirmation of hybridisation and recent inbreeding in a vector-isolated Leishmania population. PLoS Genet. 10, e1004092 (2014).
Inbar, E. et al. Whole genome sequencing of experimental hybrids supports meiosis-like sexual recombination in Leishmania. PLoS Genet. 15, e1008042 (2019).
Van den Broeck, F. et al. Ecological divergence and hybridization of neotropical Leishmania parasites. Proc. Natl Acad. Sci. USA 117, 25159–25168 (2020).
Stelzer, C. P. & Lehtonen, J. Diapause and maintenance of facultative sexual reproductive strategies. Philos. Trans. R. Soc. Lond. B Biol. Sci. 371, 20150536 (2016).
Kwok, A. J., Mentzer, A. & Knight, J. C. Host genetics and infectious disease: new tools, insights and translational opportunities. Nat. Rev. Genet. 22, 137–153 (2021).
Inbar, E. et al. The mating competence of geographically diverse Leishmania major strains in their natural and unnatural sand fly vectors. PLoS Genet. 9, e1003672 (2013).
Romano, A. et al. Cross-species genetic exchange between visceral and cutaneous strains of Leishmania in the sand fly vector. Proc. Natl Acad. Sci. USA 111, 16808–16813 (2014).
Louradour, I., Ferreira, T. R., Ghosh, K., Shaik, J. & Sacks, D. In vitro generation of Leishmania hybrids. Cell Rep. 31, 107507 (2020).
Louradour, I. et al. Stress conditions promote Leishmania hybridization in vitro marked by expression of the ancestral gamete fusogen HAP2 as revealed by single-cell RNA-seq. eLife 11, e73488 (2022).
Navin, T. R., Krug, E. C. & Pearson, R. D. Effect of immunoglobulin M from normal human serum on Leishmania donovani promastigote agglutination, complement-mediated killing, and phagocytosis by human monocytes. Infect. Immun. 57, 1343–1346 (1989).
Reyneveld, G. I., Savelkoul, H. F. J. & Parmentier, H. K. Current understanding of natural antibodies and exploring the possibilities of modulation using veterinary models. A review. Front. Immunol. 11, 2139 (2020).
Eyayu, T. et al. Evaluation of urine sample for diagnosis of visceral leishmaniasis using rK-39 immunochromatographic test in Northwest Ethiopia. PLoS ONE 17, e0263696 (2022).
Spath, G. F. et al. Persistence without pathology in phosphoglycan-deficient Leishmania major. Science 301, 1241–1243 (2003).
Silva, R. & Sacks, D. L. Metacyclogenesis is a major determinant of Leishmania promastigote virulence and attenuation. Infect. Immun. 55, 2802–2806 (1987).
Walters, L. L. Leishmania differentiation in natural and unnatural sand fly hosts. J. Eukaryot. Microbiol. 40, 196–206 (1993).
Zhang, J., Zhang, L., Wang, J., Ouyang, L. & Wang, Y. Polo-like kinase 1 inhibitors in human cancer therapy: development and therapeutic potential. J. Med. Chem. 65, 10133–10160 (2022).
Olah, J. et al. Tubulin binding and polymerization promoting properties of tubulin polymerization promoting proteins are evolutionarily conserved. Biochemistry 56, 1017–1024 (2017).
Sahasrabuddhe, A. A., Nayak, R. C. & Gupta, C. M. Ancient Leishmania coronin (CRN12) is involved in microtubule remodeling during cytokinesis. J. Cell Sci. 122, 1691–1699 (2009).
Yanase, R. et al. Formation and three-dimensional architecture of Leishmania adhesion in the sand fly vector. eLife 12, e84552 (2023).
Keyt, B. A., Baliga, R., Sinclair, A. M., Carroll, S. F. & Peterson, M. S. Structure, function, and therapeutic use of IgM antibodies. Antibodies 9, 53 (2020).
Serafim, T. D. et al. Sequential blood meals promote Leishmania replication and reverse metacyclogenesis augmenting vector infectivity. Nat. Microbiol. 3, 548–555 (2018).
Volfova, V. & Volf, P. The salivary hyaluronidase and apyrase of the sand fly Sergentomyia schwetzi (Diptera, Psychodidae). Insect Biochem. Mol. Biol. 102, 67–74 (2018).
Burkett-Cadena, N. D. in Medical and Veterinary Entomology 3rd edn (eds Mullen, G. R. & Durden, L. A.) 17–22 (Academic Press, 2019).
