Abstract
Path integration (PI) is a highly conserved, self-motion-based navigation strategy. Since the discovery of grid cells in the medial entorhinal cortex, neurophysiological data and computational models have suggested that these neurons serve PI. However, more direct empirical evidence supporting this hypothesis has been missing due to a lack of selective manipulations of grid cell activity and suitable behavioral assessments. Here we report that selective disruption of grid cell activity in mice can be achieved by removing NMDA glutamate receptors from the retro-hippocampal region and that disrupted grid cell firing accounts for impaired PI performance. Notably, the genetic manipulation did not affect the activity of other spatially selective cells in the medial entorhinal cortex and the hippocampus. By directly linking grid cell activity to PI, these results contribute to a better understanding of how grid cells support navigation and spatial memory.
Similar content being viewed by others
Main
Path integration (PI) is a basic form of navigation. It involves continuous integration of self-movement so that global vectors connecting past and present locations can be continuously updated. Animals memorize these vectors to travel between locations using straight paths1,2,3. Experiments combining brain lesions and behavioral assays pointed to the hippocampus (HP) and the retro-hippocampal (RH) regions as major players in PI4,5,6. The RH region (comprising the entorhinal cortex, subiculum, presubiculum, postsubiculum and parasubiculum)7 contains different types of spatially tuned cell types, i.e., grid, head direction, border and speed cells, which are thought to contribute to space representation8,9,10,11,12,13. Grid cells display a stable periodic hexagonal firing pattern, and they are modulated by heading direction and running speed9,14,15. Together, these features can, in principle, enable the integration of self-motion-based information about traveled distance and running direction into a metric representation of space necessary for PI14,16,17.
Evaluating the link between grid cell activity and PI entails both adequate behavioral assays and selective manipulation of neuronal activity. Recently, we developed a behavioral assay, the L-maze, to specifically test PI in mice3. Selective manipulation of grid cell activity has been difficult, as molecular markers suitable for targeting distinct spatially tuned cells are not known. Consequently, the hypothesis that grid cell activity serves PI has not been experimentally tested to date. Here we achieved selective disruption of grid cell activity by removing NMDA glutamate receptors (NMDARs) from the RH region of newborn mice and directly tested whether grid cells support PI.
Results
NMDAR ablation in the RH region
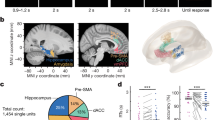
To generate mice lacking NMDARs in the RH region (NR1 RH−/− mice), we removed the constitutive NR1 subunit18 by bilaterally injecting an adeno-associated virus (AAV) expressing Cre-recombinase (Cre) and GFP into the medial entorhinal cortex (MEC) of 4-d-old mice carrying loxP-flanked NR1 alleles (henceforth: NR1 flox mice; Fig. 1a). Four weeks after injection, we confirmed NMDAR ablation in infected (GFP+) cells by performing whole-cell patch-clamp recordings in the MEC obtained from NR1 RH−/−, wild-type (WT) and WT-GFP+ mice WT: n = 3, WT-GFP+: n = 4, NR1 RH−/−: n = 4). The amplitude of AMPA receptor (AMPAR)-mediated evoked excitatory postsynaptic currents was similar in the three groups of mice (mean ± s.e.m., WT: –240.54 ± 19.54 pA, n = 9; WT-GFP+: –204.2 ± 24.21 pA, n = 10; NR1 RH−/−: –230.45 ± 19.66 pA, n = 11; one-way ANOVA: F 2,27 = 0.67, P = 0.52). In contrast, NMDAR-mediated currents were virtually abolished in NR1 RH−/− mice (mean ± s.e.m., WT: 147.13 ± 25.29 pA, n = 9; WT-GFP+: 142.2 ± 18.13 pA, n = 10; NR1 RH−/−: 6.48 ± 1.19 pA, n = 11; Kruskal-Wallis test followed by Dunn's multiple comparisons: H = 20.24, P = 0.0001, WT vs. WT-GFP+: P = 0.98; WT vs. NR1 RH−/−: P = 0.0001; WT-GFP+ vs. NR1 RH−/−: P = 0.0001). As a result, the NMDAR/AMPAR ratio in cells of NR1 RH−/− mice was significantly smaller than in cells of WT and WT-GFP+ mice (Fig. 1b).
a, Four-day-old NR1 flox mice received bilateral MEC injections of AAV-Syn-Cre-GFP. The experimental protocol included whole-cell patch-clamp recordings, as well as behavioral and in vivo electrophysiological recordings, according to the depicted schedule. b, Left: representative traces of AMPAR- and NMDAR-mediated currents (black and blue lines, respectively) obtained from MEC cells of noninjected WT mice, WT mice injected with AAV-Syn-Cre-GFP (WT-GFP+) and NR1 flox mice injected with AAV-Syn-Cre-GFP (NR1 RH−/−). d-AP5 blocked NMDAR-mediated currents (red traces). Right: mean (± s.e.m.) ratio of NMDAR/AMPAR-mediated excitatory postsynaptic currents (Kruskal–Wallis test followed by Dunn's multiple comparisons, H = 20.27, P = 0.0001; WT vs. WT-GFP+: P = 0.98; WT vs. NR1 RH−/−: P = 0.0001; WT-GFP+ vs. NR1 RH−/−: P = 0.0001; WT: n = 9 cells from 3 mice, WT-GFP+: n = 10 cells from 4 mice, NR1 RH−/−: n = 11 cells from 4 mice). Cells were used as experimental units. All tests were two-sided. ***P ≤ 0.0001. c, Slide scanner images delineating areas subsequently used to quantify the extent of viral infection. In each brain section, we first delimited the area corresponding to the RH and HP regions (magenta and light green lines, respectively) and measured the areas covered by the GFP signal (red and green lines, respectively). This quantification was performed in all mice using all brain serial sections (~100 sections per mouse). Left and right panels correspond to brain sections from NR1 RH−/− and NR1 RH&HP−/− mice, respectively. d, Relationship between the percent of infection in the RH and HP regions used to assign the mice to the experimental groups.
A second group of injected mice was subjected to behavioral tests and/or in vivo electrophysiological recordings after they reached adulthood (Fig. 1a). After completion of the experiments, the location and volume of the viral infection were determined for each mouse. As expected, the virus infection (indicated by the GFP signal) was not always confined to the MEC. In some mice, it spread to other structures within the RH region and to the HP. Using serial sections of each brain, we measured the extent of the viral infection in the RH and HP regions (henceforth, percent infection in the RH and HP regions, respectively; Fig. 1c and Supplementary Fig. 1). A threshold value of 3% was arbitrarily set to assign the mice to one of the two experimental groups: (i) NR1 RH−/− mice, in which the percent infection in the HP region was <3%, and (ii) NR1 RH&HP−/− mice, in which the percent infection in the HP region was >3% (Fig. 1d). The control group included wild-type mice injected with AAV-Syn-Cre-GFP and NR1 flox mice injected with AAV-Syn-Tomato. For further details about the viral infection quantification and its limitations, see Supplementary Fig. 1 and Methods.
Disruption of grid cell activity in NR1 RH−/− and NR1 RH&HP−/− mice
We conducted in vivo extracellular recordings to investigate the effect of region-specific NMDAR ablation on MEC neuronal activity. To this end, we unilaterally implanted five to six independently movable tetrodes in the MEC and recorded the activity of neurons as mice ran in a square arena. The number of recording sessions, the position of the tetrodes, the isolation distance of the recorded neurons and the running speed of the mice in the arena were similar across experimental groups (Supplementary Table 1 and Supplementary Fig. 2). We sorted the recorded neurons according to their mean firing rate into putative principal cells (<5 Hz) and interneurons (≥5 Hz) and found that the numbers of cells recorded per session, the percentages of cells and their firing rates did not differ across groups for both neuronal types (Supplementary Table 1). Moreover, rhythmic firing at theta frequency was preserved in NR1 RH−/− and NR1 RH&HP−/− mice (Supplementary Fig. 3a–c). Theta rhythmicity in NR1 RH&HP−/− mice tended to be higher than in the other two groups, and theta modulated neurons in mutant mice had a lower peak frequency than neurons in control mice (Supplementary Fig. 3d,e).
Next, we characterized the spatial properties of the entire population of principal neurons (unless otherwise specified, all scores were computed using mice as experimental units). To estimate their spatial selectivity, i.e., the extent to which they fired in particular locations of the arena, we calculated the spatial information score. Principal cells in NR1 RH−/− and NR1 RH&HP−/− mice showed significantly lower spatial information scores than in control mice (Fig. 2a). To measure how well multiple firing fields fitted to the typical hexagonal pattern of grid cells, we calculated the grid score. Principal cells in NR1 RH−/− and NR1 RH&HP−/− mice had significantly lower grid scores than in control mice (Fig. 2a). In addition to the comparisons across groups, we took advantage of the variability in injection size (Fig. 1d) and correlated the percent of infection and the different electrophysiological variables. Notably, both spatial information and grid scores of principal cells in mutant mice were inversely correlated with the percent of infection in the RH region but not in the HP region (Fig. 2b and Supplementary Table 2). As expected, these correlations were absent in control mice, in which the virus did not lead to the ablation of NMDARs (Supplementary Table 2). The remaining firing features of principal neurons were similar across groups (Fig. 2a). These were: (i) the number of firing fields; (ii) the head direction score, indicating how well the firing was tuned to a preferred head direction; (iii) the border score, indicating the tendency to fire along a wall of the arena; and (iv) the speed score, indicating the extent to which the firing was modulated by the running speed of the animal. Similar results were obtained when statistics were performed using cells as experimental units (Supplementary Table 3). Therefore, MEC principal neurons in mutant mice exhibited disrupted grid spatial periodicity and spatial selectivity.
