Abstract
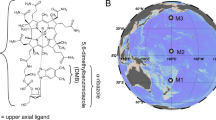
Cobalamin (vitamin B12, herein referred to as B12) is an essential cofactor for most marine prokaryotes and eukaryotes1,2. Synthesized by a limited number of prokaryotes, its scarcity affects microbial interactions and community dynamics2,3,4. Here we show that two bacterial B12 auxotrophs can salvage different B12 building blocks and cooperate to synthesize B12. A Colwellia sp. synthesizes and releases the activated lower ligand α-ribazole, which is used by another B12 auxotroph, a Roseovarius sp., to produce the corrin ring and synthesize B12. Release of B12 by Roseovarius sp. happens only in co-culture with Colwellia sp. and only coincidently with the induction of a prophage encoded in Roseovarius sp. Subsequent growth of Colwellia sp. in these conditions may be due to the provision of B12 by lysed cells of Roseovarius sp. Further evidence is required to support a causative role for prophage induction in the release of B12. These complex microbial interactions of ligand cross-feeding and joint B12 biosynthesis seem to be widespread in marine pelagic ecosystems. In the western and northern tropical Atlantic Ocean, bacteria predicted to be capable of salvaging cobinamide and synthesizing only the activated lower ligand outnumber B12 producers. These findings add new players to our understanding of B12 supply to auxotrophic microorganisms in the ocean and possibly in other ecosystems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Complete genome sequences of Colwellia sp. M166 and Roseovarius sp. M141 were uploaded into the NCBI and annotated by JGI Gold. Genomes can be inquired by their IMG genome identifiers 2828890045 for Colwellia and 2828508468 for Roseovarius. Transcriptomic sequences generated in this study have been deposited into the ENA at EMBL-EBI with the accession number PRJEB43320, using the data brokerage service of the GFBio in compliance with the Minimal Information about any (X) Sequence (MIxS) standard. The data from the sampling stations (ANT-XXVIII/4 and ANT-XXVIII/5) are available from PANGEA under the accession number PANGAEA.906247, and respective sequence data can be retrieved from the ENA under the INSDC accession number PRJEB34453. The SILVA and rfam database was used for transcriptome analysis. For bacterial gene annotation, the ProGenome database was used, and for phage gene and protein annotation, the NR database (from the NCBI), the Virus Orthologous Group database, the InterPro database and the HMM database were used. Source data are provided with this paper.
Code availability
No specialized in-house code was used for this study. All software used for the data analyses in this study are publicly available and cited in the Methods. Custom scripts and the pipeline used have been deposited into GitHub (https://github.com/LeonDlugosch/MetaSeq-Toolkit).
References
Croft, M. T., Lawrence, A. D., Raux-Deery, E., Warren, M. J. & Smith, A. G. Algae acquire vitamin B12 through a symbiotic relationship with bacteria. Nature 438, 90–93 (2005).
Sañudo-Wilhelmy, S. A., Gómez-Consarnau, L., Suffridge, C. & Webb, E. A. The role of B vitamins in marine biogeochemistry. Annu. Rev. Mar. Sci. 6, 339–367 (2014).
Shelton, A. N. et al. Uneven distribution of cobamide biosynthesis and dependence in bacteria predicted by comparative genomics. ISME J. 13, 789–804 (2019).
Lu, X., Heal, K. R., Ingalls, A. E., Doxey, A. C. & Neufeld, J. D. Metagenomic and chemical characterization of soil cobalamin production. ISME J. 14, 53–66 (2020).
Menzel, D. W. & Spaeth, J. P. Occurrence of vitamin B12 in the Sargasso Sea. Limnol. Oceanogr. 7, 151–154 (1962).
Suffridge, C. P. et al. B vitamins and their congeners as potential drivers of microbial community composition in an oligotrophic marine ecosystem. J. Geophys. Res. Biogeosci. 123, 2890–2907 (2018).
Cruz-López, R. & Maske, H. The vitamin B1 and B12 required by the marine dinoflagellate Lingulodinium polyedrum can be provided by its associated bacterial community in culture. Front. Microbiol. 7, 560 (2016).
