Abstract
To systematically define molecular features in human tumor cells that determine their degree of sensitivity to human allogeneic natural killer (NK) cells, we quantified the NK cell responsiveness of hundreds of molecularly annotated ‘DNA-barcoded’ solid tumor cell lines in multiplexed format and applied genome-scale CRISPR-based gene-editing screens in several solid tumor cell lines, to functionally interrogate which genes in tumor cells regulate the response to NK cells. In these orthogonal studies, NK cell–sensitive tumor cells tend to exhibit ‘mesenchymal-like’ transcriptional programs; high transcriptional signature for chromatin remodeling complexes; high levels of B7-H6 (NCR3LG1); and low levels of HLA-E/antigen presentation genes. Importantly, transcriptional signatures of NK cell–sensitive tumor cells correlate with immune checkpoint inhibitor (ICI) resistance in clinical samples. This study provides a comprehensive map of mechanisms regulating tumor cell responses to NK cells, with implications for future biomarker-driven applications of NK cell immunotherapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
TCGA data were acquired from the Broad Firehose web site, v.2016_07_15. Read counts from the CRISPR and PRISM screens are available in Supplementary Table 4. Raw data can be available upon request. Molecular profiling data of tumor cells treated with broad-spectrum HDAC inhibitors were accessed from Gene Expression Omnibus (GEO: GSE108191, GSE96649, GSE101708 and GSE37376). RNA-seq data of tumors from patients treated with ICI were accessed from GEO: GSE115978 (Jerby-Arnon et al.25), GSE91061 (Riaz et al.26) and GSE78220 (Hugo et al.24). The Van-Allen dataset was acquired from dbGaP (accession phs000452.v2.p1 (ref. 28)). Source data are provided with this paper.
Code availability
The study did not develop new software of custom code, but applied in sequence different previously available codes for which details are provided in the Methods and Supplementary Information. All code used in this study can be available upon request.
References
Liu, E. et al. Use of CAR-transduced natural killer cells in CD19-positive lymphoid tumors. N. Engl. J. Med. 382, 545–553 (2020).
Guillerey, C., Huntington, N. D. & Smyth, M. J. Targeting natural killer cells in cancer immunotherapy. Nat. Immunol. 17, 1025–1036 (2016).
Nayyar, G., Chu, Y. & Cairo, M. S. Overcoming resistance to natural killer cell based immunotherapies for solid tumors. Front. Oncol. 9, 51 (2019).
Shimasaki, N., Jain, A. & Campana, D. NK cells for cancer immunotherapy. Nat. Rev. Drug Discov. 19, 200–218 (2020).
Gurjao, C. et al. Intrinsic resistance to immune checkpoint blockade in a mismatch repair-deficient colorectal cancer. Cancer Immunol. Res. 7, 1230–1236 (2019).
Sade-Feldman, M. et al. Resistance to checkpoint blockade therapy through inactivation of antigen presentation. Nat. Commun. 8, 1136 (2017).
Yu, C. et al. High-throughput identification of genotype-specific cancer vulnerabilities in mixtures of barcoded tumor cell lines. Nat. Biotechnol. 34, 419–423 (2016).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Ghandi, M. et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature 569, 503–508 (2019).
Brandt, C. S. et al. The B7 family member B7-H6 is a tumor cell ligand for the activating natural killer cell receptor NKp30 in humans. J. Exp. Med. 206, 1495–1503 (2009).
Kaifu, T., Escaliere, B., Gastinel, L. N., Vivier, E. & Baratin, M. B7-H6/NKp30 interaction: a mechanism of alerting NK cells against tumors. Cell. Mol. Life Sci. 68, 3531–3539 (2011).
Braud, V. M. et al. HLA-E binds to natural killer cell receptors CD94/NKG2A, B and C. Nature 391, 795–799 (1998).
Andre, P. et al. Anti-NKG2A mAb is a checkpoint inhibitor that promotes anti-tumor immunity by unleashing both T and NK cells. Cell 175, 1731–1743.e13 (2018).
Kamiya, T., Seow, S. V., Wong, D., Robinson, M. & Campana, D. Blocking expression of inhibitory receptor NKG2A overcomes tumor resistance to NK cells. J. Clin. Invest. 129, 2094–2106 (2019).