a, Firing properties of MEC principal cells for the three genotypes. Mean (± s.e.m.) spatial information (inf.) score (Kruskal–Wallis test followed by Dunn's multiple comparisons, H = 13.69, P = 0.001; control vs. NR1 RH−/−: P = 0.02; control vs. NR1 RH&HP−/−: P = 0.0007; NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.44), grid score (one-way ANOVA followed by Tukey's multiple comparisons, F 2,35 = 6.32, P = 0.004; control vs. NR1 RH−/−: P = 0.004; control vs. NR1 RH&HP−/−: P = 0.01; NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.96), number of firing fields (Kruskal–Wallis test, H = 2.73, P = 0.26), HD score (one-way ANOVA, F 2,35 = 0.13, P = 0.88), border score (one-way ANOVA, F 2,35 = 1.43, P = 0.25) and speed score (one-way ANOVA, F 2,35 = 0.44, P = 0.65). b, Relationship between the spatial information (inf.) scores and grid scores of principal cells and the percent infection in the RH and HP regions (Pearson correlation, inf. score: percent RH: r = –0.31, P = 0.04; percent HP: r = –0.2, P = 0.28; grid score: percent RH: r = –0.35, P = 0.03; percent HP: r = –0.09, P = 0.64; n = 30). c, Mean (± s.e.m.) percentage of grid cells (one-way ANOVA followed by Tukey's multiple comparisons, F 2,35 = 4.45, P = 0.02; control vs. NR1 RH−/−: P = 0.02; control vs. NR1 RH&HP−/−: P = 0.03; NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.97), ISS cells (one-way ANOVA followed by Tukey's multiple comparisons, F 2,35 = 4.66, P = 0.02; control vs. NR1 RH−/−: P = 0.03; control vs. NR1 RH&HP−/−: P = 0.005; NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.26), border cells (one-way ANOVA, F 2,35 = 0.51, P = 0.60), HD cells (one-way ANOVA, F 2,35 = 0.07, P = 0.93), SM cells (one-way ANOVA, F 2,35 = 0.18, P = 0.83) and NSS cells (one-way ANOVA, F 2,35 = 0.92, P = 0.41). d, Relationship between percent grid cells and percent infection in the RH and HP regions (Pearson correlation, percent RH: r = –0.41, P = 0.02; percent HP: r = –0.19, P = 0.30; n = 30). For a and c, control: n = 8, NR1 RH−/−: n = 19, NR1 RH&HP−/−: n = 11. Mice were used as experimental units. All tests were two-sided. **P ≤ 0.01, *P < 0.05, ns: not significant. Yellow, red and blue symbols correspond to control, NR1 RH−/− and NR1 RH&HP−/− mice, respectively.
Within MEC principal neurons, we defined six mutually exclusive cell populations: grid cells, irregular spatially selective (ISS) cells, border cells, head direction (HD) cells, speed modulated (SM) cells and nonspatially selective (NSS) cells (Fig. 2c; see Methods for details). We determined the percentage of each cell type per mouse and found that only the percentages of grid cells and ISS cells changed across genotypes, i.e., the percentages of grid cells and ISS cells were significantly lower in mutant than in control mice (Fig. 2c). Therefore, in mutant mice, disrupted grid periodicity and spatial selectivity (Fig. 2a) led to a decrease in the percentages of grid cells and ISS cells, which seemed to lead to a concomitant increase in the percentage of NSS cells, although the differences did not reach significance (Fig. 2c). The percentages of border cells, HD cells and SM cells were similar across genotypes (Fig. 2c). Furthermore, we found that in mutant mice the percentage of grid cells was inversely correlated with the percent infection in the RH region but not in the HP region (Fig. 2d, Supplementary Table 2). Such correlation was absent in control mice (Supplementary Table 2).
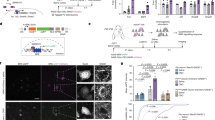
Next, we further characterized the firing properties of each of the six principal cell populations (Fig. 3, Table 1 and Supplementary Table 3). Grid cells were found in 86% of control mice (7 of 8) and in 79–72% of mutant mice (15 of 19 in NR1 RH−/− mice and 8 of 11 in NR1 RH&HP−/− mice). Critically, in the mutant mice in which grid cells were found, the percentage of grid cells was significantly smaller than in controls (mean ± s.e.m., control: 14.26 ± 4.25%, NR1 RH−/−: 4.69 ± 0.61%, NR1 RH&HP−/−: 4.61 ± 0.98%; one-way ANOVA followed by Tukey's multiple comparisons: F 2,27 = 6.55, P = 0.005; control vs. NR1 RH−/−, P = 0.004; control vs. NR1 RH&HP−/−, P = 0.03; NR1 RH−/− vs. NR1 RH&HP−/−, P = 0.92). Moreover, in around 25% of mutant mice, only one grid cell was found (4 of 15 in NR1 RH−/− mice, 2 of 8 in NR1 RH&HP−/− mice), which was never the case in control mice (Supplementary Fig. 4). In sum, mutants had significantly fewer grid cells than control mice (Fig. 2c and Supplementary Fig. 4). The few grid cells recorded in mutant mice showed grid scores similar to those in controls (Fig. 3a and Table 1), but they were less spatially selective, i.e., grid cells in mutant mice showed significantly lower spatial information scores than those in control mice (Fig. 3a and Table 1). Also, grid cells in mutant mice had a lower peak firing rate (Table 1). These differences across groups were also evident when cells were used as experimental units (Supplementary Table 3). Additionally, grid cells in mutant mice were less stable than in controls, but only when cells were used as experimental units (Supplementary Table 3). In mutant mice, the spatial information score, the number of fields and the peak firing rate of grid cells were negatively correlated with the percent infection in the RH region but not in the HP region (Supplementary Table 2). All other firing features of grid cells were similar across groups (Table 1and Supplementary Table 3). Overall, we conclude that mutant mice had fewer grid cells and that the remaining grid cells were less spatially selective, less stable, had a lower peak firing rate and tended to have fewer firing fields than those recorded in controls. Thus, ablation of NMDAR in the RH region disrupted the normal firing of grid cells.
a, Firing rate maps (top) and mean (± s.e.m.) grid and spatial information (inf.) scores (bottom) of grid cells for each genotype (one-way ANOVA followed by Tukey's multiple comparisons, grid score: F 2,27 = 0.07, P = 0.9; inf. score: F 2,27 = 5.0, P = 0.01; control vs. NR1 RH−/−: P = 0.02; control vs. NR1 RH&HP−/−: P = 0.005; NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.31; control: n = 7, NR1 RH−/−: n = 15, NR1 RH&HP−/−: n = 8). b, Firing rate maps (top) and mean (± s.e.m.) grid and spatial information (inf.) scores (bottom) of ISS cells for each genotype (one-way ANOVA followed by Tukey's multiple comparisons, grid score: F 2,27 = 4.94; P = 0.01; control vs. NR1 RH−/−: P = 0.03; control vs. NR1 RH&HP−/−: P = 0.005; NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.24; inf. score: F 2,27 = 0.7, P = 0.5; control: n = 7, NR1 RH−/−: n = 15, NR1 RH&HP−/−: n = 8). In a and b, numbers above each firing rate map correspond to the grid score (left, black) and peak firing rate (right, red). c, Firing rate maps (top) and mean (± s.e.m.) border and spatial information (inf.) scores (bottom) of border cells for each genotype. Numbers above to each firing map correspond to the border score (left, black) and peak firing rate (right, red; Kruskal–Wallis test or one-way ANOVA followed by multiple comparisons, border score: H = 10.09, P = 0.006; control vs. NR1 RH−/−: P = 0.99; control vs. NR1 RH&HP−/−: P = 0.046; NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.005; inf. score: F 2,35 = 4.88, P = 0.01; control vs. NR1 RH−/−: P = 0.65; control vs. NR1 RH&HP−/−: P = 0.04; NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.004; control: n = 8, NR1 RH−/−: n = 19, NR1 RH&HP−/−: n = 11). d, Head direction firing rate polar plots (top) and mean (± s.e.m.) HD and spatial information (inf.) scores (bottom) for each genotype. Numbers above each firing map correspond to the HD score (left, black) and peak firing rate (right, red; one-way ANOVA or Kruskal–Wallis test, HD score: F 2,31 = 1.74, P = 0.19; inf. score: H = 1.88, P = 0.39; control: n = 8, NR1 RH−/−: n = 17, NR1 RH&HP−/−: n = 9). Mice were used as experimental units. The color scale for firing rate maps is from 0 Hz (blue) to peak firing rate (red). All tests were two-sided. **P ≤ 0.01, *P < 0.05, ns: not significant after post hoc multiple comparisons. Yellow, red and blue symbols correspond to control, NR1 RH−/− and NR1 RH&HP−/− mice, respectively.
ISS cells constituted a heterogeneous group, including cells that almost reached the criteria that would classify them as grid cells (Fig. 3b). Like grid cells, ISS cells also tended to be less frequent in mutant mice, as they were found in 86% of control mice (7 of 8) and in 79–72% of mutant mice (15 of 19 in NR1 RH−/− mice and 8 of 11 in NR1 RH&HP−/− mice). In addition, ISS cells in mutant mice had significantly lower grid scores than those in control mice (Fig. 3b), while the other firing features remained similar across groups (Table 1). Comparable results were obtained when statistics were performed using cells as experimental units (Supplementary Table 3).
Border cells were found in all recorded mice and in similar proportions (Fig. 2c). The firing of border cells was affected only when the virus infection extended to the HP (NR1 RH&HP−/− mice), while it remained intact in NR1 RH−/− mice (Fig. 3c, Table 1, Supplementary Table 3 and Supplementary Fig. 5). Specifically, the border score and spatial information score of border cells were significantly lower in NR1 RH&HP−/− than in NR1 RH−/− and control mice (Fig. 3c, Table 1and Supplementary Table 3). These differences were no longer detected when more conservative thresholds were used (Supplementary Table 4). When the analysis was performed using cells as experimental units, we found that border cells in NR1 RH&HP−/− mice also showed a significantly lower head direction score and stability than those in the other two groups (Supplementary Table 3). Additionally, in mutant mice the border score and stability of border cells were inversely correlated with the percent infection in the HP region but not in the RH region (Supplementary Table 2). These correlations were absent in control mice (Supplementary Table 2). These results suggest that hippocampal NMDAR ablation had a detrimental effect on the firing properties of MEC border cells.
HD cells were also found in similar proportions across groups (Fig. 2c), and their firing properties were similar across groups (Fig. 3d, Table 1 and Supplementary Fig. 6). However, HD cells in NR1 RH&HP−/− mice showed a significantly higher head direction score than those in the other two groups of mice when cells were considered as experimental units (Supplementary Table 3). Accordingly, head direction selectivity of mutant mice was positively correlated to the percent infection in the HP region but not in the RH region (Supplementary Table 2). Finally, firing properties of SM cells and NSS cells did not differ across genotypes (Fig. 2c, Table 1 and Supplementary Table 3).
