Abstract
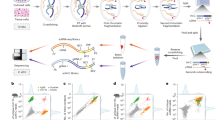
Signaling pathways that drive gene expression are typically depicted as having a dozen or so landmark phosphorylation and transcriptional events. In reality, thousands of dynamic post-translational modifications (PTMs) orchestrate nearly every cellular function, and we lack technologies to find causal links between these vast biochemical pathways and genetic circuits at scale. Here we describe the high-throughput, functional assessment of phosphorylation sites through the development of PTM-centric base editing coupled to phenotypic screens, directed by temporally resolved phosphoproteomics. Using T cell activation as a model, we observe hundreds of unstudied phosphorylation sites that modulate NFAT transcriptional activity. We identify the phosphorylation-mediated nuclear localization of PHLPP1, which promotes NFAT but inhibits NFκB activity. We also find that specific phosphosite mutants can alter gene expression in subtle yet distinct patterns, demonstrating the potential for fine-tuning transcriptional responses. Overall, base editor screening of PTM sites provides a powerful platform to dissect PTM function within signaling pathways.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw mass spectrometry data and metadata can be accessed at ftp://MSV000092965@massive.ucsd.edu. Raw RNA sequencing data can be accessed at GEO accession ID GSE244164.
Code availability
The code for the base editor design tool is available at https://github.com/mhegde/base-editor-design-tool.
References
Hunter, T. Why nature chose phosphate to modify proteins. Philos. Trans. R. Soc. Lond. B Biol. Sci. 367, 2513–2516 (2012).
Manning, G., Whyte, D. B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002).
Chen, M. J., Dixon, J. E. & Manning, G. Genomics and evolution of protein phosphatases. Sci. Signal 10, eaag1796 (2017).
Katrancha, S. M. et al. Trio haploinsufficiency causes neurodevelopmental disease-associated deficits. Cell Rep. 26, 2805–2817.e9 (2019).
Martinez-Val, A. et al. Spatial-proteomics reveals phospho-signaling dynamics at subcellular resolution. Nat. Commun. 12, 7113 (2021).
Koch, H. et al. Phosphoproteome profiling reveals molecular mechanisms of growth-factor-mediated kinase inhibitor resistance in EGFR-overexpressing cancer cells. J. Proteome Res 15, 4490–4504 (2016).
Paulo, J. A. & Gygi, S. P. A comprehensive proteomic and phosphoproteomic analysis of yeast deletion mutants of 14-3-3 orthologs and associated effects of rapamycin. Proteomics 15, 474–486 (2015).
Needham, E. J., Parker, B. L., Burykin, T., James, D. E. & Humphrey, S. J. Illuminating the dark phosphoproteome. Sci. Signal 12, eaau8645 (2019).
Hornbeck, P. V. et al. PhosphoSitePlus, 2014: mutations, PTMs and recalibrations. Nucleic Acids Res. 43, D512–D520 (2015).
Dixit, A. et al. Perturb-Seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866.e17 (2016).
Parnas, O. et al. A genome-wide CRISPR screen in primary immune cells to dissect regulatory networks. Cell 162, 675–686 (2015).
Shifrut, E. et al. Genome-wide CRISPR screens in primary human T cells reveal key regulators of immune function. Cell 175, 1958–1971.e15 (2018).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Meyers, R. M. et al. Computational correction of copy number effect improves specificity of CRISPR–Cas9 essentiality screens in cancer cells. Nat. Genet. 49, 1779–1784 (2017).
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770–788 (2018).
Hanna, R. E. et al. Massively parallel assessment of human variants with base editor screens. Cell 184, 1064–1080.e20 (2021).
Lue, N. Z. et al. Base editor scanning charts the DNMT3A activity landscape. Nat. Chem. Biol. 19, 176–186 (2023).
Li, H. et al. Assigning functionality to cysteines by base editing of cancer dependency genes. Nat. Chem. Biol. 19, 1320–1330 (2023).
Schmidt, R. et al. Base-editing mutagenesis maps alleles to tune human T cell functions. Nature 625, 805–812 (2024).
Yeh, W.-H., Chiang, H., Rees, H. A., Edge, A. S. B. & Liu, D. R. In vivo base editing of post-mitotic sensory cells. Nat. Commun. 9, 2184 (2018).