Zhou, F. Molecular mechanisms of IFN-γ to up-regulate MHC class I antigen processing and presentation. Int. Rev. Immunol. 28, 239–260 (2009).
Dendoncker, K. et al. The nature of the GRE influences the screening for GR-activity enhancing modulators. PLoS ONE 12, e0181101 (2017).
Rafehi, H. et al. Vascular histone deacetylation by pharmacological HDAC inhibition. Genome Res. 24, 1271–1284 (2014).
Sanchez, G. J. et al. Genome-wide dose-dependent inhibition of histone deacetylases studies reveal their roles in enhancer remodeling and suppression of oncogenic super-enhancers. Nucleic Acids Res. 46, 1756–1776 (2018).
Pech, M. F. et al. Systematic identification of cancer cell vulnerabilities to natural killer cell-mediated immune surveillance. eLife 8, e47362 (2019).
Tawbi, H. A. et al. Pembrolizumab in advanced soft-tissue sarcoma and bone sarcoma (SARC028): a multicentre, two-cohort, single-arm, open-label, phase 2 trial. Lancet Oncol. 18, 1493–1501 (2017).
Burgess, M. A. et al. Clinical activity of pembrolizumab (P) in undifferentiated pleomorphic sarcoma (UPS) and dedifferentiated/pleomorphic liposarcoma (LPS): final results of SARC028 expansion cohorts. J. Clin. Oncol. 37, 11015–11015 (2019).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Jerby-Arnon, L. et al. A cancer cell program promotes T cell exclusion and resistance to checkpoint blockade. Cell 175, 984–997.e24 (2018).
Riaz, N. et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171, 934–949.e16 (2017).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Schwartzkopff, S. et al. Tumor-associated E-cadherin mutations affect binding to the killer cell lectin-like receptor G1 in humans. J. Immunol. 179, 1022–1029 (2007).
Freeman, A. J. et al. Natural killer cells suppress T cell-associated tumor immune evasion. Cell Rep. 28, 2784–2794.e5 (2019).
Zhuang, X., Veltri, D. P. & Long, E. O. Genome-wide CRISPR screen reveals cancer cell resistance to NK cells induced by NK-derived IFN-γ. Front. Immunol. 10, 2879 (2019).
Leung, W. H. et al. Modulation of NKG2D ligand expression and metastasis in tumors by spironolactone via RXRγ activation. J. Exp. Med. 210, 2675–2692 (2013).
Mou, X. et al. The regulatory effect of UL-16 binding protein-3 expression on the cytotoxicity of NK cells in cancer patients. Sci. Rep. 4, 6138 (2014).
Xu, X., Rao, G. & Li, Y. Xanthine oxidoreductase is required for genotoxic stress-induced NKG2D ligand expression and gemcitabine-mediated antitumor activity. Oncotarget 7, 59220–59235 (2016).
Ferrari de Andrade, L. et al. Antibody-mediated inhibition of MICA and MICB shedding promotes NK cell-driven tumor immunity. Science 359, 1537–1542 (2018).
Woods, D. M. et al. HDAC inhibition upregulates PD-1 ligands in melanoma and augments immunotherapy with PD-1 blockade. Cancer Immunol. Res. 3, 1375–1385 (2015).
Zhu, S. et al. The narrow-spectrum HDAC inhibitor entinostat enhances NKG2D expression without NK cell toxicity, leading to enhanced recognition of cancer cells. Pharm. Res. 32, 779–792 (2015).
McMillin, D. W. et al. Tumor cell-specific bioluminescence platform to identify stroma-induced changes to anticancer drug activity. Nat. Med. 16, 483–489 (2010).
McMillin, D. W. et al. Compartment-specific bioluminescence imaging platform for the high-throughput evaluation of antitumor immune function. Blood 119, e131–e138 (2012).
Tsafrir, D. et al. Sorting points into neighborhoods (SPIN): data analysis and visualization by ordering distance matrices. Bioinformatics 21, 2301–2308 (2005).
Shirasaki, R. et al. Functional genomics identify distinct and overlapping genes mediating resistance to different classes of heterobifunctional degraders of oncoproteins. Cell Rep. 34, 108532 (2021).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Lozupone, F. et al. Effect of human natural killer and γδ T cells on the growth of human autologous melanoma xenografts in SCID mice. Cancer Res. 64, 378–385 (2004).