Grant, M. A., Kazamia, E., Cicuta, P. & Smith, A. G. Direct exchange of vitamin B12 is demonstrated by modelling the growth dynamics of algal–bacterial cocultures. ISME J. 8, 1418–1427 (2014).
Cooper, M. B. et al. Cross-exchange of B-vitamins underpins a mutualistic interaction between Ostreococcus tauri and Dinoroseobacter shibae. ISME J. 13, 334–345 (2019).
Sokolovskaya, O. M., Shelton, A. N. & Taga, M. E. Sharing vitamins: cobamides unveil microbial interactions. Science 369, eaba0165 (2020).
Wienhausen, G. et al. Availability of vitamin B12 and its lower ligand intermediate α-ribazole impact prokaryotic and protist communities in oceanic systems. ISME J. 16, 2002–2014 (2022).
Sultana, S., Bruns, S., Wilkes, H., Simon, M. & Wienhausen, G. Vitamin B12 is not shared by all marine prototrophic bacteria with their environment. ISME J. 17, 836–845 (2023).
Stupperich, E., Steiner, I. & Eisinger, H. J. Substitution of co alpha-(5-hydroxybenzimidazolyl)cobamide (factor III) by vitamin B12 in Methanobacterium thermoautotrophicum. J. Bacteriol. 169, 3076–3081 (1987).
Crofts, T. S., Seth, E. C., Hazra, A. B. & Taga, M. E. Cobamide structure depends on both lower ligand availability and CobT substrate specificity. Chem. Biol. 20, 1265–1274 (2013).
Yi, S. et al. Versatility in corrinoid salvaging and remodeling pathways supports corrinoid-dependent metabolism in Dehalococcoides mccartyi. Appl. Environ. Microbiol. 78, 7745–7752 (2012).
Banerjee, R. & Ragsdale, S. W. The many faces of vitamin B12: catalysis by cobalamin-dependent enzymes. Annu. Rev. Biochem. 72, 209–247 (2003).
Hazra, A. B., Tran, J. L. A., Crofts, T. S. & Taga, M. E. Analysis of substrate specificity in CobT homologs reveals widespread preference for DMB, the lower axial ligand of vitamin B12. Chem. Biol. 20, 1275–1285 (2013).
Helliwell, K. E. et al. Cyanobacteria and eukaryotic algae use different chemical variants of vitamin B12. Curr. Biol. 26, 999–1008 (2016).
Heal, K. R. et al. Two distinct pools of B12 analogs reveal community interdependencies in the ocean. Proc. Natl Acad. Sci. USA 114, 364–369 (2017).
Gray, M. J. & Escalante-Semerena, J. C. A new pathway for the synthesis of α-ribazole-phosphate in Listeria innocua. Mol. Microbiol. 77, 1429–1438 (2010).
Gray, M. J. & Escalante-Semerena, J. C. The cobinamide amidohydrolase (cobyric acid-forming) CbiZ enzyme: a critical activity of the cobamide remodeling system of Rhodobacter sphaeroides. Mol. Microbiol. 74, 1198–1210 (2009).
Wienhausen, G., Noriega-Ortega, B. E., Niggemann, J., Dittmar, T. & Simon, M. The exometabolome of two model strains of the Roseobacter group: a marketplace of microbial metabolites. Front. Microbiol. 8, 1985 (2017).
Johnson, W. M., Kido Soule, M. C. & Kujawinski, E. B. Evidence for quorum sensing and differential metabolite production by a marine bacterium in response to DMSP. ISME J. 10, 2304–2316 (2016).
Crofts, T. S., Men, Y., Alvarez-Cohen, L. & Taga, M. E. A bioassay for the detection of benzimidazoles reveals their presence in a range of environmental samples. Front. Microbiol. 5, 592 (2014).
Butzin, N. C., Secinaro, M. A., Swithers, K. S., Gogarten, J. P. & Noll, K. M. Thermotoga lettingae can salvage cobinamide to synthesize vitamin B12. Appl. Environ. Microbiol. 79, 7006–7012 (2013).