Lee, S. H. et al. Mannose receptor high, M2 dermal macrophages mediate nonhealing Leishmania major infection in a Th1 immune environment. J. Exp. Med. 215, 357–375 (2018).
Chaves, M. M. et al. The role of dermis resident macrophages and their interaction with neutrophils in the early establishment of Leishmania major infection transmitted by sand fly bite. PLoS Pathog. 16, e1008674 (2020).
Prieto Barja, P. et al. Haplotype selection as an adaptive mechanism in the protozoan pathogen Leishmania donovani. Nat. Ecol. Evol. 1, 1961–1969 (2017).
Negreira, G. H. et al. High throughput single-cell genome sequencing gives insights into the generation and evolution of mosaic aneuploidy in Leishmania donovani. Nucleic Acids Res. 50, 293–305 (2022).
Goodenough, U. & Heitman, J. Origins of eukaryotic sexual reproduction. Cold Spring Harb. Perspect. Biol. 6, a016154 (2014).
Hayakawa, K., Hardy, R. R. & Herzenberg, L. A. Peritoneal Ly-1 B cells: genetic control, autoantibody production, increased lambda light chain expression. Eur. J. Immunol. 16, 450–456 (1986).
Haury, M. et al. The repertoire of serum IgM in normal mice is largely independent of external antigenic contact. Eur. J. Immunol. 27, 1557–1563 (1997).
Hayakawa, K. et al. Positive selection of natural autoreactive B cells. Science 285, 113–116 (1999).
Bendelac, A., Bonneville, M. & Kearney, J. F. Autoreactivity by design: innate B and T lymphocytes. Nat. Rev. Immunol. 1, 177–186 (2001).
Khasbiullina, N. R. & Bovin, N. V. Hypotheses of the origin of natural antibodies: a glycobiologist’s opinion. Biochemistry 80, 820–835 (2015).
Maddur, M. S. et al. Natural antibodies: from first-line defense against pathogens to perpetual immune homeostasis. Clin. Rev. Allergy Immunol. 58, 213–228 (2020).
Oliveira, F. et al. A sand fly salivary protein vaccine shows efficacy against vector-transmitted cutaneous leishmaniasis in nonhuman primates. Sci. Transl. Med. 7, 290ra290 (2015).
Aslan, H. et al. A new model of progressive visceral leishmaniasis in hamsters by natural transmission via bites of vector sand flies. J. Infect. Dis. 207, 1328–1338 (2013).
Spath, G. F. et al. Lipophosphoglycan is a virulence factor distinct from related glycoconjugates in the protozoan parasite Leishmania major. Proc. Natl Acad. Sci. USA 97, 9258–9263 (2000).
Spath, G. F., Garraway, L. A., Turco, S. J. & Beverley, S. M. The role(s) of lipophosphoglycan (LPG) in the establishment of Leishmania major infections in mammalian hosts. Proc. Natl Acad. Sci. USA 100, 9536–9541 (2003).
Capul, A. A., Barron, T., Dobson, D. E., Turco, S. J. & Beverley, S. M. Two functionally divergent UDP-Gal nucleotide sugar transporters participate in phosphoglycan synthesis in Leishmania major. J. Biol. Chem. 282, 14006–14017 (2007).
Sacks, D. L. & Perkins, P. V. Identification of an infective stage of Leishmania promastigotes. Science 223, 1417–1419 (1984).
Guo, H. et al. Genetic metabolic complementation establishes a requirement for GDP-fucose in Leishmania. J. Biol. Chem. 292, 10696–10708 (2017).
Vickers, T. J., Murta, S. M., Mandell, M. A. & Beverley, S. M. The enzymes of the 10-formyl-tetrahydrofolate synthetic pathway are found exclusively in the cytosol of the trypanosomatid parasite Leishmania major. Mol. Biochem. Parasit. 166, 142–152 (2009).
Serafim, T. D. & Valenzuela, J. G. A detailed procedure to maximize IgM-induced Leishmania major mating in vitro. Protoc. Exch. https://doi.org/10.21203/rs.3.pex-2345/v1 (2023).
Afonso, L. C. et al. The adjuvant effect of interleukin-12 in a vaccine against Leishmania major. Science 263, 235–237 (1994).
Rogers, M. E., Ilg, T., Nikolaev, A. V., Ferguson, M. A. & Bates, P. A. Transmission of cutaneous leishmaniasis by sand flies is enhanced by regurgitation of fPPG. Nature 430, 463–467 (2004).