Bobisse, S. et al. Reprogramming T lymphocytes for melanoma adoptive immunotherapy by T-cell receptor gene transfer with lentiviral vectors. Cancer Res. 69, 9385–9394 (2009).
Eisenberg, V. et al. Targeting multiple tumors using T-cells engineered to express a natural cytotoxicity receptor 2-based chimeric receptor. Front. Immunol. 8, 1212 (2017).
Gunnarsdottir, F. B., Kiessling, R. & Pico de Coana, Y. Establishment of melanoma tumor xenograft using single cell line suspension and co-injection of patient-derived T cells in immune-deficient NSG mice. Methods Mol. Biol. 1913, 207–215 (2019).
Jakka, G. et al. Antigen-specific in vitro expansion of functional redirected NY-ESO-1-specific human CD8+ T-cells in a cell-free system. Anticancer Res. 33, 4189–4201 (2013).
Acknowledgements
This work was supported by the Stand Up To Cancer (SU2C) Convergence 2.0 Grant (M.S., A.C.C., C.S.M.), a collaboration by SU2C and the Society for Immunotherapy of Cancer; the SU2C Phillip A. Sharp Award for Innovation in Collaboration (grant number: SU2C-AACR-PS-22) (M.S., C.S.M.); the Claudia Adams Barr Program in Innovative Basic Cancer Research at Dana–Farber Cancer Institute (M.S., E.D., C.S.M., E.L.); the Human Frontier Science Program (HFSP) Fellowship (LT000834/2014-L, M.S.); the Leukemia and Lymphoma Society (LLS) Scholar Award (C.S.M.); and the Ludwig Center at Harvard (C.S.M.). The collaborating laboratories have also been partially supported by grants NIH R01 CA050947 (C.S.M.), CA196664 (C.S.M.) and U01 CA225730 (C.S.M.); de Gunzburg Myeloma Research Fund (C.S.M.); International Myeloma Society (C.S.M.), the Shawna Ashlee Corman Investigatorship in Multiple Myeloma Research (C.S.M.); Cobb Family Myeloma Research Fund (C.S.M.); the Department of Defense grants W81XWH-15-1-0012 (C.S.M.) and W81XWH-15-1-0013 (A.C.); the Finnish Cancer Organizations, Sigrid Juselius Foundation, Academy of Finland (S.M.); the Health Foundation Limburg, Nijbakker-Morra Stichting (N.B.); and the Dutch Cancer Association (KWF Kanker Bestrijding, UM2012-5375) (L. Wieten). We thank M. Sade-Feldman for technical advice and discussion, A. Rotem for the gift of Malme-3M cell line and Y. Maruvka for assistance with MSI profiling.
Author information
Authors and Affiliations
Contributions
M.S. and C.S.M. conceived and designed the study overall. M.S., E.L., N.B., S.N.-A.A, M.B., C.C.M, J.A.R., O. Dashevsky, R.D.M.S., E.D., C.Y., Y.H., S.G., T.G., C.Z., L. Wang, L. Wieten and C.S.M. contributed to design of individual experiments. C.C.M., J.A.R., C.Y. and T.G. provided critical reagents. M.S., E.L., N.B., S.N.-A.A., M.B., O. Dashevsky, S.B., C.Z. and L. Wang performed experiments. M.S., E.L., N.B., J.G.B., S.S.F., A.T., R.D.M.S. and A.C.C. performed data analyses. M.S., J.A.R., A.T., J.G.B., O. Dufva, M.G., I.T., G.M.K., V.S., E.F., T.G., R.R., S.M., A.C.C., L. Wieten and C.S.M. contributed to data interpretation. M.S. and C.S.M. wrote the manuscript with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
C.C.M., J.A.R., C.Y., T.G., L. Wieten, M.S. and C.S.M. are authors of a patent application related to antitumor activity of NK cells. C.S.M. also discloses consultant/honoraria from Fate Therapeutics, Ionis Pharmaceuticals and FIMECS; employment of a relative with Takeda; and research funding from Janssen/Johnson & Johnson, TEVA, EMD Serono, Abbvie, Arch Oncology, Karyopharm, Sanofi, Nurix and H3 Biomedicine. M.G. receives research funding from Bristol-Myers Squibb and Merck. A.T. is a consultant for Tango Therapeutics. S.M. has received honoraria and research funding from Novartis, Pfizer and Bristol-Myers Squibb (not related to this study). The remaining authors declare no competing interests.