Abraham, R. T. & Weiss, A. Jurkat T cells and development of the T-cell receptor signalling paradigm. Nat. Rev. Immunol. 4, 301–308 (2004).
Larange, A. et al. A regulatory circuit controlled by extranuclear and nuclear retinoic acid receptor α determines T cell activation and function. Immunity 56, 2054–2069 (2023).
Abelin, J. G. et al. Workflow enabling deepscale immunopeptidome, proteome, ubiquitylome, phosphoproteome, and acetylome analyses of sample-limited tissues. Nat. Commun. 14, 1851 (2023).
Krug, K. et al. A curated resource for phosphosite-specific signature analysis. Mol. Cell Proteom. 18, 576–593 (2019).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Richter, M. F. et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat. Biotechnol. 38, 883–891 (2020).
Huang, T. P., Newby, G. A. & Liu, D. R. Precision genome editing using cytosine and adenine base editors in mammalian cells. Nat. Protoc. 16, 1089–1128 (2021).
Kluesner, M. G. et al. EditR: a method to quantify base editing from Sanger sequencing. CRISPR J. 1, 239–250 (2018).
Helou, Y. A., Nguyen, V., Beik, S. P. & Salomon, A. R. ERK positive feedback regulates a widespread network of tyrosine phosphorylation sites across canonical T cell signaling and actin cytoskeletal proteins in Jurkat T cells. PLoS ONE 8, e69641 (2013).
Bottini, N. et al. Activation of ZAP-70 through specific dephosphorylation at the inhibitory Tyr-292 by the low molecular weight phosphotyrosine phosphatase (LMPTP). J. Biol. Chem. 277, 24220–24224 (2002).
Di Bartolo, V. et al. Tyrosine 319, a newly identified phosphorylation site of ZAP-70, plays a critical role in T cell antigen receptor signaling. J. Biol. Chem. 274, 6285–6294 (1999).
Jutz, S. et al. Assessment of costimulation and coinhibition in a triple parameter T cell reporter line: Simultaneous measurement of NF-κB, NFAT and AP-1. J. Immunol. Methods 430, 10–20 (2016).
Li, J. et al. Functional phosphoproteomics in cancer chemoresistance using CRISPR-mediated base editors. Adv. Sci. 9, e2200717 (2022).
Pihlajamaa, P., Kauko, O., Sahu, B., Kivioja, T. & Taipale, J. A competitive precision CRISPR method to identify the fitness effects of transcription factor binding sites. Nat. Biotechnol. 41, 197–203 (2023).
Quesada, A. E. et al. Clinico-pathologic characteristics and outcomes of the World Health Organization (WHO) provisional entity de novo acute myeloid leukemia with mutated RUNX1. Mod. Pathol. 33, 1678–1689 (2020).
Huang, K. et al. Genome-wide CRISPR-Cas9 screening identifies NF-κB/E2F6 responsible for EGFRvIII-associated temozolomide resistance in Glioblastoma. Adv. Sci. 6, 1900782 (2019).
Cheng, F. H. C. et al. E2F6 functions as a competing endogenous RNA, and transcriptional repressor, to promote ovarian cancer stemness. Cancer Sci. 110, 1085–1095 (2019).
Johnson, J. L. et al. An atlas of substrate specificities for the human serine/threonine kinome. Nature 613, 759–766 (2023).
Mognol, G. P. et al. Targeting the NFAT:AP-1 transcriptional complex on DNA with a small-molecule inhibitor. Proc. Natl Acad. Sci. USA 116, 9959–9968 (2019).
Wang, B. et al. Integrative analysis of pooled CRISPR genetic screens using MAGeCKFlute. Nat. Protoc. 14, 756–780 (2019).
Raudvere, U. et al. g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47, W191–W198 (2019).
Hogan, P. G., Chen, L., Nardone, J. & Rao, A. Transcriptional regulation by calcium, calcineurin, and NFAT. Genes Dev. 17, 2205–2232 (2003).
Ortega-Pérez, I. et al. c-Jun N-terminal kinase (JNK) positively regulates NFATc2 transactivation through phosphorylation within the N-terminal regulatory domain. J. Biol. Chem. 280, 20867–20878 (2005).