Dostalova, A. & Volf, P. Leishmania development in sand flies: parasite–vector interactions overview. Parasit. Vector 5, 276 (2012).
Cecilio, P. et al. Exploring Lutzomyia longipalpis sand fly vector competence for Leishmania major parasites. J. Infect. Dis. 222, 1199–1203 (2020).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Zaqout, S., Becker, L. L. & Kaindl, A. M. Immunofluorescence staining of paraffin sections step by step. Front. Neuroanat. 14, 582218 (2020).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Faust, G. G. & Hall, I. M. SAMBLASTER: fast duplicate marking and structural variant read extraction. Bioinformatics 30, 2503–2505 (2014).
Shaik, J. S., Dobson, D. E., Sacks, D. L. & Beverley, S. M. Leishmania sexual reproductive strategies as resolved through computational methods designed for aneuploid genomes. Genes 12, 167 (2021).
Okonechnikov, K., Conesa, A. & Garcia-Alcalde, F. Qualimap 2: advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics 32, 292–294 (2016).
Coutinho-Abreu, I. V. et al. Distinct gene expression patterns in vector-residing Leishmania infantum identify parasite stage-enriched markers. PLoS Negl. Trop. Dis. 14, e0008014 (2020).
Zhao, T. & Wang, Z. GraphBio: a shiny web app to easily perform popular visualization analysis for omics data. Front. Genet. 13, 957317 (2022).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Marques, C. A., Dickens, N. J., Paape, D., Campbell, S. J. & McCulloch, R. Genome-wide mapping reveals single-origin chromosome replication in Leishmania, a eukaryotic microbe. Genome Biol. 16, 230 (2015).
Plate, T. & Heiberger, R. abind: Combine Multidimensional Arrays. R package version 1.4-5 https://CRAN.R-project.org/package=abind (2016).
James, D. & Hornik, K. chron: Chronological Objects which Can Handle Dates and Times. R package version 2.3-56 https://cran.r-project.org/web/packages/chron/index.html (2020).
Dowle, M. & Srinivasan, A. data.table: Extension of ‘data.frame‘. R package version 1.14.0 https://CRAN.R-project.org/package=data.table (2021).
Meyer, D., Dimitriadou, E., Hornik, K., Weingessel, A. & Leisch, F. e1071: Misc Functions of the Department of Statistics, Probability Theory Group (Formerly: E1071), TU Wien. R package version 1.7-8 https://CRAN.R-project.org/package=e1071 (2021).
Lenth, R. V. emmeans: Estimated Marginal Means, aka Least-Squares Means. R package version 1.6.2-1 https://CRAN.R-project.org/package=emmeans (2021).
Bocinsky, R. K. FedData: Functions to Automate Downloading Geospatial Data Available from Several Federated Data Sources. R package version 2.5.7 https://CRAN.R-project.org/package=FedData (2019).
Grothendieck, G. gsubfn: Utilities for Strings and Function Arguments. R package version 0.7 https://CRAN.R-project.org/package=gsubfn (2018).
Iannone, R., Cheng, J. & Schloerke, B. gt: Easily Create Presentation-Ready Display Tables. R package version 0.3.1 https://CRAN.R-project.org/package=gt (2021).
Nolan, R. & Padilla-Parra, S. ijtiff: an R package providing TIFF I/O for ImageJ users. J. Open Source Softw. 3, 663 (2018).
Firke, S. janitor: Simple Tools for Examining and Cleaning Dirty Data. R package version 2.1.0 https://CRAN.R-project.org/package=janitor (2021).
Grolemund, G. & Wickham, H. Dates and times made easy with lubridate. J. Stat. Softw. 40, 1–25 (2011).
Venables, W. N. & Ripley, B. D. Modern Applied Statistics with S 4th edn (Springer, 2002).
Wickham, H. The aplit-apply-combine strategy for data analysis. J. Stat. Softw. 40, 1–29 (2011).
Mangiafico, S. rcompanion: Functions to Support Extension Education Program Evaluation. R package version 2.4.1 https://CRAN.R-project.org/package=rcompanion (2021).
Wickham, H. & Bryan, B. readxl: Read Excel Files. R package version 1.3.1 https://CRAN.R-project.org/package=readxl (2019).
Wickham, H. Reshaping data with the reshape Package. J. Stat. Softw. 21, 1–20 (2007).
