Abstract
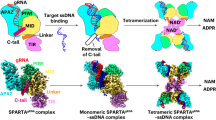
Short prokaryotic Ago accounts for most prokaryotic Argonaute proteins (pAgos) and is involved in defending bacteria against invading nucleic acids. Short pAgo associated with TIR-APAZ (SPARTA) has been shown to oligomerize and deplete NAD+ upon guide-mediated target DNA recognition. However, the molecular basis of SPARTA inhibition and activation remains unknown. In this study, we determined the cryogenic electron microscopy structures of Crenotalea thermophila SPARTA in its inhibited, transient and activated states. The SPARTA monomer is auto-inhibited by its acidic tail, which occupies the guide-target binding channel. Guide-mediated target binding expels this acidic tail and triggers substantial conformational changes to expose the Ago–Ago dimerization interface. As a result, SPARTA assembles into an active tetramer, where the four TIR domains are rearranged and packed to form NADase active sites. Together with biochemical evidence, our results provide a panoramic vision explaining SPARTA auto-inhibition and activation and expand understanding of pAgo-mediated bacterial defense systems.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Atomic coordinates, maps and structure factors of the reported cryo-EM structures have been deposited in the PDB under accession codes 8J84 (SPARTA-apo), 8J9G (monomeric SPARTA bound with guide-target, state 1: Monomer-GT1), 8J8H (monomeric SPARTA bound with guide-target, state 2: Monomer-GT2), 8J9P (SPARTA dimer bound with guide-target) and 8JAY (SPARTA tetramer bound with guide-target) and in the Electron Microscopy Data Bank under accession codes EMD-36059 (SPARTA-apo), 36095 (monomeric SPARTA bound with guide-target, state 1: Monomer-GT1), 36070 (monomeric SPARTA bound with guide-target, state 2: Monomer-GT2), 36114 (SPARTA dimer bound with guide-target) and 36138 (SPARTA tetramer bound with guide-target). The structures used for comparison in this study are available under PDB accession codes 5GQ9, 5UZB and 6O0Q. Source data are provided with this paper.
References
Hall, T. M. T. Structure and function of argonaute proteins. Structure 13, 1403–1408 (2005).
Meister, G. Argonaute proteins: functional insights and emerging roles. Nat. Rev. Genet. 14, 447–459 (2013).
Peters, L. & Meister, G. Argonaute proteins: mediators of RNA silencing. Mol. Cell 26, 611–623 (2007).
Makarova, K. S., Wolf, Y. I., Van der Oost, J. & Koonin, E. V. Prokaryotic homologs of Argonaute proteins are predicted to function as key components of a novel system of defense against mobile genetic elements. Biol. Direct 4, 29 (2009).
Swarts, D. et al. The evolutionary journey of Argonaute proteins. Nat. Struct. Mol. Biol. 21, 743–753 (2014).
Ryazansky, S., Kulbachinskiy, A. & Aravin, A. A. The expanded universe of prokaryotic Argonaute proteins. MBio 9, e01935–18 (2018).
Hegge, J., Swarts, D. & Van der Oost, J. Prokaryotic Argonaute proteins: novel genome-editing tools? Nat. Rev. Microbiol. 16, 5–11 (2018).
Willkomm, S., Makarova, K. S. & Grohmann, D. DNA silencing by prokaryotic Argonaute proteins adds a new layer of defense against invading nucleic acids. FEMS Microbiol. Rev. 42, 376–387 (2018).
Koopal, B., Mutte, S. K. & Swarts, D. C. A long look at short prokaryotic Argonautes. Trends Cell Biol. 33, 605–618 (2022).
Koopal, B. et al. Short prokaryotic Argonaute systems trigger cell death upon detection of invading DNA. Cell 185, 1471–1486.e19 (2022).
Garb, J. et al. Multiple phage resistance systems inhibit infection via SIR2-dependent NAD+ depletion. Nat. Microbiol. 7, 1849–1856 (2022).
Zaremba, M. et al. Short prokaryotic Argonautes provide defence against incoming mobile genetic elements through NAD+ depletion. Nat. Microbiol. 7, 1857–1869 (2022).
Wang, X. et al. Structural insights into mechanisms of Argonaute protein-associated NADase activation in bacterial immunity. Cell Res. 33, 699–711 (2023).
Medzhitov, P., Preston-Hurlburt, C. & Janeway, C. A. Jr. A human homologue of the Drosophila Toll protein signals activation of adaptive immunity. Nature 388, 394–397 (1997).
Van de Weyer, A. L. et al. A species-wide inventory of NLR genes and alleles in Arabidopsis thaliana. Cell 178, 1260–1272 (2019).
Newman, R. M., Salunkhe, P., Godzik, A. & Reed, J. C. Identification and characterization of a novel bacterial virulence factor that shares homology with mammalian Toll/interleukin-1 receptor family proteins. Infect. Immun. 74, 594–601 (2006).