Since the MEC is one of the major cortical inputs to the HP and grid cells have been shown to project directly to the HP19, we also asked whether NMDAR ablation in the RH region affected the activity of hippocampal place cells. We bilaterally implanted six independently movable tetrodes in each HP of control and mutant mice and recorded the activity of place cells as the mice ran in a square arena. The firing rates of place cells were similar across groups (Table 1). The spatial information score and the stability of place cells remained intact in NR1 RH−/− mice and were reduced only when the virus infection extended to the HP (NR1 RH&HP−/− mice; Fig. 4, Table 1 and Supplementary Fig. 7). Similar differences across groups were observed when cells were used as experimental units (Supplementary Table 3). In addition, in mutant mice, the spatial information score and the stability of place cells were inversely correlated with the percent infection in the HP region but not in the RH region (Supplementary Table 2). Hence, place cell firing in NR1 RH−/− mice was indistinguishable from controls, while it was impaired in NR1 RH&HP−/− mice, which is in agreement with previous studies showing that normal place cell firing depends on hippocampal NMDAR signaling20,21,22.
a, Firing rate maps of representative place cells for each genotype. Numbers above each firing rate map correspond to the spatial information score (black, left) and the peak firing rate (red, right). The color scale for firing rate maps is from 0 Hz (blue) to peak firing rate (red). b, Mean (± s.e.m.) spatial information (inf.) score and stability of active HP cells (8 Hz > mean firing rate ≥ 0.25 Hz) for each genotype (Kruskal–Wallis test or one-way ANOVA followed by multiple comparisons, inf. score: H = 9.33, P = 0.009; control vs. NR1 RH−/−: P = 0.99; control vs. NR1 RH&HP−/−: P = 0.03; NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.01; stability: F 2,25 = 6.36, P = 0.006; control vs. NR1 RH−/−: P = 0.93; control vs. NR1 RH&HP−/−: P = 0.01; NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.01; control: n = 8, NR1 RH−/−: n = 13, NR1 RH&HP−/−: n = 7). Hemispheres were used as experimental unit (see Methods). All tests were two-sided. **P ≤ 0.01, *P < 0.05, ns: not significant after post hoc multiple comparisons. Yellow, red and blue symbols correspond to control, NR1 RH−/− and NR1 RH&HP−/− mice, respectively.
Altogether, our results demonstrate that NMDAR ablation in the RH region disrupted grid spatial periodicity and spatial selectivity of MEC principal neurons. This was reflected in a reduction in the number of grid cells and ISS cells, as well as in a disruption of their normal firing. At the same time, head direction selectivity, border selectivity and speed modulation of MEC principal neurons, as well as place cell activity in the HP, remained unaffected. Hence, NR1 RH−/− mice constitute a valid tool to test whether normal grid cell firing is necessary for PI computation.
Impaired path integration in NR1 RH−/− and NR1 RH&HP−/− mice
We examined the performance of control, NR1 RH−/− and NR1 RH&HP−/− mice in the L-maze assay3 (Fig. 5). In this assay, mice first swim in complete darkness and reach a submerged platform at the far end of a corridor during an initial ‘sample’ trial. Subsequently, during the ‘test’ trial, the corridor is removed and they must find the platform in the open tank in absence of external cues, relying only on PI information that they acquired during the sample trial. PI allows mice that swim along the corridor to build a global vector that, in its reversed form, enables them to quickly find the platform during a subsequent swim in the open tank, thereby minimizing the time spent in the water3. During the sample trials, mice were exposed to one of three corridors, referred to as Str. (straight), L-S (long–short) and S-L (short–long), in which the platform was located at 0°, 30° and 60° relative to the start box, respectively. Accordingly, the direction of the resulting global vector during the test trial should approach 0°, 30° and 60°, respectively (Fig. 5a). At the end of the experiment, each mouse swam between 8 and 10 times per condition, following a pseudorandom sequence. Calculation of individual mean global vectors from several test trials was performed as shown in Supplementary Fig. 8a. Performance did not improve with trial number (Supplementary Fig. 8b). In line with our previous study3, control mice clearly exhibited different global vector directions during Str., L-S and S-L corridor swims (Figs. 5b, c). This was not the case for NR1 RH−/− and NR1 RH&HP−/− mice, for which global vector directions in the Str. and L-S conditions overlapped (Figs. 5b,c). Vector directions across genotypes for each corridor configuration were also compared. In the Str. configuration, no differences were found across genotypes (Fig. 5c). Conversely, in the L-S and S-L configurations, vector directions differed across genotypes, i.e., the values were significantly smaller in mutant mice (Fig. 5c). The length of the resulting global vector within and across genotypes was also analyzed (Fig. 5d). Control mice showed similar vector length in all corridor configurations (Fig. 5d). In mutant mice, on the contrary, the vector length in the S-L configuration was shorter than in the Str. and L-S configurations (Fig. 5d). Comparison of the vector length across genotypes within each condition showed no difference for the Str. and L-S conditions (Fig. 5d). In contrast, in the S-L condition, the length of the global vector was shorter in mutants than in control mice (Fig. 5d). The differences across genotypes could not be accounted for by changes in swimming speed, since the mice from all groups swam at similar speeds during both sample and test trials in the three conditions (Fig. 5e). Therefore, the deficient performance of mutant mice in the L-S and S-L conditions (L-S: vector direction; S-L: vector direction and length) is consistent with a PI impairment (see also Supplementary Fig. 9).
a, Schematics of the L-maze assay consisting of one sample trial (corridor swim) and a test trial (open tank swim) with intertrial intervals of 15 min. The experiment was performed in complete darkness and compass information was provided only by the position of the start box. b, Polar plots depicting individual mean global vectors (blue arrows) and average global vectors (red arrows) for each genotype and condition. c, Average global vector direction (circular mean ± s.e.m.). Top: comparisons within genotypes (Watson–Williams F-tests, control: F 2,48 = 22.44, P = 1.32 × 10−7, Str. vs. L-S: P = 0.006, Str. vs. S-L: P = 8.07 × 10−8, L-S. vs. S-L: P = 0.0009; NR1 RH−/−: F 2,81 = 19.12, P = 1.58 × 10−7, Str. vs. L-S: P = 0.30, Str. vs. S-L: P = 8.35 × 10−8, L-S vs. S-L: P = 2.04 × 10−5; NR1 RH&HP−/−: F 2,81 = 16.28, P = 1.14 × 10−6, Str. vs. L-S: P = 0.86, Str. vs. S-L: P = 1.69 × 10−5, L-S vs. S-L: P = 1.6 × 10−7). Bottom: comparisons across genotypes (Watson–Williams F-tests, Str.: F 2,70 = 1.02, P = 0.37; L-S: F 2,70 = 4.21, P = 0.02, control vs. NR1 RH−/−: P = 0.01, control vs. NR1 RH&HP−/−: P = 0.009, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.82; S-L: F 2,70 = 3.54, P = 0.03, control vs. NR1 RH−/−: P = 0.02, control vs. NR1 RH&HP−/−: P = 0.02, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.67). d, Global vector length (mean ± s.e.m.) for each genotype and condition. The top and bottom panels show comparisons within and across genotypes, respectively (two-way repeated-measures ANOVA, interaction: F 4, 140 = 2.45, P = 0.04, condition effect: F 2, 140 = 2.22, P = 0.11, genotype effect: F 2,70 = 1.39, P = 0.25; top: Tukey's multiple comparisons, control: Str. vs. L-S: P = 0.88, Str. vs. S-L: P = 0.71, L-S vs. S-L: P = 0.44; NR1 RH−/−: Str. vs. L-S: P = 0.74, Str. vs. S-L: P = 0.03, L-S vs. S-L: P = 0.03; NR1 RH&HP−/−: Str. vs. L-S: P = 0.80, Str. vs. S-L: P = 0.01, L-S vs. S-L: P = 0.03; bottom: Tukey's multiple comparisons, Str.: control vs. NR1 RH−/−: P = 0.90, control vs. NR1 RH&HP−/−: P = 0.74, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.38; L-S: control vs. NR1 RH−/−: P = 0.89, control vs. NR1 RH&HP−/−: P = 0.72, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.93; S-L: control vs. NR1 RH−/−: P = 0.0007, control vs. NR1 RH&HP−/−: P = 0.009, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.59). e, Mean (± s.e.m.) swimming speed during sample (top) and test (bottom) trials of the L-maze assay (two-way repeated-measures ANOVA, sample trial: interaction: F 4, 140 = 0.18, P = 0.95, condition effect: F 2, 140 = 2.17, P = 0.08, genotype effect: F 2,70 = 0.43, P = 0.65; test trial: interaction: F 4, 140 = 0.94, P = 0.44, condition effect: F 2, 140 = 2.51, P = 0.08, genotype effect: F 2,70 = 0.64, P = 0.53). f, Mean (± s.e.m.) DOP score for each genotype (one-way ANOVA followed by Tukey's multiple comparisons, F 2,70 = 6.86, P = 0.002, control vs. NR1 RH−/−: P = 0.02, control vs. NR1 RH&HP−/−: P = 0.0004, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.12; control: n = 17, NR1 RH−/−: n = 28, NR1 RH&HP−/−: n = 28). Mice were used as experimental units. All tests were two-sided. *P < 0.05, **P ≤ 0.01, ***P ≤ 0.001, ns: not significant. White, light gray and dark gray bars correspond to the Str., L-S and S-L conditions, respectively. Yellow, red and blue bars correspond to control, NR1 RH−/− and NR1 RH&HP−/− mice, respectively.