Woodson, J. D., Zayas, C. L. & Escalante-Semerena, J. C. A new pathway for salvaging the coenzyme B12 precursor cobinamide in archaea requires cobinamide-phosphate synthase (CbiB) enzyme activity. J. Bacteriol. 185, 7193–7201 (2003).
Gray, M. J. & Escalante-Semerena, J. C. Single-enzyme conversion of FMNH2 to 5,6-dimethylbenzimidazole, the lower ligand of B12. Proc. Natl Acad. Sci. USA 104, 2921–2926 (2007).
Campbell, G. R. O. et al. Sinorhizobium meliloti bluB is necessary for production of 5,6-dimethylbenzimidazole, the lower ligand of B12. Proc. Natl Acad. Sci. USA 103, 4634–4639 (2006).
Yu, T.-Y. et al. Active site residues critical for flavin binding and 5,6-dimethylbenzimidazole biosynthesis in the flavin destructase enzyme BluB. Protein Sci. 21, 839–849 (2012).
Suh, S.-J. & Escalante-Semerena, J. C. Cloning, sequencing and overexpression of cobA which encodes ATP:corrinoid adenosyltranferase in Salmonella typhimurium. Gene 129, 93–97 (1993).
Escalante-Semerena, J. C., Suh, S. J. & Roth, J. R. cobA function is required for both de novo cobalamin biosynthesis and assimilation of exogenous corrinoids in Salmonella typhimurium. J. Bacteriol. 172, 273–280 (1990).
Mireku, S. A. et al. Conformational change of a tryptophan residue in BtuF facilitates binding and transport of cobinamide by the vitamin B12 transporter BtuCD-F. Sci Rep. 7, 41575 (2017).
Carini, P. et al. Discovery of a SAR11 growth requirement for thiamin’s pyrimidine precursor and its distribution in the Sargasso Sea. ISME J. 8, 1727–1738 (2014).
Wienhausen, G. et al. The overlooked role of a biotin precursor for marine bacteria—desthiobiotin as an escape route for biotin auxotrophy. ISME J. 16, 2599–2609 (2022).
Fang, H., Kang, J. & Zhang, D. Microbial production of vitamin B12: a review and future perspectives. Microb. Cell Fact. 16, 15 (2017).
Rodionov, D. A., Vitreschak, A. G., Mironov, A. A. & Gelfand, M. S. Comparative genomics of the vitamin B12 metabolism and regulation in prokaryotes. J. Biol. Chem. 278, 41148–41159 (2003).
Kadner, R. J. Repression of synthesis of the vitamin B12 receptor in Escherichia coli. J. Bacteriol. 136, 1050–1057 (1978).
Choi, J., Kotay, S. M. & Goel, R. Various physico-chemical stress factors cause prophage induction in Nitrosospira multiformis 25196—an ammonia oxidizing bacteria. Water Res. 44, 4550–4558 (2010).
Erez, Z. et al. Communication between viruses guides lysis–lysogeny decisions. Nature 541, 488–493 (2017).
Silpe, J. E. & Bassler, B. L. A host-produced quorum-sensing autoinducer controls a phage lysis–lysogeny decision. Cell 176, 268–280.e13 (2019).
Laganenka, L. et al. Quorum sensing and metabolic state of the host control lysogeny–lysis switch of bacteriophage T1. mBio 10, e01884–19 (2019).
Giovannoni, S. J. et al. Genome streamlining in a cosmopolitan oceanic bacterium. Science 309, 1242–1245 (2005).
Morris, J. J., Lenski, R. E. & Zinser, E. R. The Black Queen hypothesis: evolution of dependencies through adaptive gene loss. mBio 3, e00036–12 (2012).
Bertrand, E. M. et al. Vitamin B12 and iron colimitation of phytoplankton growth in the Ross Sea. Limnol. Oceanogr. 52, 1079–1093 (2007).
Bertrand, E. M. et al. Phytoplankton–bacterial interactions mediate micronutrient colimitation at the coastal Antarctic Sea ice edge. Proc. Natl Acad. Sci. USA 112, 9938–9943 (2015).