Essuman, K. et al. TIR domain proteins are an ancient family of NAD+-consuming enzymes. Curr. Biol. 28, 421–430 (2018).
Ofir, G. et al. Antiviral activity of bacterial TIR domains via immune signaling molecules. Nature 600, 116–120 (2021).
Doron, S. et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science 359, eaar4120 (2018).
Essuman, K. et al. Shared TIR enzymatic functions regulate cell death and immunity across the tree of life. Science 377, eabo0001 (2022).
Morehouse, B. R. et al. STING cyclic dinucleotide sensing originated in bacteria. Nature 586, 429–433 (2020).
Morehouse, B. R. et al. Cryo-EM structure of an active bacterial TIR–STING filament complex. Nature 608, 803–807 (2022).
Tal, N. et al. Cyclic CMP and cyclic UMP mediate bacterial immunity against phages. Cell 184, 5728–5739 (2021).
Hogrel, G. et al. Cyclic nucleotide-induced helical structure activates a TIR immune effector. Nature 608, 808–812 (2022).
Martin, R. et al. Structure of the activated ROQ1 resistosome directly recognizing the pathogen effector XopQ. Science 370, eabd9993 (2020).
Ma, S. et al. Direct pathogen-induced assembly of an NLR immune receptor complex to form a holoenzyme. Science 370, eabe3069 (2020).
Ve, T., Williams, S. J. & Kobe, B. Structure and function of Toll/interleukin-1 receptor/resistance protein (TIR) domains. Apoptosis 20, 250–261 (2015).
Nimma, S. et al. Structural evolution of TIR-domain signalosomes. Front. Immunol. 12, 784484 (2021).
Burroughs, A. M., Iyer, L. M. & Aravind, L. Two novel PIWI families: roles in inter-genomic conflicts in bacteria and mediator-dependent modulation of transcription in eukaryotes. Biol. Direct 8, 13 (2013).
Burroughs, A. M., Ando, Y. & Aravind, L. New perspectives on the diversification of the RNA interference system: insights from comparative genomics and small RNA sequencing. RNA 5, 141–181 (2014).
Willkomm, S. et al. Structural and mechanistic insights into an archaeal DNA-guided Argonaute protein. Nat. Microbiol. 2, 17035 (2017).
Tuschl, T. & Patel, D. J. Crystal structure of A. aeolicus Argonaute, a site-specific DNA-guided endoribonuclease, provides insights into RISC-mediated mRNA cleavage. Mol. Cell 19, 405–419 (2005).
Wang, Y. et al. Structure of the guide-strand-containing argonaute silencing complex. Nature 456, 209–213 (2008).
Wang, Y. et al. Nucleation, propagation and cleavage of target RNAs in Ago silencing complexes. Nature 461, 754–761 (2009).
Wang, J. et al. Reconstitution and structure of a plant NLR resistosome conferring immunity. Science 364, eaav5870 (2019).
Shi, Y. et al. Structural basis of SARM1 activation, substrate recognition, and inhibition by small molecules. Mol. Cell 82, 1643–1659 (2022).
Sporny, M. et al. Structural basis for SARM1 inhibition and activation under energetic stress. eLife 9, e62021 (2020).
Ve, T. et al. Structural basis of TIR-domain-assembly formation in MAL- and MyD88-dependent TLR4 signaling. Nat. Struct. Mol. Biol. 24, 743–751 (2017).
Sheng, G. et al. Structure-based cleavage mechanism of Thermus thermophilus Argonaute DNA guide strand-mediated DNA target cleavage. Proc. Natl Acad. Sci. USA 111, 652–657 (2014).
Ma, J. B. et al. Structural basis for 5′-end-specific recognition of guide RNA by the A. fulgidus Piwi protein. Nature 434, 666–670 (2005).
Manakova, E. et al. Structural basis for sequence-specific recognition of guide and target strands by the Archaeoglobus fulgidus Argonaute protein. Sci. Rep. 13, 6123 (2023).
Golovinas, E. et al. Prokaryotic Argonaute from Archaeoglobus fulgidus interacts with DNA as a homodimer. Sci. Rep. 11, 4518 (2021).
Zeng, Z. et al. A short prokaryotic Argonaute activates membrane effector to confer antiviral defense. Cell Host Microbe 30, 930–943 (2022).
Kim, S. Y., Jung, Y. & Lim, D. Argonaute system of Kordia jejudonensis is a heterodimeric nucleic acid-guided nuclease. Biochem. Biophys. Res. Commun. 525, 755–758 (2020).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Acknowledgements
We would like to thank the Instrument Analysis Center at Shanghai Jiao Tong University for cryo-EM data collection. This work is supported by the National Key Research and Development Program of China (2018YFA0902000, Y.X.), STI2030-Major Projects (2021ZD0203400, Y.X.), the National Natural Science Foundation of China (32271330, M.C.; 31970547, Y.X.; 32000889, M.C.; 82321005, Y.X.), the Natural Science Foundation of Jiangsu Province (BK20190552, Y.X.), the Project Program of the State Key Laboratory of Natural Medicines, China Pharmaceutical University (SKLNMZZ202014, Y.X.) and the Fundamental Research Funds for the Central Universities (2632023GR17, Z.L.).