To confirm the specificity of the behavioral impairment, mice were also tested on two additional tasks: a win–shift task on an elevated T-maze and a beaconing task in the water maze (Supplementary Figs. 10 and 11). The performances of control and NR1 RH−/− mice in the win–shift task were similar, while NR1 RH&HP−/− mice performed significantly worse (Supplementary Fig. 10b). The performance of mutant mice correlated with the percent infection in the HP region but not in the RH region (Supplementary Fig. 10c), which is in agreement with previous work showing that normal performance in this task depends on hippocampal NMDAR signaling23. In the beaconing task, mice swam in an illuminated open tank and reached a tagged platform located at 0°, 30° or 60° relative to the start box (Supplementary Fig. 11a). This control experiment served to test for sensorimotor, motivational or learning deficits that might account for the differential performance in the L-maze. Control, NR1 RH−/− and NR1 RH&HP−/− mice successfully solved this task, i.e., global vectors differed across conditions within each genotype but not across genotypes within each condition (Supplementary Fig. 11b,c). Thus, the performance of mutant mice in the beaconing task was indistinguishable from that of controls. Beaconing swims represented optimal vectors for each condition, and we used them to quantify the directional deviation from optimal performance in all corridor configurations of the L-maze assay. Accordingly, we calculated a single score called DOP (deviation from optimal performance), which provided a single value capturing the performance of each mouse across the three corridor configurations (see Methods for details). We found that the DOP scores were significantly higher in NR1 RH−/− and NR1 RH&HP−/− mice than in control mice (Fig. 5f), confirming that mutant mice were unable to compute accurate global vectors in the L-maze. In summary, the results of the three different behavioral tests show that NMDAR ablation in the RH region led to PI impairment and that such impairment was task-specific and did not result from sensorimotor, motivational or learning deficits. In addition, taken together, the results of the behavioral and electrophysiological experiments indicated that PI impairment correlates with disrupted grid cell activity.
To further strengthen our findings linking grid cell activity and PI performance, we took advantage of the heterogeneity of the mutant groups (variability in injection size) and correlated the DOP score with all electrophysiological variables. The DOP score was negatively correlated with the percent grid cells, the spatial information score of principal cells and the grid scores of principal cells, grid cells and ISS cells (Fig. 6 and Supplementary Table 5). No correlation was found between the DOP score and the remaining electrophysiological properties of MEC and HP neurons (Supplementary Table 5). Altogether, our results demonstrate that the severity of grid cell activity disruption correlated with the extent of PI impairment.
a, Relationship between DOP score and percentage of grid cells and NSS cells (Pearson correlation, percent grid cells: r = –0.42, P = 0.02; percent NSS cells: r = 0.29, P = 0.08; n = 23). b, Relationship between DOP score and the spatial information (inf.) score and grid score of MEC principal neurons (Pearson correlation, inf. score: r = –0.43, P = 0.02; grid score: r = –0.52, P = 0.005; n = 23). c, Relationship between DOP score and the spatial information (inf.) score and grid score of grid cells (Pearson correlation, inf. score: r = –0.21, P = 0.20; grid score: r = –0.47, P = 0.02; n = 18). d, Relationship between DOP score and the spatial information (inf.) score and grid score of ISS cells (Pearson correlation, inf. score: r = –0.29, P = 0.11; grid score: r = –0.41, P = 0.03; n = 20). All other correlations between DOP score and the remaining electrophysiological properties of MEC neurons were not significant (Supplementary Table 5). Mice were used as experimental units. All tests were two-sided. *P < 0.05, **P ≤ 0.01, ns: not significant.
Discussion
Here we demonstrate that NMDAR ablation in the RH region disrupts grid cell firing and PI performance. The link between the degrees of grid cell and PI impairment provides strong experimental support for the hypothesis that grid cell activity underlies PI navigation.
We found that NMDAR ablation in the RH region reduced spatial selectivity and grid spatial periodicity of MEC principal cells. The manipulation led to a strong reduction in the number of grid cells and ISS cells and also disrupted their normal firing, i.e., it reduced spatial selectivity, peak firing rate, stability and number of fields of grid cells, as well as the grid score of ISS cells. Yet the few grid cells recorded in mutant mice showed grid scores similar to those found in controls. This might be due to the fact that only cells with high grid scores were included in this group (ceiling effect). It could also be that grid cells with normal periodicity were recorded from MEC areas that were not infected. However, our method does not allow us to confirm this. The detrimental effect on grid cell activity did not reflect a global change in the MEC network state, since head direction selectivity, border selectivity, speed and theta modulation, as well as the firing properties of NSS cells and interneurons, remained unaltered. The role of NMDAR signaling in the HP has been addressed before, but its contribution to MEC neuronal activity has remained unknown. NMDAR ablation in the HP impairs long-term potentiation and depression in CA1 pyramidal cells24 and also disrupts place cell activity, leading to enlarged, diffuse and less stable place fields20,21,22. Here we confirm these earlier findings by showing that place cells in NR1 RH&HP−/− mice are less spatially selective and stable than in controls.
Disruption of grid cell activity was previously observed in rats in which the medial septum, HP or anterior thalamic nuclei were pharmacologically inactivated25,26,27,28, as well as in mice lacking GluA1-containing AMPA receptors (GluA1 −/− mice)3. Most of these studies3,25,26,27 also reported some degree of cell-type-specific functional dissociation within the MEC network, i.e., normal head direction cells and border cells in the absence of grid cell spatial periodicity, supporting the notion that distinct mechanisms underlie the generation of grid cell periodicity, head direction and border selectivity. One limitation of previous pharmacological and genetic approaches, however, is the lack of confinement to the grid cell network, thus precluding a direct test of the link between grid cell activity and PI. Here we overcame this limitation by ablating NMDARs in the RH region. The genetic manipulation disrupted grid periodicity while leaving intact the activity of the remaining MEC neurons and that of hippocampal cells. Our manipulation affected also other subregions within the RH region (subiculum, presubiculum, postsubiculum and parasubiculum) that contain spatially tuned cells8,11,12. We did not investigate the firing properties of these neurons. However, it is reasonable to assume that the effect of NMDAR ablation in these regions would be similar to the one found in the MEC, i.e., it is unlikely that the activity of spatially tuned cells other than grid cells is affected in these areas but not in the MEC upon virus infection. Although further experiments are required to confirm this assumption, NR1 RH−/− mice represent a powerful tool for testing the function of grid cells in general and the role of grid cell function in PI in particular.
MEC neurons in NR1 RH−/− and NR1 RH&HP−/− mice exhibited similar firing properties, except for border cells and HD cells. On one hand, border cells of NR1 RH&HP−/− mice had a lower border score and spatial selectivity (although this effect was no longer detectable when more conservative thresholds were applied). This finding points toward a putative contribution of the HP to the firing of MEC border cells. A previous study reported that HP inactivation did not affect MEC border cell firing, but the number of recorded border cells was small (n = 7)25. Further experiments would be necessary to determine whether or not normal HP activity is required for normal firing of MEC border cells. On the other hand, HD cells in NR1 RH&HP−/− mice showed higher head direction selectivity, which is in line with previous reports showing that HP inactivation results in an increase in head direction modulation of MEC neurons25.
Notably, NMDAR ablation in the RH region also spared the firing of place cells in the HP. This is in agreement with previous studies showing that grid cell spatial periodicity is not essential for normal place cell firing26,27,29 and also with the observation that place cells mature (acquire adult-like firing properties) earlier than grid cells30,31. Hence, our results do not support models proposing grid cells as the primary source of spatial information for the formation of place cells32. One alternative, however, is that residual spatial information of MEC neurons suffices for the generation of normal place fields in the HP.
In rats, grid cell activity emerges postnatally after eye opening and a short period of exploratory behavior30,31, suggesting that the system might not be hardwired and that its development might require synaptic plasticity33. We removed functional NMDARs in young mice before eye opening and exploratory experience, most likely before grid cells emerged. It has been shown that NMDARs are necessary for the development of modularly arranged neural circuits in the visual and somatosensory cortices34,35,36. Grid cells in the RH region are also organized in a discretized manner14,37, and our results indicate that proper development of this network depends on NMDAR signaling. Hence, our findings support the hypothesis that an NMDAR-dependent learning process is required for normal development and/or functioning of the grid cell network33. Our results also suggest that network interactions between grid cells are likely to be necessary to maintain their periodic firing. Previous experimental data indicate that such network interactions could work either via direct connections between grid cells in layer III38 or via a recurrent network of inhibitory interneurons present in layer II of the MEC39,40. It is noteworthy that NMDAR ablation, and most likely the developmental plasticity conferred by the receptor, affected grid cells specifically.
Irrespective of the underlying mechanism leading to grid cell firing disruption, NMDAR ablation in the RH region allowed us to investigate the link between grid cell firing and PI. We used the L-maze assay, which has several advantages compared to the commonly used food-carrying task4. First, the L-maze does not require previous training. Naïve mice swim proficiently and minimize the time in the water from the very first trial, which reduces interindividual variations arising from learning and motivation components. Second, due to the initial corridor swim, the outbound path is imposed by the experimenter instead of the animal. This allows experimenters to control the accumulation of errors inherent to PI, reduces the variability among animals and allows clear predictions about the direction and length of the expected global vectors. Last, the assay allows control of mouse-borne olfactory cues.
Our results clearly showed that mutant mice were unable to compute proper global vectors in the L-maze (they performed vectors of altered direction in the L-S condition and vectors of altered direction and length in the S-L condition). This was true even when the viral infection did not affect the complete RH region. Thus, NMDAR ablation in just a part of the RH network was enough to induce a significant PI impairment. Taken together, our data demonstrate that NMDAR ablation in the RH region disrupts both grid cell firing and PI performance. Notably, we were able to confirm the link between these two variables by using the same set of mice for behavioral and in vivo electrophysiological recordings. While our findings are, to our knowledge, the first to demonstrate an important role of grid cell firing in PI, they do not exclude the possibility that other, yet-unknown, cell types within and outside the RH region also contribute to this function41.
Our results are relevant for several reasons. First, they offer new insight into the function of NMDARs in the RH area, both at the cellular and behavioral level. Second, they support the idea that NMDAR-dependent plasticity is required for the normal development and/or functioning of the grid cell network. Third, they suggest that grid cell firing is not essential for the development or functioning of hippocampal place cells. Fourth, ever since the discovery of grid cells, theoretical considerations based on an overwhelming amount of neurophysiological and computational data have led to the hypothesis that these spatially selective neurons support PI. Here we provide experimental evidence supporting this hypothesis.