Bengtsson, H. R.utils: Various Programming Utilities. R package version 2.10.1 https://CRAN.R-project.org/package=R.utils (2020).
Ren, K. rlist: A Toolbox for Non-Tabular Data Manipulation. R package version 0.4.6.1 https://CRAN.R-project.org/package=rlist (2016).
Hope, R. M. Rmisc: Rmisc: Ryan Miscellaneous. R package version 1.5 https://CRAN.R-project.org/package=Rmisc (2013).
Kassambara, A. rstatix: Pipe-Friendly Framework for Basic Statistical Tests. R package version 0.7.0 https://CRAN.R-project.org/package=rstatix (2021).
The R Development Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2021).
Wickham, H. stringr: Simple, Consistent Wrappers for Common String Operations. R package version 1.4.0 https://CRAN.R-project.org/package=stringr (2019).
Wickham, H. et al. Welcome to the tidyverse. J. Open Source Softw. 4, 1686 (2019).
Mair, P. & Wilcox, R. Robust statistical methods in R using the WRS2 package. Behav. Res. Methods 52, 464–488 (2020).
Dragulescu, A. & Arendt, C. xlsx: Read, Write, Format Excel 2007 and Excel 97/2000/XP/2003 Files. R package version 0.6.5 https://CRAN.R-project.org/package=xlsx (2020).
Acknowledgements
We would like to thank Y. Rangel Gonzalez, R. Salas Carrillo and B. Mills from VMBS, NIAID for technical support; S. Doe and D. Akueson from VMBS, NIAID for sand fly insectary support; C. Percopo from LMVR, NIAID for flow cytometry support; A. de Souza from the Department of Medicine, Division of Nephrology and Hypertension, Georgetown University for critical reading of the manuscript; G. A. Nardone, M. Suzuki and L. R. Olano of the Research Technologies Branch, NIAID, for MS analysis; C. Brodskin from Fiocruz, Bahia, Brazil, and P. Lima dos santos, Programa de Pós-graduação em Ciencias da Saúde (Graduate Program in Health Science), Federal University of Sergipe for Leishmania-infected human serum; C. Petersen from the University of Iowa, USA, for Leishmania-infected canine serum; and S. Kuhn of the NCBR Research Technologies Branch, NIAID, for support with RNA sequencing analysis. This research was supported by the Intramural Research Program of the NIH, National Institute of Allergy and Infectious Diseases, and NIH grant R01-AI29646 to S.M.B. The schematics in Figs. 4b and 5a and Extended Data Figs. 2a, 8a,c and 9a were created using BioRender (https://biorender.com).
Author information
Authors and Affiliations
Contributions
T.D.S. developed the hypothesis. T.D.S., S.K., J.G.V., S.M.B., I.V.C.-A., C.B.-M., J.A. and F.O. contributed to experimental design. E.I. planned the experiments. T.D.S., E.I., J.S.P.D., A.B.F.B., J.A., M.D., V.N., M.S., P.C., T.W. and T.L.A.S. performed the experiments. T.D.S., J.M.C.R., J.S.P.D., J.L., A.B.F.B., J.G.V., S.K., S.M.B., E.I., T.L.A.S. and J.V.-R. analysed the data. C.M. performed sand fly insectary work. J.G.V., S.K. and S.M.B. supervised the project. All authors wrote the manuscript. P.C. performed the in vitro IgM experiment with two different Leishmania species, L. major and L. tropica, and was able to replicate the IgM-mediated hybridization process five times, independently. A.B.F.B. and P.C. contributed equally.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Álvaro Acosta-Serrano, Jeremy Mottram and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Identification of IgM in blood serum as the Leishmania clumping factor.
(a) Gel filtration chromatography of inactivated dog plasma on Sephacryl S-200. Buffer: 20 mM sodium phosphate (pH 7.4), 150 mM NaCl, 5 mM EDTA. Bar represents HPLC fractions corresponding to Leishmania clumping activity. (b) SDS-PAGE of fractions obtained from gel filtration chromatography shown in panel A. Bar represents fractions containing clumping activity that were pooled for ion exchange chromatography. (c) Ion exchange chromatography of pooled fractions 11–13 from B. HPLC gradient, a sodium chloride gradient of 0 – 1 M in 20 mM sodium phosphate, pH 6.0. Bar represents fractions pooled for high resolution gel filtration chromatography. (d) High-resolution gel filtration chromatography on Superdex-200. Buffer: 20 mM sodium phosphate (pH 7.4), 150 mM NaCl. Bar represents fractions pooled for protein identification. (e) Mass spectrum analysis of the most abundant proteins in pooled fractions from (d).