Additional information
Peer review information Nature Genetics thanks Joseph Sun, Sidi Chen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Genes associated with tumour cell responses to NK cells in PRISM studies.
a Scatter plots for AUC derived from PRISM study per time point (x-axis) and from cytotoxic assays of individual cell lines (y-axis). The Spearman correlation coefficients are specified for each time point. b Volcano plot for the -log10(p-value) of the Spearman correlation between AUC and gene expression (y-axis) vs log2 fold changes of expression for the respective genes between NK cell-resistant (upper third of AUC distribution) and NK cell-sensitive (lower third of AUC distribution) cell lines (see Methods). Results are shown for 24hr-AUC as representative of results for other time points. Genes with sgRNA enrichment are labeled red and sgRNA depletion are labeled blue. Genes in bold were significant based on MaGECK and STARS analyses in at least one CRISPR screen.
Extended Data Fig. 2 Key regulators of tumor cell responses to NK cells in CRISPR studies.
Plots for sgRNA enrichment (upper panels) or depletion (lower panels) for each one of the genome-scale CRISPR gene-editing screens. Highlighted genes were identified as significant based on MaGECK and STARS in at least one screen.
Extended Data Fig. 3 Stratified PRISM-based analyses for global (all cell lines), epithelial-like or mesenchymal-like groups of lines.
Normalized transcript levels for genes which have significant correlation (Spearman correlation coefficient, adjusted p-values<0.05, see Methods) with AUC values in ≥2 time points for the entire set of PRISM cell lines. A gene was considered to correlate with AUC in the epithelial-like or the mesenchymal-like subset of lines if p-value ≤0.05 in ≥2 time points in the respective subgroup analysis. Horizontal black lines represent genes which exhibited significant sgRNA enrichment or depletion in at least one of the CRISPR studies (based on MAGECK and STARS) and in a manner concordant with the results of the PRISM analyses. Group A was enriched for these genes, compared with groups B-D (p = 0.022, χ2 test, df = 1).
Extended Data Fig. 4 Integration of results for PRISM and CRISPR-based identification of regulators of tumor cell response to NK cells.
Functional clusters of genes that were associated with tumour cell responses to NK cells (based on genes that were significant for both MaGECK and STARS in at least one screen). Gene highlighted in bold were also significant for PRISM.
Extended Data Fig. 5 B7-H6 and HLA-E as key regulators of NK cell responses.
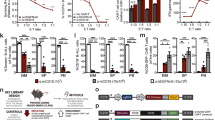
a, In vivo testing of NK cell activity against SW620 tumour cells with vs. without B7-H6 CRISPR knockout: Bioluminescence intensity (photons/sec/cm2) measured at week 2. P-values are calculated using two- sided Mann-Whitney tests between B7-H6– vs. B7-H6+ tumours in each group. Data are represented as mean ± SD, n = 10 per group. b, B7-H6 log2-fold changes of tumour vs. normal samples in RNA-seq data of the TCGA dataset. Data are represented as box/dot plots, demarcating the first and third quartiles of the distribution, with the median shown in the center and whiskers covering data within 1.5x the interquartile range of the box (significant FDR-adjusted p-values for upregulation are shown in red; two-sided Mann-Whitney tests). c, Heatmap for z-scores of transcript levels for antigen presentation genes, showing co-expression in the different cancer types of the TCGA. The gene expression levels were normalized across the samples of each cancer type. d, Western blot analyses for STAT1 protein levels in STAT1 knockouts vs. OR10A2 as control knockouts in HT29 and SW620 cell lines. Staining for GAPDH on the same uncropped membrane serves as loading control. n = 2 distinct experiments. e, Flow cytometry analysis for surface HLA-E levels in HT29 cells with sgRNA for HLA-E vs. control sgRNA with vs. without IFNγ (5ng/ml) overnight incubation, n = 2 experiments.