Methods
Animals
Experiments were performed using NR1 2lox/2lox mice (henceforth; NR1 flox mice)42 and wild-type littermates with a C57BL/6 background (64 males and 36 females; the male/female ratio was 1.63 for control, 1.85 for NR1 RH−/− and 1.83 for NR1 RH&HP−/− mice). Mice were kept on a 12-h light/dark schedule with all testing performed during the light phase. All experiments were approved by the Regierungspräsidium Karlsruhe in compliance with the European guidelines for the care and use of laboratory animals (licenses G-248/14 and G-50/14).
Virus-mediated NR1 gene ablation in young mice
Four-day-old (P4) NR1 flox mice received bilateral injections into MEC of AAV-Syn-GFP-T2A-Cre. In this vector, the green fluorescent protein (GFP) and the protein Cre-recombinase (Cre) are expressed under the control of the neuron-specific synapsin promoter (Syn). Anesthesia was induced and maintained with isoflurane (1–2.5%). We injected 1 µL of the viral construct in each MEC (±3 mm lateral from bregma, 0.1 mm anterior from the transverse sinus, at a 3° angle in the sagittal plane, with the tip pointing in the anterior direction). Following virus injection, Cre-recombinase excises the loxP-flanked exons 11–18 of the NR1 2lox/2lox alleles and transforms them to NR1 lox alleles, causing ablation of NR1 specifically in the area of injection. The control group included (i) wild-type littermate mice injected with AAV-Syn-GFP-T2A-Cre and (2) NR1 flox mice injected with AAV-Syn-Tomato (in which the synapsin promoter directs the expression of the fluorescent protein Tomato). Viral constructs were produced in-house. After virus injection, mice stayed with their mother until weaning (~4 weeks), at which time they were separated from the mother and divided by gender (4 mice per cage). Mice were housed in Plexiglas home cages (25 × 19 × 13 cm) with free access to food and water.
Electrophysiological recordings in brain slices
Whole-cell patch-clamp recordings were performed in slices obtained from a subgroup of NR1 RH−/− and wild-type mice 4–8 weeks after injection, (WT: n = 9 cells from 3 mice, WT-GFP+: n = 10 cells from 4 mice, NR1 RH−/−: n = 11 cells from 4 mice). The experimenter was not blind to the genotype of the mice. Mice were deeply anaesthetized with isoflurane, followed by transcardial perfusion with 30 mL sucrose solution containing (in mM) 212 sucrose, 26 NaHCO3, 1.25 NaH2PO4, 3 KCl, 7 MgCl2, 10 glucose and 0.2 CaCl2, cooled to 4 °C and oxygenated with carbogen gas (95% O2/5% CO2, pH 7.4). Acute sagittal sections (300 µm) containing the MEC were prepared using a vibratome (Slicer HR2, Sigmann Elektronik, Germany), and the tissue was incubated at room temperature for 1 h in oxygenated extracellular solution containing (in mM) 12.5 NaCl, 2.5 NaHCO3, 0.125 NaH2PO4, 0.25 KCl, 2 CaCl2, 1 MgCl2 and 25 glucose. For recording, individual slices were transferred into a submerged chamber superfused (perfusion rate = 3.0 mL/min) with oxygenated extracellular solution (heated to 30–32 °C). Fluorescently labeled (GFP+) cells in the MEC were visualized with epifluorescence and DIC optics. Recording pipettes, pulled from borosilicate capillaries with tip resistances of 5–8 MΩ, were filled with a Cs+-based solution, containing (in mM) 120 Cs+-gluconate, 10 CsCl, 10 HEPES, 0.2 EGTA, 8 NaCl, 10 phosphocreatine, 2 Mg-ATP and 0.3 GTP (pH 7.3 adjusted with CsOH). Liquid junction potentials were not corrected. Series resistance was maximally compensated and cells were discarded if it changed more than 20% or was higher than 40 MΩ. Evoked excitatory postsynaptic currents (eEPSCs) were recorded in voltage-clamp mode by placing the stimulation electrode (glass capillary, 3 MΩ and filled with extracellular solution) in layer I in the vicinity of the recorded cells (200–300 µm distance). AMPAR-mediated currents were recorded at a holding potential of –70 mV in presence of the GABAA receptor blocker gabazine (10 µm). NMDA receptor mediated currents were recorded at +40 mV in presence of the AMPA/kainate receptor blocker CNQX (10 µm) and gabazine. NMDAR-mediated currents were verified using d-AP5 (10 µM), a potent NMDAR blocker. To evaluate the ratio of NMDAR/AMPAR, the stimulus intensity and amplitude of AMPAR-mediated currents were kept constant (8–10% of 1 mA and 200–230 pA, respectively). For analysis of signal amplitudes, an average of 15 sweeps was used, and only monosynaptic inputs with latencies lower than 2 ms were considered. All recordings were obtained using HEKA PatchMaster EPC 10, and signals were filtered at 3 kHz and digitized at 20 kHz. Data were analyzed offline with HEKA software FitMaster. Statistics were performed using cells as experimental units
Behavioral tests
Behavioral experiments started between 11 and 20 weeks after virus injection. The experimenter was blind to the genotype of the mice. In all experiments, mice were first video-tracked at 25 frames per second and their movements subsequently analyzed using a position tracking system (Ethovision XT9, Noldus). After task completion, mice rested for at least 10 d before being exposed to a new task. Half of the mice were first tested in the L-maze assay and next in the win–shift task, while the other half was first tested in the win–shift task and next in the L-maze assay. The beaconing task was always performed after the L-maze assay.
L-maze assay for path integration
The test was carried out as previously reported3, with slight variations. The experiment was performed in a circular tank (120 cm in diameter and 60 cm tall) filled with water (21 ± 1 °C) to a depth of 22 cm. The water was made opaque by the addition of milk. An L-shaped maze was present inside the tank during sample trials (see below). The maze was made of Plexiglas and consisted of a corridor that had a long (82.5 × 10 × 35 cm) and a short segment (42.5 × 10 × 35 cm) joined at a 90° angle. Start and end boxes (13 × 10 × 35 cm) were attached at the end of each segment. The start box had a sliding door to confine the mouse at the beginning of each trial. The end box contained a circular platform (6.5-cm diameter) submerged 1 cm below the water surface. The entire experiment (including the pretraining phase, see below) was performed in darkness. The tank was illuminated with infrared light and a video camera with an infrared filter was located above its center. Two loudspeakers producing white noise were located below the tank to mask potential directional auditory cues. First, mice underwent a pretraining phase consisting of 9 trials. At the beginning of each trial, a mouse was placed inside the start box for 10 s before the sliding door was removed, and it would swim in the open tank to find one of three hidden platforms, whose distance from the start box increased gradually across trials. All mice successfully found one of the three platforms at the end of the pretraining phase. During this phase, mice learned the basic rules of the task, i.e., to swim and find a hidden platform in a dark pool to escape from the water. After this preparatory phase, the L-maze assay was performed. It consisted of one sample trial (corridor swim) and a test trial (open tank swim) with intertrial intervals of 15 min. At the beginning of the sample trial, the mouse was placed inside the start box (10 s before the sliding door was removed) and had to swim along the corridor to find the hidden platform located in the end box. During sample trials, the corridor could be arranged according to one of three configurations: (i) straight (Str.): the start box and the end box were both connected to opposite ends of the long segment; thus, mice had to swim in a straight line to find the platform at its end; (ii) long–short (L-S): the start box was connected to the long segment followed by the short segment; thus, mice had to swim along the long segment first, then turn 90° and swim along the shorter segment to find the end platform; and (iii) short–long (S-L): the start box was connected to the short segment followed by the long segment; in this case, mice had to swim first along the short segment, then turn 90° and swim along the long segment to find the platform. The position of the platform relative to the long axis of the start box was at 0°, 30° and 60° in the Str., L-S and S-L corridors, respectively. After the sample trial, the corridor was removed, leaving in place only the start box and the hidden platform. During the test trial, the mouse was placed again inside the start box, but this time it had to swim in the open tank to find the hidden platform. Both sample and test trials lasted until the mouse found the platform or 1 min elapsed. If the mouse failed to find the platform within 1 min, it was placed onto it by the experimenter. The mouse was left on the platform for 20 s before it was returned to its home cage. Mice performed 4–5 sample trials of a given condition each day, and this was repeated twice following a pseudorandom sequence. Test trials with a vector length shorter than 20 cm and/or a mean swimming speed lower than 5 cm/s were excluded from the analysis.
Beaconing task
The experiment was performed in the same circular tank filled with opaque water, but this time it was illuminated with visible light and the position of the platform was tagged with a flag that extended above the water surface. As before, at the beginning of each trial, mice were placed inside the start box and required to swim in the open tank to find the platform located at 0°, 30° or 60° relative to the long axis of the start box. The start box and the platform were moved around the tank in each trial. In this task, the flag attached to the platform was the only conspicuous cue reliably signaling the location of the goal. Each mouse swam 6–8 times per condition following a pseudorandom sequence.
Win–shift task
The experiment was performed in a T-maze (95 × 135 cm) illuminated with visible light and surrounded by conspicuous visual cues. Mice were maintained at 80–85% of their normal body weight with free access to water. Each trial consisted of a sample run and a choice run. On sample runs, the mouse was forced to turn either left or right by blocking access to one goal arm with a wooden block. Food reward was available at the end of the goal arm. The block was then removed, and the mouse was placed back at the beginning of the start arm to begin the choice run. During the choice run, the mouse was rewarded for choosing the previously unvisited arm. The delay between the sample run and the choice run was 10–15 s. Mice ran 10 daily trials (intertrial interval: ~30 min) for 7 d.
Data analysis
Circular statistics were used to compute individual mean global vectors from 5 to 8 swims performed by each mouse in each condition during the L-maze and beaconing assays. First, a trial vector (tv) connecting the start and the end of each trajectory was calculated. The end of the trajectory was defined as the position of the mouse when: (i) it touched the wall of the pool, (ii) its trajectory accumulated a turn >95° or (iii) it reached the platform. Next, for each mouse and condition, an individual mean global vector (igv) was calculated as the mean global vector weighted by the length of each tv. The individual mean global vectors (igv) were then used to calculate an average global vector (agv) for each condition and genotype. The direction and lengths of average global vectors were compared within and across genotypes.