Extended Data Fig. 2 IgM promotes Leishmania hybrid formation in vitro.
(a) Workflow for in vitro crossing of Leishmania parental lines (L. major or L. tropica). Mating wells were prepared with 50 µg/mL IgM in complete Schneider’s media. An improved detailed step-by-step procedure to maximize IgM-induced Leishmania major mating in vitro is available at https://doi.org/10.21203/rs.3.pex-2345/v1. (b) Data summary of hybrid formation from in vitro crossing of L. major parental lines: Parental 1, WR-SSU-HYG; Parental 2, FVI-FKP40-BSD; parental 3, FVI-FTL-SAT. L. major hybrids genotyping by PCR targeting parental selectable drug markers available at Fig. 1h. (c) Data summary of hybrid formation from in vitro crossing of L. tropica parental lines: Parental 4, K27-SSU-HYG; Parental 5, K27-SSU-SAT. (d) L. tropica hybrids genotyping by PCR targeting parental selectable drug markers HYG, and SAT. Parental 4, K27-SSU-HYG; Parental 5, K27-SSU-SAT; ntc, no template control; L, 1kb-plus ladder.
Extended Data Fig. 3 IgM natural antibodies and sera from naive animals bind to Leishmania parasites.
(a,b) Sera from naive or Leishmania-infected adult mammals were used as indicated. IgG (a) or IgM from naïve serum (b) ELISA performed using recombinant K39 (rK39) protein or Leishmania lysate (LL) as target antigens. Statistical analysis by Mann-Whitney test between groups, *P < 0.05, **P ≤ 0.01. n = 4. (c) Leishmania clumping pattern 2 h after incubation with naïve or Leishmania-infected sera. L. major parasites were incubated in complete Schneider’s supplemented with 5% naïve or Leishmania-infected sera from adult mammals as indicated (n = 4-5). Scale bars = 50 µm. (d) Western blot of L. major enriched cell wall membrane preparation incubated with naïve adult human sera detected by anti-IgM (left panel). Right panel, L. major enriched cell wall membrane preparation incubated with only secondary antibody as control. n = 2. (e) Purified naïve IgM (IgMn) binding to Leishmania performed ELISA using LL or a negative control protein (bovine serum albumin) as target antigens. Statistical analysis by Mann-Whitney test between groups, ****P ≤ 0.0001. n = 3. (f) PNA agglutination does not promote Leishmania hybrid formation. Images taken 12 h after incubating parasites with varying concentrations of PNA. Scale bars = 50 µm. (g) Data summary of hybrid formation from in vitro crossing of L. major parental lines 1 and 2. Mating wells were prepared with 50 µg/mL IgM or with 6.25, 12.5, 25 and 50 µg/mL PNA in complete Schneider’s media. ELISAs data represented by individual values scatter plots. Dashed bars, medians.
Extended Data Fig. 4 IgM promotes Leishmania mating clump formation and hybridization inside the mating clump.
(a) SEM of L. major (upper panel) and L. tropica (lower panel) mating clump showing the 3D organization of promastigote forms. Scale bar, 15 µm. (b) TEM of a L. major mating clump showing a cell with 3 nuclei. Scale bar, 3 µm. (c) TEM images of a L. tropica mating clump showing cells with 3 nuclei. Scale bars: 5 µm. Arrows point to individual nuclei within a parasite cell body. Three-nucleated cells are suggestive of an earlier step that precedes the merging of nuclei. The cell cycle progression follows a strict pattern were Nucleus, Kinetoplast, Flagellum (NKF) division is always constant and goes from 1N1K1F to 2N2K2F culminating with cytokinesis. Note the presence of 3 nuclei without changes to either the kinetoplast or flagellum suggestive of a potential early fusion event inside the LMC. (d, e) Leishmania mating clump formation within the sand fly midgut. Twelve days post-infection with L. major; Lu. longipalpis females were provided with a second reconstituted blood meal containing IgM (500 µg/mL). (d) Initiation of Leishmania mating clump (LMC) formation 30 min after imbibing a second blood meal (blue circles). (e) Parasites merge to form larger LMCs within 24 h after imbibing a second blood meal. Scale bars = 50 μm. LMCs formation in sand flies can also be observed in Supplementary Videos 11 and 12. (f) Confocal immunofluorescence of a 5 µm transversal section from L. tropica mating clump. Promastigotes were grown to stationary phase in the presence of EdU or BrdU then extensively washed and used at a 1:1 ratio to assemble the LMC with IgMn. EdU, green; BrdU, red. White arrows point to yellow nuclei indicative of fusion and exchanged genetic material. Scale bars, 3 µm.