Ishitani, T. et al. The TAK1-NLK mitogen-activated protein kinase cascade functions in the Wnt-5a/Ca(2+) pathway to antagonize Wnt/beta-catenin signaling. Mol. Cell. Biol. 23, 131–139 (2003).
MacDonnell, S. M. et al. CaMKII negatively regulates calcineurin-NFAT signaling in cardiac myocytes. Circ. Res. 105, 316–325 (2009).
Anshabo, A. T., Milne, R., Wang, S. & Albrecht, H. CDK9: a comprehensive review of its biology, and its role as a potential target for anti-cancer agents. Front Oncol. 11, 678559 (2021).
Phee, H. et al. Pak2 is required for actin cytoskeleton remodeling, TCR signaling, and normal thymocyte development and maturation. eLife 3, e02270 (2014).
Pareek, T. K. et al. Cyclin-dependent kinase 5 activity is required for T cell activation and induction of experimental autoimmune encephalomyelitis. J. Exp. Med. 207, 2507–2519 (2010).
Ochoa, D. et al. The functional landscape of the human phosphoproteome. Nat. Biotechnol. 38, 365–373 (2020).
Chen, M. et al. Identification of PHLPP1 as a tumor suppressor reveals the role of feedback activation in PTEN-mutant prostate cancer progression. Cancer Cell 20, 173–186 (2011).
Nitsche, C. et al. The phosphatase PHLPP1 regulates Akt2, promotes pancreatic cancer cell death, and inhibits tumor formation. Gastroenterology 142, 377–87.e1–5 (2012).
Cohen Katsenelson, K. et al. PHLPP1 counter-regulates STAT1-mediated inflammatory signaling. eLife 8, e48609 (2019).
Patterson, S. J. et al. Cutting edge: PHLPP regulates the development, function, and molecular signaling pathways of regulatory T cells. J. Immunol. 186, 5533–5537 (2011).
Balasuriya, N. et al. Genetic code expansion and live cell imaging reveal that Thr-308 phosphorylation is irreplaceable and sufficient for Akt1 activity. J. Biol. Chem. 293, 10744–10756 (2018).
Feske, S. Calcium signalling in lymphocyte activation and disease. Nat. Rev. Immunol. 7, 690–702 (2007).
Gwack, Y. et al. A genome-wide Drosophila RNAi screen identifies DYRK-family kinases as regulators of NFAT. Nature 441, 646–650 (2006).
Liu, H. et al. NFATc1 phosphorylation by DYRK1A increases its protein stability. PLoS ONE 12, e0172985 (2017).
Thompson, B. J. et al. DYRK1A controls the transition from proliferation to quiescence during lymphoid development by destabilizing Cyclin D3. J. Exp. Med. 212, 953–970 (2015).
Chen, J. et al. NR4A transcription factors limit CAR T cell function in solid tumours. Nature 567, 530–534 (2019).
Beltrao, P. et al. Systematic functional prioritization of protein posttranslational modifications. Cell 150, 413–425 (2012).
Beltrao, P., Bork, P., Krogan, N. J. & van Noort, V. Evolution and functional cross-talk of protein post-translational modifications. Mol. Syst. Biol. 9, 714 (2013).
Liu, N., Guo, Y., Ning, S. & Duan, M. Phosphorylation regulates the binding of intrinsically disordered proteins via a flexible conformation selection mechanism. Commun. Chem. 3, 1–9 (2020).
Nicolaou, S. T., Hebditch, M., Jonathan, O. J., Verma, C. S. & Warwicker, J. PhosIDP: a web tool to visualize the location of phosphorylation sites in disordered regions. Sci. Rep. 11, 9930 (2021).
Trinidad, J. C. et al. Global identification and characterization of both O-GlcNAcylation and phosphorylation at the murine synapse. Mol. Cell Proteom. 11, 215–229 (2012).
Ren, X. et al. High-throughput PRIME-editing screens identify functional DNA variants in the human genome. Mol. Cell 83, 4633–4645.e9 (2023).
Hiatt, J. et al. Efficient generation of isogenic primary human myeloid cells using CRISPR-Cas9 ribonucleoproteins. Cell Rep. 35, 109105 (2021).