Tuttle, M. J. & Buchan, A. Lysogeny in the oceans: lessons from cultivated model systems and a reanalysis of its prevalence. Environ. Microbiol. 22, 4919–4933 (2020).
Gude, S., Pherribo, G. J. & Taga, M. E. Emergence of metabolite provisioning as a by-product of evolved biological functions. mSystems 5, e00259-20 (2020).
Suttle, C. A. Marine viruses—major players in the global ecosystem. Nat. Rev. Microbiol. 5, 801–812 (2007).
Alejandre-Colomo, C., Harder, J., Fuchs, B. M., Rosselló-Móra, R. & Amann, R. High-throughput cultivation of heterotrophic bacteria during a spring phytoplankton bloom in the North Sea. Syst. Appl. Microbiol. 43, 126066 (2020).
Zech, H. et al. Growth phase-dependent global protein and metabolite profiles of Phaeobacter gallaeciensis strain DSM 17395, a member of the marine Roseobacter clade. Proteomics 9, 3677–3697 (2009).
Osterholz, H., Niggemann, J., Giebel, H.-A., Simon, M. & Dittmar, T. Inefficient microbial production of refractory dissolved organic matter in the ocean. Nat. Commun. 6, 7422 (2015).
Lunau, M., Lemke, A., Dellwig, O. & Simon, M. Physical and biogeochemical controls of microaggregate dynamics in a tidally affected coastal ecosystem. Limnol. Oceanogr. 51, 847–859 (2006).
Bakenhus, I. et al. Composition of total and cell-proliferating bacterioplankton community in early summer in the North Sea—Roseobacters are the most active component. Front. Microbiol. 8, 1771 (2017).
Fuchs, B. M., Glöckner, F. O., Wulf, J. & Amann, R. Unlabeled helper oligonucleotides increase the in situ accessibility to 16S rRNA of fluorescently labeled oligonucleotide probes. Appl. Environ. Microbiol. 66, 3603–3607 (2000).
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Res. 32, 1363–1371 (2004).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.J 17, 10–12 (2011).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Vaser, R., Sović, I., Nagarajan, N. & Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27, 737–746 (2017).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Karst, S. M., Kirkegaard, R. H. & Albertsen, M. mmgenome: a toolbox for reproducible genome extraction from metagenomes. Preprint at bioRxiv https://doi.org/10.1101/059121 (2016).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2021).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Parks, D. H. et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat. Biotechnol. 36, 996–1004 (2018).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Methodol. 57, 289–300 (1995).
Markowitz, V. M. et al. IMG: the Integrated Microbial Genomes database and comparative analysis system. Nucleic Acids Res. 40, D115–122 (2012).
Raes, J., Korbel, J. O., Lercher, M. J., von Mering, C. & Bork, P. Prediction of effective genome size in metagenomic samples. Genome Biol. 8, R10 (2007).
Brown, C. T. et al. Unusual biology across a group comprising more than 15% of domain Bacteria. Nature 523, 208–211 (2015).
Dlugosch, L. et al. Significance of gene variants for the functional biogeography of the near-surface Atlantic Ocean microbiome. Nat. Commun. 13, 456 (2022).
Nurk, S., Meleshko, D., Korobeynikov, A. & Pevzner, P. A. metaSPAdes: a new versatile metagenomic assembler. Genome Res. 27, 824–834 (2017).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Menzel, P., Ng, K. L. & Krogh, A. Fast and sensitive taxonomic classification for metagenomics with Kaiju. Nat. Commun. 7, 11257 (2016).
Mende, D. R. et al. proGenomes: a resource for consistent functional and taxonomic annotations of prokaryotic genomes. Nucleic Acids Res. 45, D529–D534 (2017).
Brussaard, C. P., Payet, J. P., Winter, C. & Weinbauer, M. G. Quantification of aquatic viruses by flow cytometry. MAVE 11, 102–109 (2010).