Extended Data Fig. 5 Hybrid genotyping and infection status in sand flies after a naturally acquired Leishmania infection from mice lesions composed of two parental lines.
(a) Leishmania major hybrids formed in sand flies given one infectious blood meal (BM1) or provided 6 days later with an additional uninfected bloodmeal (BM2) in the presence (+IgM) or absence (−IgM) of IgM were genotyped by PCR targeting parental selectable drug markers HYG (Hygromycin), BSD (Blasticidin) and SAT (Nourseothricin). Parental 1, WR-SSU-HYG; Parental 2, FVI-FKP40-BSD; Parental 3, FVI-FTL-SAT; ntc, no template control; L, 1kb-plus ladder. Double drug resistant hybrid lines were cloned before genotyping. A single hybrid exemplar is shown for positive events from each group. “n” detailed on Fig. 4d. Parasite number (b,c,d) and percentage of metacyclic promastigotes (e,f,g) in L. major-infected Lu. longipalpis. At 6 days post-infection, a proportion of the sand flies were provided a second uninfected blood meal via a membrane feeder composed of rabbit red blood cells reconstituted with fetal bovine serum with (+IgM) or without (−IgM) 500 µg/ml adult bovine IgM or allowed to feed on uninfected mice as a blood source for natural IgM. Infection status of individual sand flies was assessed at 14 days after the first blood meal (8 days after the second bloodmeal). Parental line combination 1×2 (b,e) n = 4, 1×3 (c,f) n = 3, 2×3 (d,g) n = 3. data represented by individual values scatter plots. Dashed bars, medians. (h) An antibiotic cocktail (ABTs) was used to control contamination with sand fly gut microbiota when isolating parasites from sand fly midguts. ABTs has no effect on parasite growth and viability. ABTs: Penicillin-Streptomycin (100 U/mL); Gentamicin (50 µg/mL); Caspofungin (15 µg/mL); 5-fluorocytosine (30 µg/mL). Growth curve represented by median ± interquartile range. n = 3.
Extended Data Fig. 6 Resistance markers whole-genome analysis of parental and hybrid lines.
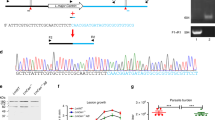
Schematic display of an artificial chromosome containing arbitrary loci of the three antibiotic resistance genes (Hygromycin – HYG, Blasticidin – BSD, Nourseothricin – SAT) used as parental markers in this study. (a) Leishmania major F1 hybrids. (b) L. major backcross hybrids. (c) L. tropica F1 hybrids. Raw reads were processed and displayed using Geneious prime software v2021.2.2. Upper case letters in green provide a unique identifier for each Leishmania parental or hybrid. Red abbreviations present on first panel of (a) are applicable to all panels in the figure: Cov, coverage; r-Chr, artificial resistance chromosome; A-r, aligned reads. Panel I was repeated from (a) to (b) for illustrative purposes as this was the selected F1 hybrid for backcrosses.
Extended Data Fig. 7 Whole-genome analysis of Leishmania major hybrids.
(a, b) Allele-specific chromosomal copy number as inferred by read depth and allelic proportions. Biparental ancestry was confirmed across the whole genome for crosses between parentals 1×2 and backcrosses of (1×2) F1 hybrid with parental 2′ or parental 2″ (a) and a parental 1×3 cross (b). Biparental inheritance of backcrosses for 1×2 crosses are also shown in Fig. 5d. Upper case letters provide a unique identifier for each Leishmania parental or hybrid. (2×3) crosses not present because within strains there were very few fixed differences between the parents, and therefore not enough informative sites to call ancestry. Parental 1 plot was repeated from (a) to (b) for illustrative purposes as this parental composed both F1 crosses setup. (c) Somy heatmap of sequenced Leishmania major parentals and hybrids. Whole-genome analysis shows characteristic Leishmania major aneuploidy chromosome distribution in all samples. (a-c) Parental 1, WR-SSU-HYG; Parental 2, FVI-FKP40-BSD; Parental 3, FVI-FTL-SAT. Parental 2′, FV1-FKP40-SAT; Parental 2″, FV1-SSU-SAT. Upper case letters provide a unique identifier of each Leishmania parental or hybrid.