Mari, T. et al. In vitro Kinase-to-Phosphosite database (iKiP-DB) predicts kinase activity in phosphoproteomic fatasets. J. Proteome Res 21, 1575–1587 (2022).
Hwang, G.-H. et al. Web-based design and analysis tools for CRISPR base editing. BMC Bioinform. 19, 542 (2018).
Chen, P. J. et al. Enhanced prime editing systems by manipulating cellular determinants of editing outcomes. Cell 184, 5635–5652.e29 (2021).
Acknowledgements
We thank A. Haber, P. Vijayanand, E. Kvedaraite, B. Hamilton, A. Rubin, T.M. Yaron and M. Gentili for useful discussion. We also thank G. and S. Clouse and S. Carr for support, as well as P. Guo and the Nikon Imaging Center at the University of California San Diego for the support on microscopy experiments. This work was supported by National Institutes of Health (NIH) grant nos. R35GM147554 and R01CA279795 (S.A.M.); NIH grant no. R35GM122523 (A.C.N.); NIH grant nos. U01AI142756, R35GM118062, RM1HG009490 and HHMI (D.R.L.); NIH grant nos. R01AI040127 and R01AI109842 (P.G.H.); Stem Cell Network Jump Start Award (no. ECR-C4R1-7) for C.G.d.B. who is a Michael Smith Health Research BC Scholar; and the University of California San Diego Graduate Training Program in Cellular and Molecular Pharmacology (grant no. T32 GM007752) and the National Science Foundation Graduate Research Fellowship Program (no. DGE-1650112) (A.C.J.). The NovaSeq 6000 was acquired through the Shared Instrumentation Grant Program (S10) S10OD025052; La Jolla Institute for Immunology Next-Generation Sequencing Core Facility RRID:SCR_023107. FACSAria-3 was acquired through the Shared Instrumentation Grant Program (S10): RR027366; La Jolla Institute for Immunology Flow Cytometry Core RRID:SCR_014832.
Author information
Authors and Affiliations
Contributions
S.A.M. conceptualized the study. P.H.K., A.C.J., M.H., C.G.d.B., G.A.N. and S.A.M. developed the methodology. M.H. and J.G.D. wrote the software. P.H.K., A.A.D.S., A.C.J. and S.A.M. validated the results. A.A.D.S., M.E.O., R.B., S.A., R.A.G., G.A.N. and S.A.M. performed the formal analyses. P.H.K., A.A.D.S., M.B., A.C.J., M.E.O., M.I.M., N.P., P.G.H., R.B., A.C.N., S.A., R.A.G., C.G.d.B., G.A.N. and S.A.M. performed the investigations. M.B., N.P., J.L., T.L., P.G.H., D.R.L., J.G.D., G.A.N., C.G.d.B. and S.A.M. provided resources. A.A.D.S., M.E.O., M.H., M.I.M., R.B., J.G.D., S.A., R.A.G. and S.A.M. curated the data. All authors wrote the paper. P.H.K., A.A.D.S., A.C.J., M.E.O., S.A. and S.A.M. visualized the findings. S.A.M. supervised the project and was the project administrator. S.A.M. acquired funding.
Corresponding author
Ethics declarations
Competing interests
D.R.L. is a consultant and/or equity owner for Prime Medicine, Beam Therapeutics, Pairwise Plants, Chroma Medicine and Nvelop Therapeutics—companies that use or deliver genome editing or epigenome engineering agents. The other authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Rita Strack, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 associated with Fig. 1 Kennedy et al.
Phosphoproteomics quality controls. a) Principal component analysis of all phosphoproteomics samples prior to differential expression analysis. b) Multi-scatter plot comparing all samples to each other pairwise. Pearson’s r is shown. Colors of samples are the same as in Extended Data Fig. 1a.
Extended Data Fig. 2 associated with Fig. 2 Kennedy et al.
Gating strategy for CD69 staining analyzed by flow cytometry.
Extended Data Fig. 3 associated with Fig. 3 Kennedy et al.
Extended Data Fig. 4 associated with Fig. 4 Kennedy et al.