Noguchi, H., Taniguchi, T. & Itoh, T. MetaGeneAnnotator: detecting species-specific patterns of ribosomal binding site for precise gene prediction in anonymous prokaryotic and phage genomes. DNA Res. 15, 387–396 (2008).
Charif, D. & Lobry, J. R. in Structural Approaches to Sequence Evolution. Biological and Medical Physics, Biomedical Engineering (eds Bastolla, U. et al.) 207–232 (Springer, 2007).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Grazziotin, A. L., Koonin, E. V. & Kristensen, D. M. Prokaryotic virus orthologous groups (pVOGs): a resource for comparative genomics and protein family annotation. Nucleic Acids Res. 45, D491–D498 (2017).
Steinegger, M. et al. HH-suite3 for fast remote homology detection and deep protein annotation. BMC Bioinformatics 20, 473 (2019).
Kiening, M. et al. Conserved secondary structures in viral mRNAs. Viruses 11, 401 (2019).
Finn, R. D. et al. InterPro in 2017—beyond protein family and domain annotations. Nucleic Acids Res. 45, D190–D199 (2017).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Barrero‐Canosa, J., Moraru, C., Zeugner, L., Fuchs, B. M. & Amann, R. Direct-geneFISH: a simplified protocol for the simultaneous detection and quantification of genes and rRNA in microorganisms. Environ. Microbiol. 19, 70–82 (2017).
Moraru, C. Gene-PROBER—a tool to design polynucleotide probes for targeting microbial genes. Syst. Appl. Microbiol. 44, 126173 (2021).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Barrero-Canosa, J. & Moraru, C. Linking microbes to their genes at single cell level with direct-geneFISH. Methods Mol. Biol. 2246, 169–205 (2021).
Lunau, M., Lemke, A., Walther, K., Martens-Habbena, W. & Simon, M. An improved method for counting bacteria from sediments and turbid environments by epifluorescence microscopy. Environ. Microbiol. 7, 961–968 (2005).
Bruns, S., Wienhausen, G., Scholz-Böttcher, B., Heyen, S. & Wilkes, H. Method development and quantification of all B vitamins and selected biosynthetic precursors in winter and spring samples from the North Sea and de novo synthesized by Vibrio campbellii. Mar. Chem. 256, 104300 (2023).
Bruns, S., Wienhausen, G., Scholz-Böttcher, B. & Wilkes, H. Simultaneous quantification of all B vitamins and selected biosynthetic precursors in seawater and bacteria by means of different mass spectrometric approaches. Anal. Bioanal. Chem. 414, 7839–7854 (2022).
Mukherjee, S. et al. Genomes OnLine Database (GOLD) v.8: overview and updates. Nucleic Acids Res. 49, D723–D733 (2021).
Silvester, N. et al. The European Nucleotide Archive in 2017. Nucleic Acids Res. 46, D36–D40 (2018).
Diepenbroek, M. et al. Towards an Integrated Biodiversity and Ecological Research Data Management and Archiving Platform: The German Federation for the Curation of Biological Data (GFBio) (Gesellschaft für Informatik, 2014).
Yilmaz, P. et al. Minimum information about a marker gene sequence (MIMARKS) and minimum information about any (x) sequence (MIxS) specifications. Nat. Biotechnol. 29, 415–420 (2011).
Acknowledgements
We are grateful to C. Alejandre-Colomo, R. Mosseló-Móra, R. Amann and V. Bischoff for providing the North Sea collection of bacterial isolates; B. Kuerzel and M. Wolterink for technical assistance in the HPLC analysis of amino acids and enumerating VLPs; the captains, their crews and the scientific groups of the cruises ANT XXVIII/4 and XXVIII/5 of RV Polarstern and RV Senckenberg for their support; S. Sañudo-Wilhelmy for stimulating discussions regarding B12 analyses; and D. Kirchman for carefully revising a previous version of this publication This work was supported by Deutsche Forschungsgemeinschaft within the Transregional Collaborative Research Center Roseobacter (TRR51) and the Gordon and Betty Moore Foundation (to F.A.).