The directions of the average global vectors obtained in the beaconing task (dagv beac.) were used to quantify the directional deviation from optimal performance in the L-maze. The DOP (deviation from optimal performance) score was calculated for each mouse as follows:
where digv Str , digv LS and digv SL are the directions of the individual mean global vector after Str., L-S and S-L corridor swims, respectively, and dagv beac.0°, dagv beac.30° and dagv beac.60° are the directions of the average global vector obtained in the beaconing task (for each genotype) when the platform was located at 0°, 30° and 60° relative to the start box, respectively.
In the win–shift task, the probability of alternating between the two arms of the maze (i.e., the number of trials with alternation divided by the total number of trials assayed) was calculated for each mouse.
In vivo electrophysiological recordings
After behavioral testing, mice underwent surgery for tetrode implantation. The experimenter was blind to the genotype of the mice. Surgery was performed under anesthesia (isoflurane 1–2.5%). For MEC recordings, a craniotomy above the right MEC (+3.1 mm lateral from bregma, 0.2 mm anterior from the transverse sinus) followed by implantation of a microdrive that allowed independent movement of five or six tetrodes. The tetrodes were lowered 0.8 mm below the brain surface. For HP recordings, the craniotomy was performed bilaterally above each HP (2 mm posterior and ±1.4 lateral from bregma). A microdrive with two tetrode bundles, each aiming at one hemisphere and containing six individually movable tetrodes, was implanted. The tetrodes were lowered 0.6–0.8 mm below the brain surface. Microdrives were fixed to the skull with dental acrylic. After surgery mice were singly housed. The tetrodes permitted recording unitary and LFP (local field potential) signals while the animal explored a square arena. After recovering from surgery (~1 week), mice were maintained at 80–85% of their normal body weight with free access to water, and trained during 2–4 weeks to forage inside the arena (70 × 70 × 30 cm). Tetrodes were lowered to the recording position during the training period and were considered to be in the MEC when large theta oscillations were observed. For hippocampal recordings, sharp wave-ripples were used to guide tetrodes into the pyramidal cell layer of the HP. The recording period started when mice readily explored the whole arena. A recording session consisted of two 20-min exploratory trials in the square arena and a 20-min rest trial in a rest box (23 × 23 × 50 cm, no pellets dispensed) in between both trials in the square arena. Mice were recorded once a day during a period of 3–6 weeks for MEC recordings and 1–2 weeks for HP recordings. After each recording session, tetrodes were lowered ~25–50 µm. Brain signals were recorded using dacqUSB (Axona) or RHD2000 (Intan Technologies) data acquisition systems and signals were sampled at 24 kHz and 20 kHz, respectively. The position of the animal in the arena was tracked using a camera attached to the ceiling and a custom-made tracking program that detected the position of three colored LEDs attached to the microdrive. Spike detection was performed offline using principal component analysis43. Spikes were subjected to automatic clustering (https://github.com/klusta-team/klustakwik) before being manually refined using a graphical cluster cutting program. Only clusters with stable features and a clear refractory period in their spike-time autocorrelation were analyzed. Separation between clusters was estimated by the isolation distance44. Positional information during arena exploration was synchronized with the brain signals, allowing analysis of the recorded neurons. Analysis of the firing properties of the recorded units was performed using custom-written C++ programs, shell scripts and R scripts. Only trials in which the mouse covered more than 80% of the arena and cells that fired at least 300 spikes in both trials were considered. For each recorded unit, a firing rate map was calculated. The recording environment was divided into 2 × 2-cm bins, the time spent in each bin was calculated, and the resulting occupancy map was smoothed with a Gaussian kernel (s.d. = 3 cm). The number of spikes in each bin was then divided by the corresponding bin of the occupancy map. The resulting firing rate maps were smoothed with the Gaussian kernel. Only data recorded when mice ran faster than 3 cm/s were considered. These firing rate maps were used to calculate different spatial properties of the recorded units.
Characterization of the recorded units:
-
1)
The spatial information score, used to quantify spatial selectivity of each cell45, was calculated as:
$${\rm{I}}=\sum _{i=1}^{N}{p}_{i}\frac{{\lambda }_{i}}{\lambda }{log}_{2}\frac{{\lambda }_{i}}{\lambda }$$where p i is the occupancy probability of bin i in the firing map, λ i is the firing rate in bin i, and λ is the mean firing rate of the neuron.
-
2)
Firing fields were defined as regions of the firing rate map containing at least one bin with a firing rate higher than 5 Hz and a minimum of 10 adjacent bins (40 cm2) with a firing rate higher than 20% of the highest firing rate in the field.
-
3)
The grid score was calculated using spatial autocorrelation matrices computed from the firing rate14,15. Each bin of a spatial autocorrelation matrix represented a Pearson correlation coefficient between all possible pairs of bins in a firing rate map with a given spatial lag. The spatial lag associated with a particular bin of the matrix was determined by the location of that bin relative to the matrix center. Pearson correlations at spatial lags for which < 20 pairs of firing rates were available were not considered. A peak in the autocorrelation matrix was defined as more than 10 adjacent bins with a correlation coefficient higher than a peak detection threshold, which was set to 0.1. To determine the periodicity of the spatial autocorrelation matrix, a circular region of the spatial autocorrelation matrix was defined containing up to six peaks but excluding the central peak. Pearson correlation coefficients (r) were calculated between that circular region of the matrix and a rotated version of it (by 30°, 60°, 90°, 120° and 150°). Then the grid score was calculated as:
$${\rm{Grid}}\,{\rm{score}}=\left(\frac{r6{0}^{\circ }+r12{0}^{\circ }}{2}\right)-\left(\frac{r3{0}^{\circ }+r9{0}^{\circ }+r15{0}^{\circ }}{3}\right)$$ -
4)
The border score was calculated as:
$${\rm{Border}}\,{\rm{score}}=\frac{CM-DM}{CM+DM}$$For each firing field (defined as groups of adjacent pixels with a firing rate larger than 30% of the peak firing rate of the map and covering at least 80 cm2) the proportion of the pixels along a wall that were also part of the field was calculated for the four walls separately. CM was defined as the maximum proportion obtained over all possible field–wall combinations. DM was the mean shortest distance to a wall for pixels that were part of a firing field, weighted by the firing rate in each pixel. DM was then normalized so that it ranged from 0 to 1. For each pixel in the map, the shortest distance to a wall was calculated. The largest value obtained over all map pixels was the value used for the normalization of DM.
-
5)
The head direction score was computed as the mean vector length of the polar histogram of the firing rate of a cell as a function of the head direction of the mouse (10°/bin).
-
6)
The speed score was obtained as the Pearson product–moment correlation between instantaneous firing rate of a neuron and running speed of the mouse13. The instantaneous firing rate was obtained by counting the number of spikes in 1-ms time windows. Next, the convolution between the spike count array and a Gaussian kernel (s.d. = 5 ms) was performed. The resulting vector was then integrated over 5-ms time windows and the values were multiplied by 200 (1,000/5) to transform the probability into Hz.
-
7)
The theta rhythmicity score was calculated using the power spectrum of the instantaneous firing rate of each unit as:
$${\rm{Theta}}\,{\rm{rhythmicity}}=\frac{{\rm{\theta }}-{\rm{\beta }}}{{\rm{\theta }}+{\rm{\beta }}}$$where θ is the mean power at theta frequency range (6–10 Hz) and β is the mean power within two frequency intervals (3–5 and 11–13 Hz). A score higher than zero indicates more power at theta frequency than for adjacent frequencies. For neurons with a theta rhythmicity score >0.5, the peak theta frequency was calculated as the frequency with the highest power within the frequency range between 6 and 10 Hz.
-
8)
The stability of the spatial firing patterns between the two open-field trials was calculated as the Pearson correlation coefficient (r) between firing rate maps from the two open-field trials in each session, and it ranged from –1 to +1.
For MEC recordings, recorded units with a mean firing rate ≥5 Hz were considered interneurons, whereas those with a mean firing rate <5 Hz were considered principal cells. Within principal cells, six mutually exclusive categories were defined: grid, border, head direction (HD), speed-modulated (SM), irregular spatially selective (ISS) and nonspatially selective (NSS) cells. Units with grid scores≥0.5 were considered grid cells. Units with border score ≥0.5 and grid score <0.5 were considered border cells. Units with head direction score ≥0.3 and grid and border scores <0.5 were considered HD cells. Units with speed score ≥0.1, head direction score <0.3 and grid and border scores <0.5 were considered SM cells. Units with information score ≥0.4, grid and border scores <0.5, head direction score <0.3 and speed score <0.1 were considered ISS cells. Units with information score <0.4, grid and border scores <0.5, head direction score <0.3 and speed score <0.1 were considered NSS cells. For hippocampal recordings, only pyramidal cells (≤8 Hz) that were active (mean firing rate ≥ 0.25 Hz) were considered for analysis. The spatial information score was calculated for the first recording trial, and map stability was calculated for pyramidal cells that reached the activity threshold during at least the first recording trial.
Score threshold values were obtained after shuffling procedures consisting of shifting the spike times of each cell by a minimum of 20 s before recalculating the corresponding scores. This was repeated 500 times for each unit, and thresholds were defined as the values associated with a probability lower than 1%. Decimals were rounded to tenths.
For MEC recordings, all scores were computed for each single unit and subsequently averaged within each mouse. Therefore, statistics were performed using mice as experimental units. In the case of HP recordings, all scores were computed for each single unit and subsequently averaged within each hemisphere, so statistics were performed using hemispheres as experimental units. This is because we performed bilateral recordings and injections varied between hemispheres within single animals.
Histology
After the electrophysiological recordings, mice were perfused transcardially with phosphate buffered saline, followed by 4% paraformaldehyde. Brains were removed and sliced sagittally in 50-µm sections on a vibratome (Leica VT1000S, Heidelberg, Germany). Free-floating sections were permeabilized in 0.2% Triton-PBS for 30 min, washed three times with PBS and incubated for 5 min with DAPI (4′,6-diamidino-2-phenylindole, 1:1,000). Sections were mounted in Mowiol. Visualization of viral spread and location of the tetrodes was performed using a confocal microscope (Zeiss LSM 700) and a slide scanner (Zeiss Axio Scan.Z1). The experimenter was blind to the genotype of the mice.