Author information
Authors and Affiliations
Contributions
L.G. and P.H. purified proteins and performed in vitro reconstitution of SPARTA complexes. Z.L. and M.C. determined cryo-EM structures. L.G. performed NAD+ hydrolysis assay, gel filtration assay and EMSA assay to biochemically characterize SPARTA. P.Y. prepared the SPARTA mutants. M.C., Y.X., L.G., P.H., Z.L., Y.C.S. and M.L. analyzed the data. P.H. prepared the figures. M.C. and Y.X. conceived experiments, supervised the work and wrote the manuscript. All authors discussed the results and contributed to the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Andrey Kulbachinskiy, Dinshaw Patel and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The reconstitution of SPARTA complexes.
a. NADase activity at 37 °C and 50 °C. The experiments were repeated in triplicates, and the data represents mean values ± s.d. b. The size exclusion chromatography of reconstituted SPARTA complexes.
Extended Data Fig. 2 Single-particle cryo-EM data processing.
Data processing scheme of SPARTA-apo (a), monomeric SPARTA bound with guide-target (Monomer-GT) (b) and SPARTA tetramer sample (c).
Extended Data Fig. 3 Cryo-EM density maps of the SPARTA complexes.
Maps of SPARTA-apo (a), monomeric SPARTA bound with guide-target (Monomer-GT1) (b), monomeric SPARTA bound with guide-target (Monomer-GT2) (c), SPARTA dimer (d) and SPARTA tetramer (e) are colored by local resolution, and gold-standard FSC of 0.143 resolution graphs is indicated for each complex.
Extended Data Fig. 5 The affinity of SPARTA to RNA guide.
a. The size exclusion chromatography of wild-type CrtSPARTA and C-tail truncated SPARTA (SPARTA_ΔCtail). The absorption at 260 nm and 280 nm is monitored and shown in red and blue, respectively. b. The binding affinity quantified by Electrophoretic Mobility Shift Assay (EMSA). 1 μM guide RNA was incubated with increasing amount of protein. The complex and 21nt free guide were resolved in Native-PAGE. The samples derived from the same experiment and gels were processed in parallel. The intensity of the free guide bands was measured, and the binding affinity was then analyzed using GraphPad Prism 9.3.0. The intensity of target band was quantified from single gel for Kd calculation.
Extended Data Fig. 6 Conformational changes induced by guide-target binding and propagation.
a. The cartoon representation of helixentry withdrawal upon guide-target binding. b. D-loop flipping upon the propagation of guide-target. c. Structure superimposition of SPARTA-apo, Monomer-GT1, Monomer-GT2 and dimer shows consecutive and unidirectional movement of the D-loop. d. The conformational changes likely coupling C-tail/TIR release with guide-target propagation. The structure of Monomer-GT2 (light blue) is superimposed onto Monomer-GT1 (grey), and the regions of substantial conformational changes in Monomer-GT2 are colored in yellow, orange, blue, and magenta.
Extended Data Fig. 7 The size exclusion chromatography of SPARTA mutants.
The mutant of D-loop truncation (a), and the Ago-Ago interface mutant SPARTA_ERDAAA (b) and SPARTA_YKAA (c), incubated with guide RNA and target DNA.
Extended Data Fig. 8 The Cryo-EM maps showing the flexibility of the TIR domain.
Comparison of TIR density in Monomer-GT1 (a), Monomer-GT2 (b), SPARTA dimer (c), and SPARTA tetramer (d).
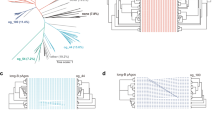
Extended Data Fig. 9 The typical assemblies of TIR domains.
a. The structure of TIR domain in SPARTA. The secondary structures are indicated. Eukaryotic TIR domains are primarily arranged either as ‘scaffold assembly’ (b) or ‘enzymatic assembly’ (c), corresponding to parallel or anti-parallel two-stranded assembly via distinct interfaces, respectively. The involved interfaces are indicated by the frames.
Supplementary information
Supplementary Information
Supplementary Tables 1–3 and Supplementary Fig. 1.
Source data
Source Data Extended Data Fig. 1
Unprocessed gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guo, L., Huang, P., Li, Z. et al. Auto-inhibition and activation of a short Argonaute-associated TIR-APAZ defense system. Nat Chem Biol 20, 512–520 (2024). https://doi.org/10.1038/s41589-023-01478-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-023-01478-0
This article is cited by
-
Insights into the modulation of bacterial NADase activity by phage proteins
Nature Communications (2024)
-
DNA-targeting short Argonautes complex with effector proteins for collateral nuclease activity and bacterial population immunity
Nature Microbiology (2024)
-
Structural basis of antiphage immunity generated by a prokaryotic Argonaute-associated SPARSA system
Nature Communications (2024)