Author information
Authors and Affiliations
Contributions
G.W. designed the study, carried out the experiments, the genome analyses for B12 biosynthetic pathways, the transcriptomic analyses and wrote the manuscript. C.M. and G.W. carried out the direct-geneFISH analyses for prophage induction and the prophage-related genomic analyses. D.Q.T. contributed to the CARD-FISH analyses. S.B. and H.W. analysed B12. S.S. assisted in the tripartite consortium experiment. L.D. carried out the transcriptome mapping and mapping of taxa with different B12 genetic traits to the Atlantic Ocean microbiome. F.A. partially supervised the co-culture experiments. M.S. assisted in designing the study, partially supervised the experiments and data analyses and wrote the manuscript, together with G.W. All authors revised the manuscript to finalize it.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
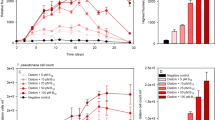
Extended Data Fig. 1 Growth characteristics of Colwellia and Roseovarius when supplemented with and without B12 and respective building blocks and methionine and Chaetoceros muelleri in tri-culture with Colwellia and Roseovarius.
a, Maximum growth yield as means of triplicates ± SD as total cell numbers per ml assessed by flow cytometry of Roseovarius; b, of Colwellia when supplemented with B12 (at 1, 10 or 100 pM) or DMB / Cbi (at 1, 10 or 100 pM); Maximum growth yield as means of triplicates ± SD as determined by optical density of c, Roseovarius and d, Colwellia. Each isolate was supplemented with B12 (100 pM), no addition (plain bar) and methionine (1 µM); e, Chlorophyll a fluorescence and total cell numbers enumerated by hemocytometer of axenic Chaetoceros muelleri over time when cultivated without any addition, f, supplemented with B12 at 10 pM and g, 100 pM final concentration and h, when grown in tri-culture with Colwellia or Roseovarius. Experiments a-d were conducted in n = 3 and experiments e-h in n = 4 biological independent samples each in 1 independent experiment.
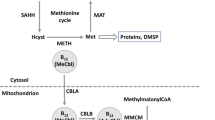
Extended Data Fig. 2 Metabolic pathways related to cobalamin biosynthesis and transcript expression patterns of Colwellia and Roseovarius grown in co-culture.
Differential gene expression between the control, represented by the mono-cultures (Roseovarius & Colwellia) with B12 added, and the co-culture, without B12 addition, was quantified with the DESeq2 package as implemented in R v4.0.5 (R-Core-Team, 2022). In each treatment n = 3 biological independent samples were used. Coloring of genes refers to their LOG-2FC regulation, defined as downregulated (light red to red), upregulated (light green to dark green) and similarly regulated (orange). Genes that are significantly regulated (LOG2-FC larger than 2 and smaller than −2 with an BH adj. p-value < 0.05) are marked with an asterisk. When an illustration represents more than one gene (all genes > (-) 2 LOG-2FC and adj. p-value < 0.05), the asterisk is placed in brackets. Genes that were not identified in the genome (grey). Since the bluB gene in Roseovarius is missing a large section of the sequence, we have indicated this by outlining the gene in gray. Corresponding gene expression significances (adjusted p-value) and further details on genes are presented in Supplementary Data 2. Our documented microbial interactions including the coincidence of prophage induction, host cell lysis and release of B12, whose causative relationships still need to be proven, are illustrated and explained by the dashed arrows and the explanations 1–6 in the legend.
Extended Data Fig. 3 Growth and substrate use of Colwellia and Roseovarius when supplemented with B12 building blocks.
Presented are total bacterial cell numbers assessed by flow cytometry and glutamate (glutamic acid) concentrations of a-d, Colwellia and e-h, Roseovarius supplemented with a, e, B12, b, f, DMB and c, g, Cbi at 1 nM final concentrations each and d, h, without any supplementation and of as well as i, Colwellia + Roseovarius when cultivated in co-culture without any addition. Shown are means of triplicates ± SD. Each condition was carried out in n = 3 biological independent samples and growth characteristics were confirmed in independently conducted experiments.