Quantification of brain area/volume infected by the virus and estimation of tetrode location
To estimate the boundaries and volume of single injections, we took pictures (5×) of each brain serial section (~100 sections per mouse) using a slide scanner (Zeiss Axio Scan.Z1). The images were subsequently analyzed using the software Zen 2lite (Zeiss Microscopy). In each brain section, we first defined two areas of interest: (i) the retro-hippocampal region (which includes the entorhinal area, subiculum, parasubiculum, presubiculum and postsubiculum) and (ii) the hippocampal region (which includes Ammon's horn and the dentate gyrus). Next, we measured the area covered by the GFP signal (i.e., area of viral expression) within each region. All regions were manually defined. The percentage of infection (GFP expression) inside each region was calculated as:
where rh i and hp i are the areas expressing GFP in the RH and HP regions of the brain section i, respectively, and RH i and HP i are the areas covered by the RH and HP regions in the brain section i, respectively. The percent infection in RH and HP regions was calculated per hemisphere and per mouse for the electrophysiological and behavioral experiments, respectively. To calculate the percent infection per mouse the following formulas were used:
A threshold value of 3% was arbitrarily set to assign hemispheres/mice to the experimental groups. Hemispheres/mice with less than 3% infection in the HP region were included in the NR1 RH−/− group. Hemispheres/mice with more than 3% infection in the HP region were included in the NR1 RH&HP−/− group. This quantification is a coarse measure that is invariant to superinfection (since fluorescence level does not add to its magnitude). Quantifying the actual fraction of infected neurons would increase the exactness of our estimate. However, this would not improve the subsequent analysis as inaccuracies in estimation of viral infection are unlikely to be greater than interindividual variations.
Tetrode location was estimated using the same brain sections and software. The dorsoventral position of the tetrode tips was defined as the distance between the dorsal border of the MEC and the tetrode tip. The mediolateral position was obtained from a stereotaxic atlas (Allen Brain Atlas). Missimplanted animals were excluded from the analysis. The experimenter was blind to the genotype of the mice.
Statistical analysis
No statistical methods were used to predetermine the sample size, but our sample sizes are similar to (or larger than) those generally employed in the field. Statistical analysis was performed using Prism 6 (GraphPad Software) and Oriana 4.02 (Kovach Computing). Data were tested for normality and homoscedasticity using the KS normality test and the Brown–Forsythe test, respectively. We used t tests, one-way ANOVA with Tukey's multiple comparison tests (with Bonferroni adjustments for multiple comparisons) and two-way repeated-measures ANOVA when data passed normality and homoscedasticity assumptions. Kruskal–Wallis tests and Dunn's multiple comparisons tests (with Bonferroni adjustment for multiple comparisons) were used when this was not the case. Watson–Williams F-tests (with Bonferroni adjustments for multiple comparisons) for circular data were used to compare the directions of global vectors. Pearson correlations were used to analyze the relationship between variables. All tests were two-sided. Unless otherwise specified, mice were used as experimental units. During the experiments, we used 73 NR1 flox mice injected with AAV-Cre-GFP. Of these, four mice were used for whole-cell patch-clamp recordings, and the remaining 69 mice for behavioral tests and in vivo electrophysiology (NR1 RH−/− n = 37, NR1 RH&HP−/− n = 33). In addition, 27 control mice were injected (19 WT mice injected with AAV-Cre-GFF and 8 NR1 flox mice injected with AAV-Syn-Tomato). Of the control mice, 4 (plus 3 noninjected WT mice) were used for whole-cell patch-clamp recordings, and the remaining 24 mice were used for behavioral tests and in vivo electrophysiology. Some mice were used in either the behavioral tests or electrophysiological recordings, while others were used in both types of experiments. L-maze assay: control: n = 17, NR1 RH−/−: n = 28, NR1 RH&HP−/−: n = 28. Beaconing task: control: n = 13, NR1 RH−/−: n = 11, NR1 RH&HP−/−: n = 10 (all these mice also performed the L-maze assay). Win–shift task: control: n = 14, NR1 RH−/−: n = 19, NR1 RH&HP−/−: n = 20 (all these mice also performed the L-maze assay). MEC electrophysiological recordings: control: n = 8, NR1 RH−/−: n = 19, NR1 RH&HP−/−: n = 11 (some of these mice also took part in the L-maze assay: control: n = 4, NR1 RH−/−: n = 13, NR1 RH&HP−/−: n = 10). HP electrophysiological recordings: control: n = 8 hemispheres from 4 mice, NR1 RH−/−: n = 13 hemispheres from 9 mice, NR1 RH&HP−/−: n = 7 hemispheres from 6 mice (some of these mice also took part in the L-maze assay: control: n = 3, NR1 RH−/−: n = 4, NR1 RH&HP−/−: n = 5).
Life sciences reporting summary
Further information on experimental design is available in the Life Sciences Reporting Summary.
Data and code availability
Codes for characterization of the in vivo recorded units can be provided by the corresponding author upon reasonable request. The data that support the findings of this study are available from the corresponding author on reasonable request.
References
Mittelstaedt, M. L. & Mittelstaedt, H. Homing by path integration in a mammal. Naturwissenschaften. 67, 566–567 (1980).
Wehner, R. Desert ant navigation: how miniature brains solve complex tasks. J. Comp. Physiol. A. Neuroethol. Sens. Neural. Behav. Physiol. 189, 579–588 (2003).
Allen, K. et al. Impaired path integration and grid cell spatial periodicity in mice lacking GluA1-containing AMPA receptors. J. Neurosci. 34, 6245–6259 (2014).
Maaswinkel, H., Jarrard, L. E. & Whishaw, I. Q. Hippocampectomized rats are impaired in homing by path integration. Hippocampus. 9, 553–561 (1999).
Parron, C. & Save, E. Evidence for entorhinal and parietal cortices involvement in path integration in the rat. Exp. Brain. Res. 159, 349–359 (2004).
Van Cauter, T. et al. Distinct roles of medial and lateral entorhinal cortex in spatial cognition. Cereb. Cortex. 23, 451–459 (2013).
Blackstad, T. W. Commissural connections of the hippocampal region in the rat, with special reference to their mode of termination. J. Comp. Neurol. 105, 417–537 (1956).
Taube, J. S., Muller, R. U. & Ranck, J. B. Jr. Head-direction cells recorded from the postsubiculum in freely moving rats. I. Description and quantitative analysis. J. Neurosci. 10, 420–435 (1990).
Fyhn, M., Molden, S., Witter, M. P., Moser, E. I. & Moser, M.-B. Spatial representation in the entorhinal cortex. Science 305, 1258–1264 (2004).
Solstad, T., Boccara, C. N., Kropff, E., Moser, M.-B. & Moser, E. I. Representation of geometric borders in the entorhinal cortex. Science 322, 1865–1868 (2008).
Lever, C., Burton, S., Jeewajee, A., O'Keefe, J. & Burgess, N. Boundary vector cells in the subiculum of the hippocampal formation. J. Neurosci. 29, 9771–9777 (2009).
Boccara, C. N. et al. Grid cells in pre- and parasubiculum. Nat. Neurosci. 13, 987–994 (2010).
Kropff, E., Carmichael, J. E., Moser, M.-B. & Moser, E. I. Speed cells in the medial entorhinal cortex. Nature 523, 419–424 (2015).
Hafting, T., Fyhn, M., Molden, S., Moser, M.-B. & Moser, E. I. Microstructure of a spatial map in the entorhinal cortex. Nature 436, 801–806 (2005).
Sargolini, F. et al. Conjunctive representation of position, direction, and velocity in entorhinal cortex. Science 312, 758–762 (2006).
McNaughton, B. L., Battaglia, F. P., Jensen, O., Moser, E. I. & Moser, M.-B. Path integration and the neural basis of the ‘cognitive map’. Nat. Rev. Neurosci. 7, 663–678 (2006).
Moser, E. I., Kropff, E. & Moser, M.-B. Place cells, grid cells, and the brain's spatial representation system. Annu. Rev. Neurosci. 31, 69–89 (2008).
Monyer, H. et al. Heteromeric NMDA receptors: molecular and functional distinction of subtypes. Science 256, 1217–1221 (1992).
Zhang, S. J. et al. Optogenetic dissection of entorhinal-hippocampal functional connectivity. Science 340, 1232627 (2013).
McHugh, T. J., Blum, K. I., Tsien, J. Z., Tonegawa, S. & Wilson, M. A. Impaired hippocampal representation of space in CA1-specific NMDAR1 knockout mice. Cell 87, 1339–1349 (1996).
Nakazawa, K. et al. Hippocampal CA3 NMDA receptors are crucial for memory acquisition of one-time experience. Neuron. 38, 305–315 (2003).
Cabral, H. O., Fouquet, C., Rondi-Reig, L., Pennartz, C. M. & Battaglia, F. P. Single-trial properties of place cells in control and CA1 NMDA receptor subunit 1-KO mice. J. Neurosci. 34, 15861–15869 (2014).
McHugh, S. B., Niewoehner, B., Rawlins, J. N. & Bannerman, D. M. Dorsal hippocampal N-methyl-D-aspartate receptors underlie spatial working memory performance during non-matching to place testing on the T-maze. Behav. Brain. Res. 186, 41–47 (2008).
Tsien, J. Z., Huerta, P. T. & Tonegawa, S. The essential role of hippocampal CA1 NMDA receptor-dependent synaptic plasticity in spatial memory. Cell 87, 1327–1338 (1996).
Bonnevie, T. et al. Grid cells require excitatory drive from the hippocampus. Nat. Neurosci. 16, 309–317 (2013).
Koenig, J., Linder, A. N., Leutgeb, J. K. & Leutgeb, S. The spatial periodicity of grid cells is not sustained during reduced theta oscillations. Science 332, 592–595 (2011).
Brandon, M. P. et al. Reduction of theta rhythm dissociates grid cell spatial periodicity from directional tuning. Science 332, 595–599 (2011).
Winter, S. S., Clark, B. J. & Taube, J. S. Spatial navigation. Disruption of the head direction cell network impairs the parahippocampal grid cell signal. Science 347, 870–874 (2015).
Miao, C. et al. Hippocampal remapping after partial inactivation of the medial entorhinal cortex. Neuron. 88, 590–603 (2015).