Extended Data Fig. 8 Evaluation of parental lines and hybrids in infected mouse tissue.
(a) Experimental design. Mouse footpads were injected with a 1:1 mixture of two L. major parental line combinations 1 and 2, 1 and 3, or 2 and 3. After lesion development, 3 to 5 weeks post injection, infected footpads were exposed to sand flies for their first blood meal. Tissue from the infected footpad (F) and draining lymph node (LN) were then disrupted and seeded in complete Schneider’s media (6 mL) for 3 days to allow for amastigote differentiation into promastigotes. Half of the material was then cultured in double drug pressure for selection of hybrids. (b) The second half of the tissue from (a) was used for DNA extraction to confirm the presence of both parental lines by genotyping. n = 12 (1×2), 9 (1×3 and 2×3). (c) Experimental design. Mouse footpads ware injected with a 1:1 mixture of (1×2) F1 hybrid and parental 2 harboring a different resistance marker (SAT) at the same (2′) or different (2″) chromosome locus. Sand fly feeding on mouse lesions and tissue processing were carried out as outlined above. Half of the tissue from the footpad and draining lymph node was then cultured in triple drug pressure for selection of hybrids. (d) The second half of the tissue from (c) was assessed for the presence of both parental lines by genotyping. n = 9 [(1×2)x2′], 6 [(1×2)×2″]. Parental 1, WR-SSU-HYG; Parental 2, FV1-FKP40-BSD; Parental 3 = FV1-FTL-SAT; Parental 2′, FV1-FKP40-SAT; Parental 2″, FV1-SSU-SAT.
Extended Data Fig. 9 IgM promotes Leishmania backcross hybrids in the gut of sand flies.
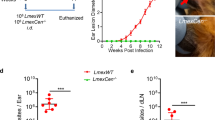
(a) Diagram outlining the generation of backcrosses. Sand flies were fed on mice lesions composed of a parental and a hybrid line. L. major Parental 1 (WR-SSU-HYG) and Parental 2 (FVI-FKP40-BSD) resistant to hygromycin (HYG) or blasticidin (BSD), respectively, were crossed to produce F1 hybrids. HYG/BSD double resistant F1 hybrids were backcrossed to L. major Parental 2′ (FV1-FKP40-SAT) or 2″ (FV1-SSU-SAT), both resistant to Nourseothricin (SAT) inserted at loci in chromosome 16 and 27, respectively. This resulted in the recovery of backcross hybrids resistant to HYG/BSD/SAT. (b) Leishmania major backcross hybrid genotyping by PCR targeting parental selectable drug markers HYG, BSD and SAT. (1×2) F1 hybrid (a cross between L. major Parental 1 and Parental 2); Parental 2′, FV1-FKP40-SAT; Parental 2″, FV1-SSU-SAT.; ntc, no template control; L, 1kb-plus ladder. Triple drug resistant hybrid lines were cloned before genotyping. Only sand flies provided a second bloodmeal containing IgM produced backcross hybrids. A single backcross hybrid exemplar is shown for positive events from each group. “n” detailed on Fig. 5c. Parasite number (c,d) and percentage of metacyclic promastigotes (e,f) in L. major-infected Lu. longipalpis. At 6 days post-infection, a proportion of the sand flies were provided a second uninfected blood meal via a membrane feeder composed of rabbit red blood cells reconstituted with fetal bovine serum with (+IgM) or without (−IgM) 500 µg/ml adult bovine IgM. Infection status of individual sand flies was assessed at 14 days after the first blood meal (8 days after the second bloodmeal). Parental line combination (1×2)×2′ (c,e) n = 3; (1×2)x2′ (d,f) n = 2.
Extended Data Fig. 10 Ploidy histograms of sequenced Leishmania parentals and hybrids.
Cloned parasite lines were evaluated by FACS to determine DNA content. Parental 1, WR-SSU-HYG; Parental 2, FVI-FKP40-BSD; Parental 3, FVI-FTL-SAT. Parental 2′, FV1-FKP40-SAT; Parental 2″, FV1-SSU-SAT; Parental 4, K27-SSU-HYG; Parental 5, K27-SSU-SAT. Ploidy indicated in red uppercase number/letter. Upper case green letters on lower right corner provide a unique identifier of each Leishmania parental or hybrid.