Extended Data Fig. 4 Growth characteristics of Thalassiosira pseudonana and Colwellia and Roseovarius cultivated in consortium.
a, Growth of Colwellia and Roseovarius as monitored by flow cytometric cell counts and calculated by CARD-FISH analyses. Further, numbers of virus-like particles measured by flow cytometry are presented. b, cell numbers of T. pseudonana, assessed microscopically using a hemocytometer, and bacterial cells (black circle) assessed by flow cytometry. c, Total bacterial cell abundance and the ratio of virus-like particle counts to the number of Colwellia. d, Total bacterial cell abundance and the ratio of virus-like particle counts to the number of Roseovarius. e, Relative proportions of Colwellia and Roseovarius cells (% of total abundance) during the culturing time. f. Descriptive illustration of the growth progression of the participating partners of the consortium, T. pseudonana, Roseovarius and Colwellia as well as the phage induction over a period of ten weeks. g, Maximum relative fluorescence of T. pseudonana when cultured with Colwellia and Roseovarius, with addition of 100 pM B12, without any addition, in co-culture with Roseovarius and in co-culture with Colwellia. a-f, Means of triplicates g, and ± SD. Measured values of the respective triplicates can be viewed in the corresponding source data file. Each treatment was carried out in n = 3 biological independent samples and growth characteristics were confirmed in independently conducted tests.
Extended Data Fig. 5 Lower ligand building block concentration in the exudate of Colwellia at B12 (20 pM) depleted growth conditions and in North Sea seawater samples.
a, means of triplicates + SD of growth of Colwellia supplemented with 1 nM B12, 20 pM B12 and no addition, monitored by flow cytometric cell counts over time. Extracellular concentrations of α-ribazole and 5,6-dimethylbenzimidazole (DMB) were measured for the growth-limited culture when supplemented with 20 pM B12. b, concentrations of B12, cobinamide, DMB and α -ribazole in North Sea seawater samples as means of triplicates + SD. c, Map of the German Bight of the North Sea. Points mark the sampling stations of the analyzed seawater samples. Experiment a was conducted in n = 3 biological independent samples. Experiment b was conducted on n = 3 independent sampling locations and each sample was detected in n = 3 technical replicates. Experiments a-b were conducted in 1 independent experiment. The map was created using Ocean Data View.
Extended Data Fig. 6 Predicted cobamide biosynthesis phenotypes in genomes of marine bacteria and their relative abundance in the Atlantic Ocean.
a, Map showing 22 stations (black dots) from 62°S in the Southern Ocean sector to 47°N in the north Atlantic Ocean sampled at 20 m depth during cruises ANTXXVIII/4 and /5 with RV Polarstern. The percentage given along each pie chart represents the proportion of genome-sequenced prokaryotes of the entire prokaryotic community (identified by 16S rRNA gene). The pie chart of every station sampled presents relative proportions of prokaryotes encoding i) the complete B12 biosynthesis pathway (B12 producers; red), ii) corrin ring biosynthesiser (dark red), iii) cobinamide salvagers (lower ligand producers) (pink) iv) prokaryotes identified as B12 non-producers (grey). Stations are overlayed on a map with annual mean concentrations of chlorophyll a at the surface (https://oceandata.sci.gsfc.nasa.gov). b-e, B12 biosynthesis genes of 1,904 genomes of marine bacteria were analyzed and classified into cobamide biosynthesis phenotypes: as described above for a. Presented are relative abundance of b, all analyzed bacteria, c, phyla and classes, d, orders of Alphaproteobacteria and e, orders of Gammaproteobacteria.
Extended Data Fig. 7 Proportions of genes encoding reactions of B12 biosynthesis in genomes of Rhodobacterales, Alteromonadales, Vibrionales and Betaproteobacteria.
Presence (as percentage) of individual B12 biosynthesis genes in Rhodobacterales (220), Alteromonadales (96), Vibrionales (52) and Betaproteobacteria (126) genomes. Percent of gene abundance is given in color-coded 10% increments from red (0%) to dark green (100%). Genes depicted in grey were not identified in examined genomes.