Quality control and characterization of phosphosite base editing coupled to NFAT activity reporters. a) Pairwise Spearman correlations between all normalized log transformed read counts across replicates and experimental conditions. 0.4 is the lower limit cut off in black. b) Mean (across replicates) sgRNA counts for individual sgRNAs prior to collapsing redundant phosphosite targets in the GFP high and low bins. Regression line is shown. c) Percentage of phosphosite targets with one or more protospacer sequences. d) g:Profiler (gene-centric) analysis of genes with phosphosite mutations enriched in the GFP low or GFP high bins. For the x-axis the normalized enrichment score (NES) was multiplied by the -log10 FDR. e) GSEA (gene-centric) analysis of gene sets enriched in the GFP high bin. TCR Calcium Pathway is bolded. f) Proportion of phosphosite targets that contain a putative bystander edit in the library as a whole and in the sorted GFP bins. Student’s two sample T test P value is shown. g) Scatterplot comparing the F statistic from the phosphoproteomic analysis, a proxy for magnitude and reproducibility of abundance changes across the four time points, and the log2 fold change GFPhigh/low bins calculated by MAGeCK. Horizontal red dashed line delineates nominal p value of < 0.05 from the moderated F test of the phosphoproteomics data. h) Scatterplot comparing the log2 fold change GFPhigh/low bins calculated by MAGeCK to the predicted functional score from the machine learning analysis in Ochoa et al.49. Inset shows the full data structure while the scatter plot is a zoom of points above a predicted functional score of 0.5. Horizontal red dashed line delineates a score threshold determined in Ochoa et al.49. i) Distribution of predicted functional scores from Ochoa et al.49 for all data points in the GFP screen, the phosphosite mutants that increased (‘up’ in red) or decreased GFP levels (‘down’ in purple). P values for comparison to the whole data set are shown. Data points represent the mean log2 FC (GFPhigh/GFP low) of four transduction replicates. P values for an ANOVA test followed by uncorrected Fisher’s least significant difference for multiple comparisons.
Extended Data Fig. 5 associated with Fig. 5, Kennedy et al.
EditR software analysis28 plots outlining bystander base editing levels for PHLPP1 S118P and MAPK1 Y187C prior to single cell clone isolation.
Extended Data Fig. 6 associated with Fig. 6, Kennedy et al.
a) Log2 fold change of select T cell genes differentially expressed between PHLPP1 S118P and MAPK1 Y187C mutant cells, compared to HEK3 control cells. b) Gating strategy for intracellular GZMB staining and analysis by flow cytometry.
Supplementary information
Supplementary Data 1
The .json file of phosphoproteomic data in Fig. 1b, which can be explored using Morpheus https://software.broadinstitute.org/morpheus/.
Supplementary Data 2
The .json file of the transcriptional data in Fig. 5. Differentially expressed genes of phosphosite mutants at 0 and 6 h post T cell activation, where values are in log2 fold changes to the mean. Only statistically significant genes are plotted. These data can be explored using https://software.broadinstitute.org/morpheus/.
Supplementary Tables
Supplementary Table 1. Phosphoproteomics analyses processed by Spectrum Mill and statistically tested by Protigy using the moderated F-test. ‘modF’ provides all analysis results and measurement values. ‘Class vector’ provides sample key for TMT channels. ‘Description of table header’ refers to modF and describes where which analysis comes from. Supplemental Table 2A. ABE8e-targetable phosphosites. Supplementary Table 2B. BE4-targetable phosphosites. Supplementary Table 3. Differential analysis of sgRNA abundances between pre- and post-ABE8e protein introduction. MAGeCK P value is shown. Supplementary Table 4. Differential analysis of sgRNA abundances between GFP high and GFP low bins–NFAT activity screen. MAGeCK P value is shown. Supplementary Table 5. RNA sequencing analysis of activated Jurkat T cells with various phosphosite mutations. P values were determined in using a variant of the negative binomial exact test in Cell Ranger (10x Genomics). Supplementary Table 6. Oligonucleotide sequences for sgRNA in vitro transcription and cloning into pRDA_118 vector for lentivirus production.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kennedy, P.H., Alborzian Deh Sheikh, A., Balakar, M. et al. Post-translational modification-centric base editor screens to assess phosphorylation site functionality in high throughput. Nat Methods (2024). https://doi.org/10.1038/s41592-024-02256-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41592-024-02256-z