Supplementary information
Supplementary Information
A full guide to the Supplementary information, Supplementary Figs. 1 and 2 and Supplementary Tables 3–7.
Supplementary Table 1
Transcripts from RNA sequence of IgM-treated or untreated (control) L. major.
Supplementary Table 2
Proteins from proteomics analysis of IgM-treated or untreated (control) L. major.
Supplementary Table 8
Proteomics raw files metadata.
Supplementary Data 1
Supplementary sequence 1: nucleotide sequence of the artificial chromosome used for Extended Data Fig. 6.
Supplementary Video 1
L. major metacyclic promastigotes in culture medium supplemented with 20% FBS and 5% inactivated adult dog serum. Video taken 30 min after seeding. Event 1. Purified metacyclic promastigotes were used to exclude in vitro multiplication rosettes and for clear visualization of parasite–parasite interaction. Scale bar, 50 μm.
Supplementary Video 2
L. major metacyclic promastigotes in culture medium supplemented with 20% FBS and 5% inactivated adult dog serum. Video taken 30 min after seeding. Event 2. Scale bar, 50 μm.
Supplementary Video 3
L. major metacyclic promastigotes in culture medium supplemented with 20% FBS and 5% inactivated adult dog serum. Video taken 180 min after seeding. Scale bar, 20 μm.
Supplementary Video 4
L. major stationary phase promastigotes in culture medium supplemented with 20% FBS and bovine IgM (50 μg ml–1). Video taken 24 h after seeding. Scale bar, 50 μm.
Supplementary Video 5
L. major metacyclic promastigotes in culture medium supplemented with 20% FBS. Video taken 30 min after seeding. Scale bar, 50 μm.
Supplementary Video 6
L. major stationary phase promastigotes in culture medium supplemented with 20% FBS. Video taken 24 h after seeding. Scale bar, 50 μm.
Supplementary Video 7
L. major stationary phase promastigotes in culture medium supplemented with 20% FBS and PNA (50 μg ml–1). Video taken 24 h after seeding. Scale bar, 50 μm.
Supplementary Video 8
In vivo LMC formation. Event 1. A sand fly midgut was dissected approximately 30 min after imbibing a second uninfected blood meal containing IgM (500 μg ml–1). Intact red blood cells are visible. Clumping formation (arrows) follows a pattern similar to that observed in vitro (Supplementary Videos 1 and 2). Scale bar, 50 μm.
Supplementary Video 9
In vivo LMC formation. Event 2. A sand fly midgut was dissected approximately 30 min after imbibing a second uninfected blood meal containing IgM (500 μg ml–1). Intact red blood cells are visible. Clumping formation (arrows) follows a pattern similar to that observed in vitro (Supplementary Videos 1 and 2). Scale bar, 50 μm.
Supplementary Video 10
In vivo LMC formation. Event 1. A sand fly midgut was dissected 24 h after imbibing a second uninfected blood meal containing IgM (500 μg ml–1). Partially digested red blood cells are visible. Fully formed LMCs can be seen at the centre of the video. Scale bar, 50 μm.
Supplementary Video 11
In vivo LMC formation. Event 2. A sand fly midgut was dissected 24 h after imbibing a second uninfected blood meal containing IgM (500 μg ml–1). Partially digested red blood cells are visible. Fully formed LMCs can be seen at the centre of the video (arrows). As shown for in vitro LMC formation (Supplementary Videos 1–3), this video captures the fusion of smaller clumps into one large clump. Scale bar, 50 μm.
Supplementary Video 12
In vivo LMC formation. Event 3. A sand fly midgut was dissected 24 h after imbibing a second uninfected blood meal containing IgM (500 μg ml–1). Partially digested red blood cells are visible. Fully formed LMCs can be seen at the centre of the video (arrows). As showed for in vitro LMC formation (Supplementary Videos 1–3), the fusion of smaller clumps into a bigger one can be observed. Scale bar, 50 μm.
Rights and permissions
About this article
Cite this article
Serafim, T.D., Iniguez, E., Barletta, A.B.F. et al. Leishmania genetic exchange is mediated by IgM natural antibodies. Nature 623, 149–156 (2023). https://doi.org/10.1038/s41586-023-06655-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06655-8
This article is cited by
-
Host IgM facilitates mating in Leishmania
Nature Reviews Immunology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.