Extended Data Fig. 8 (Pro)-phage targeting direct-geneFISH signal intensities.
Shown are the mean relative intensities of the (pro)-phage targeting direct-geneFISH signals of Roseovarius in triplicate co-culture samples withdrawn after 96 h and 120 h and the mono-culture Roseovarius samples withdrawn after 96 h as boxplots. Individual, small dots represent the (pro)-phage targeting direct-geneFISH signal intensities of the individually measured regions of interest (ROI; approximate one cell). At least 465 ROIs were defined and validated for each sample. The red dashed line defines the threshold for the definition of induced cells. The percentage below the samples indicates the proportion of induced cells in the total cells analyzed. In each treatment n = 3 biological independent samples were run. The lower and upper hinges of the box correspond to the first and third quartiles (the 25th and 75th percentiles). The central line in the box is the median. The upper whisker extends from the hinge to the largest value no further than 1.5 * IQR from the hinge (where IQR is the inter-quartile range, or distance between the first and third quartiles). The lower whisker extends from the hinge to the smallest value at most 1.5 * IQR of the hinge. Data beyond the end of the whiskers are called “outlying” points and are plotted individually. Prophage induction events, indicated by a significant increase of the phage signal intensity in regions of interest (ROI), were observed in 2 independent experiments.
Extended Data Fig. 9 Enumeration of bacterial cells and virus-like particles of Colwellia and Roseovarius grown in co- and mono-culture.
a, Bacterial cell numbers (white triangle) and ratio of virus-like particles to bacterial cell number (black circle) of Colwellia and Roseovarius growing in co-culture, c, Roseovarius and e, Colwellia in mono-culture with B12 supplementation. b, Bacterial cell numbers (white triangle) and virus-like particles (black circle) of Colwellia and Roseovarius growing in co-culture, d, Roseovarius and f, Colwellia in mono-culture with B12 supplementation. Shown are means of triplicates + SD. Each treatment was carried out in n = 3 biological independent samples and growth characteristics were confirmed in independently conducted tests.
Supplementary information
Supplementary Figures
Supplementary Figs. 1–5.
Supplementary Table 1
Growth rate, growth yield and required molecules per cell of Colwellia and Roseovarius growing in mono-culture when supplemented with B12 or respective B12 building blocks.
Supplementary Table 2
Intracellular vitamin B12 recovery (Hydroxycobalamin, Cyanocobalamin, Adenosylcobalamin, Methylcobalamin) by LC–MS of Colwellia and Roseovarius growing in mono-culture when supplemented with B12 or respective B12 building blocks.
Supplementary Table 3
Growth rate and growth yield of C. muelleri growing in mono-culture when supplemented with B12 or in consortium with Colwellia and Roseovarius.
Supplementary Table 4
Composition of synASW medium and synASW medium trace elements solution, used for combined diatom–bacteria cultivation of C. muellieri or T. pseudonana with Roseovarius and Colwellia.
Supplementary Table 5
Settings used for selected reaction monitoring.
Supplementary Data 1
Identification of B12 biosynthesis genes, B12-dependent reactions, B12-indipendent reactions, B12 transporter genes and B12 salvage genes in the genomes of Roseovarius and Colwellia.
Supplementary Data 2
Transcriptional gene regulation of both bacterial isolates in co-culture compared with gene regulation when supplemented with the addition of B12 (1 nM) cultivated in mono-culture. Genes transcribed at a log2(FC) below –2 and above 2 are outlined in separate sheets, highlighting respective cellular functions.
Supplementary Data 3
Classification of B12 pathway synthesizing groups of publicly available aquatic bacterial genomes.
Supplementary Data 4
Gene annotation of the prophage Roseophage ICBM167.
Supplementary Data 5
Polynucleotide probe mixture for the detection of Roseophage ICBM167 by targeted direct-geneFISH.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wienhausen, G., Moraru, C., Bruns, S. et al. Ligand cross-feeding resolves bacterial vitamin B12 auxotrophies. Nature (2024). https://doi.org/10.1038/s41586-024-07396-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41586-024-07396-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.