Extended Data Fig. 6 Functional relationship of B7-H6 and MHC class I molecules.
a, Comparison of transcript levels between Wild-Type (WT) and B2M-mutated cell lines of the PRISM collection (coding mutations), Mann-Whitney two-sided test. b, Spearman correlation coefficients between B7-H6 and B2M (upper panel) or HLA-E (lower panel) in TCGA and CCLE datasets. c, Heatmap of the normalized expression levels of B7-H6 and HLA-E in PRISM cell lines sorted based on difference (normalized log2 fold) of B7-H6 compared with HLA-E. d, Percentages of MSI cell lines (brown) vs. MSS cell lines (orange) in the group of HLA-low (n = 35) cell lines compared with all other lines (n = 348) of the PRISM-based surface proteomic analysis. e, HLA-A/B/C surface protein scores or B2M transcript levels between the groups of MSS vs. MSI cell lines; two-sided Mann-Whitney test. f, Left: AUC for MSI lines with low-HLA-A/B/C (n = 13) vs. all other lines (n = 370) of the PRISM-based surface proteomic analysis. Right: AUC for MSI lines with low B2M expression (n = 37) vs. all other lines (n = 490); two-sided Mann-Whitney test. Results are shown for 24hr-AUC and are representative of those for other time points. g, HLA-E and B2M log2-fold changes of tumour vs. normal samples in the RNA-seq data of the TCGA dataset (significant FDR-adjusted p-values for downregulation are shown in red; two-sided Mann-Whitney). Data in a,e,f,g, are represented as box/dot plots, demarcating the first and third quartiles of the distribution, with the median shown in the center and whiskers covering data within 1.5x the interquartile range of the box.
Extended Data Fig. 7 Transcriptional signatures in tumor cells associated with responses to NK cells.
a, Left: GSEA NES scores for selected gene sets that had FDR adjusted p<0.05 in at least two time points in analyses that included all PRISM cell lines: NES are also shown for GSEA of only epithelial-like and only mesenchymal-like lines (see Methods). Right: GSEA NES scores for same gene sets as in left panel in tumors with ICI resistance (see Methods). Red and blue represent enrichment and suppression, respectively, of gene sets in the corresponding analyses. Lack of significant changes (p>0.05) was colored white. b,c, Normalized expression levels of core genes from pathways correlating with AUC (see Methods) in b, epithelial-like or mesenchymal-like cell lines and c, TCGA tumours, sorted by the expression of CDH1 and ZEB2 using SPIN40. Tumours with low vs. high CDH1 are separated with a dotted line. d, SW620 cells were pre-treated for 12 hrs with 50nM of the HDAC inhibitor panobinostat, 5 ng/ml IFNγ or DMSO. The cells were stained with anti-HLA-E (left) and anti-B7-H6 (right) antibodies. Unlabeled controls are colored grey. n = 1 experiments. e, Flow cytometry assays to quantify NK cell cytotoxicity against tumour cells (see Methods) pre-treated with HDAC inhibitor or DMSO. Cells were treated with 12.5nM panobinostat and cocultured with NK cells in two E:T ratios. n = 4 distinct experiments. f, GSEA enrichment plots for select gene sets from a, in all cell lines of PRISM (upper) and in ICI nonresponders vs. responders of the Riaz et al. dataset (lower). g, Gene scores for NK cell sensitivity in ICI nonresponders vs. responders of the Riaz et al. dataset (see Methods, two-sided Mann-Whitney test).
Supplementary information
Supplementary Information
Supplementary Note and Tables 2, 6 and 7
Supplementary Tables
Supplementary Tables 1 and 3–5
Source data
Source Data Extended Data Fig. 5
Unprocessed western blot.
Rights and permissions
About this article
Cite this article
Sheffer, M., Lowry, E., Beelen, N. et al. Genome-scale screens identify factors regulating tumor cell responses to natural killer cells. Nat Genet 53, 1196–1206 (2021). https://doi.org/10.1038/s41588-021-00889-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-00889-w
This article is cited by
-
Epigenetic regulation of CD38/CD48 by KDM6A mediates NK cell response in multiple myeloma
Nature Communications (2024)
-
Clonal dynamics limits detection of selection in tumour xenograft CRISPR/Cas9 screens
Cancer Gene Therapy (2023)
-
The NK cell checkpoint NKG2A maintains expansion capacity of human NK cells
Scientific Reports (2023)
-
Monalizumab efficacy correlates with HLA-E surface expression and NK cell activity in head and neck squamous carcinoma cell lines
Journal of Cancer Research and Clinical Oncology (2023)
-
Intensified NK cell therapy in combination with low-dose chemoradiotherapy against human colorectal cancer
Cancer Immunology, Immunotherapy (2023)