Langston, R. F. et al. Development of the spatial representation system in the rat. Science 328, 1576–1580 (2010).
Wills, T. J., Cacucci, F., Burgess, N. & O'Keefe, J. Development of the hippocampal cognitive map in preweanling rats. Science 328, 1573–1576 (2010).
Bush, D., Barry, C. & Burgess, N. What do grid cells contribute to place cell firing? Trends. Neurosci. 37, 136–145 (2014).
Widloski, J. & Fiete, I. R. A model of grid cell development through spatial exploration and spike time-dependent plasticity. Neuron. 83, 481–495 (2014).
Simon, D. K., Prusky, G. T., O'Leary, D. D. & Constantine-Paton, M. N-methyl-D-aspartate receptor antagonists disrupt the formation of a mammalian neural map. Proc Natl Acad Sci USA 89, 10593–10597 (1992).
Iwasato, T. et al. NMDA receptor-dependent refinement of somatotopic maps. Neuron. 19, 1201–1210 (1997).
Iwasato, T. et al. Cortex-restricted disruption of NMDAR1 impairs neuronal patterns in the barrel cortex. Nature 406, 726–731 (2000).
Stensola, H. et al. The entorhinal grid map is discretized. Nature 492, 72–78 (2012).
Dhillon, A. & Jones, R. S. Laminar differences in recurrent excitatory transmission in the rat entorhinal cortex in vitro. Neuroscience. 99, 413–422 (2000).
Couey, J. J. et al. Recurrent inhibitory circuitry as a mechanism for grid formation. Nat. Neurosci. 16, 318–324 (2013).
Pastoll, H., Solanka, L., van Rossum, M. C. & Nolan, M. F. Feedback inhibition enables nested oscillations and grid firing fields. Neuron. 77, 141–154 (2013).
Valerio, S. & Taube, J. S. Path integration: how the head direction signal maintains and corrects spatial orientation. Nat. Neurosci. 15, 1445–1453 (2012).
Niewoehner, B. et al. Impaired spatial working memory but spared spatial reference memory following functional loss of NMDA receptors in the dentate gyrus. Eur. J. Neurosci. 25, 837–846 (2007).
Csicsvari, J., Hirase, H., Czurkó, A., Mamiya, A. & Buzsáki, G. Oscillatory coupling of hippocampal pyramidal cells and interneurons in the behaving Rat. J. Neurosci. 19, 274–287 (1999).
Schmitzer-Torbert, N., Jackson, J., Henze, D., Harris, K. & Redish, A. D. Quantitative measures of cluster quality for use in extracellular recordings. Neuroscience. 131, 1–11 (2005).
Skaggs, W. E., McNaughton, B. L., Wilson, M. A. & Barnes, C. A. Theta phase precession in hippocampal neuronal populations and the compression of temporal sequences. Hippocampus. 6, 149–172 (1996).
Acknowledgements
We thank E. Savenkova, U. Amtmann, L. Hartmann, B. Schwalm and K. Neumeier for their help during the experiments. This work was funded through grants from the German Research Foundation (DFG, GI 1157/1-1) to M.G., the European Research Council (250047), the German Ministry of Education and Research (BMBF) (Grant 01GQ1003A) and the German Research Foundation (SFB 1134) to H.M., a Marie Sklodowska Curie Fellowship (H2020-MSCA-IF-708295) to M.I.S. and the Emmy Noether Programme of the DFG (AL 1730/1-1) to K.A.
Author information
Authors and Affiliations
Contributions
H.M. conceived the study; R.J.D.M. designed the L-maze assay, beaconing assay, behavioral data analysis and viral infection quantification; M.G. performed viral injections, tetrode implantations, behavioral assays, MEC in vivo electrophysiological recordings, histology, brain imaging and data analysis; M.A. performed MEC in vivo electrophysiological recordings and data analysis; M.I.S. performed HP in vivo electrophysiological recordings, the corresponding tetrode implantations and data processing; A.N. performed whole-cell patch-clamp recordings; K.A. developed software for in vivo electrophysiological data analysis; M.G. and H.M. wrote the paper with contributions from all coauthors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated Supplementary Information
Supplementary Figure 1 Extent of virus infection in NR1 RH–/– and NR1 RH&HP–/– mice.
a, Percentage of virus infection (% infection, mean ±s.e.m.) in the RH and HP regions of NR1 RH−/− and NR1 RH&HP−/− mice. NR1 RH&HP−/− mice showed higher % infection than NR1 RH−/− mice in both areas (t-test, HP: t = 8.27, P = 0.0001; RH: t = 4.65, P = 0.0001, NR1 RH−/−: n = 23, NR1 RH&HP−/−: n = 21). b, Percentage of virus infection (mean ±s.e.m.) in the RH sub-regions, i.e., entorhinal cortex (EC), subiculum (Sub), parasubiculum (ParaS), pre- and postsubiculum (Pre/PostS) in mutant mice. NR1 RH&HP−/− mice showed higher % infection than NR1 RH−/− mice only in the EC region (t-test, EC: t = 4.44, P = 0.0001; Sub: t = 0.27, P = 0.78; ParaS: t = 0.38; P = 0.70; Pre/PostS: t = 0.51, P = 0.61, NR1 RH−/−: n = 23, NR1 RH&HP−/−: n = 21). c, Correlation between the % infection in the whole RH region and the individual RH sub-regions (EC, Sub, ParaS and Pre/PostS) in mutant mice (Pearson correlation, NR1 RH−/− EC: r = 0.96, P = 0.0001, NR1 RH&HP−/− EC: r = 0.74, P = 0.0001, NR1 RH−/− Sub: r = 0.69, P = 0.0002, NR1 RH&HP−/− Sub: r = 0.4, P = 0.83, NR1 RH−/− ParaS: r = 0.41, P = 0.06, NR1 RH&HP−/− ParaS: r = 0.24, P = 0.39, NR1 RH−/− Pre/PostS: r = 0.43, P = 0.06, NR1 RH&HP−/− Pre/PostS: r = 0.13, P = 0.61, NR1 RH−/−: n = 23, NR1 RH&HP−/−: n = 21). d, Days post injection (dpi, mean ±s.e.m.)) for each genotype. No difference in dpi was found across groups (One-way ANOVA, F(2,62) = 2.55, P = 0.09, control: n = 17, NR1 RH−/−: n = 26, NR1 RH&HP−/−: n = 22). e, Correlations between dpi and % infection in the RH and HP regions in mutant mice. No correlations were found between these variables (Pearson correlation, RH: r = 0.18, P = 0.20; HP: r = -0.07, P = 0.64; n = 48). Likewise no correlation between dpi and % infection was found in control mice (data not shown, Pearson correlation, RH: r = -0.32, P = 0.4; HP: r = -0.59, P = 0.09; n = 9). The lack of correlation indicated that long-term infections did not result in a significant decrement of infected cells. f, Confocal image showing a long-term viral infection (275 dpi) in the RH region of an NR1 lox mouse. All tests were two-sided. ***P ≤ 0.001, P, ns: not significant.
Supplementary Figure 2 Tetrode location, isolation distance and running speed are similar across groups.
a, Confocal images showing tetrode tracks in the MEC of control, NR1 RH−/−and NR1 RH&HP−/− mice. Red circles represent the tip of the tetrode; white dashed lines delineate the MEC boundaries. Green: GFP, blue; DAPI. Scale bar, 200 µm. b, Dorso-ventral (d-v) and medio-lateral (m-l) coordinates of the tetrode tips for each genotype (upper panel: raw data, lower panel: mean ±s.e.m.). One-way ANOVA, d-v: F(2,171) = 0.66, P = 0.52; m-l: F(2,171) = 1.22, P = 0.29, control: n = 39 tetrodes in 8 mice, NR1 RH−/−: n = 92 tetrodes in 19 mice, NR1 RH&HP−/−: n = 43 tetrodes in 11 mice. c, Distribution of tetrode tips across MEC layers for each genotype. The tip of most tetrodes was located in layers I-II of the dorsal MEC. d, Isolation distance (mean ±s.e.m.) of the units recorded in each genotype. One-way ANOVA, F(2,35) = 1.91, P = 0.16. e Running speed (mean ±s.e.m.) in the recording arena for each genotype. Kruskal-Wallis test: H = 0.59, P = 0.75. In d and e, control: n = 8 mice, NR1 RH−/− : n = 19 mice, NR1 RH&HP−/−: n = 11 mice. All tests were two-sided. ns: not significant.
Supplementary Figure 3 Preserved theta rhythmicity in mutant mice.
a, Spike-time autocorrelation for principal cells and interneurons ordered from the bottom to the top according to their theta rhythmicity index for the each genotype. Red and dark blue were assigned to the highest and lowest value, respectively, of each individual autocorrelation. b, Spike-time autocorrelation (mean ±s.e.m.) for principal cells (control: n = 1581, NR1 RH−/− : n = 1881, NR1 RH&HP−/−: n = 900) and interneurons (control: n = 777, NR1 RH−/− : n = 941, NR1 RH&HP−/−: n = 524) in each genotype. Theta rhythmic activity was still present in mutant mice. c, Mean (±s.e.m.) power spectrum of spike-time autocorrelations for principal cells (control: n = 1581, NR1 RH−/− : n = 1881, NR1 RH&HP−/−: n = 900) and interneurons (control: n = 777, NR1 RH−/− : n = 941, NR1 RH&HP−/−: n = 524) in each genotype. In all groups theta was the dominant rhythmic frequency. d, Theta rhythmicity for principal cells and interneurons in each genotype. Theta rhythmicity tended to be higher in NR1 RH&HP−/− mice (Kruskal-Wallis test followed by Dunn’s comparisons, principal cells: H = 14.49, P = 0.0007, control vs. NR1 RH−/−: P = 0.18, control vs. NR1 RH&HP−/−: P = 0.07, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.0004, control: n = 1581, NR1 RH−/− : n = 1881, NR1 RH&HP−/−: n = 900; interneurons: H = 19.39, P = 0.0001, control vs. NR1 RH−/−: P = 0.23, control vs. NR1 RH&HP−/−: P = 0.02, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.0001, control: n = 777, NR1 RH−/− : n = 941, NR1 RH&HP−/−: n = 524). e, Peak theta frequency for principal cells and interneurons (with a theta rhythmicity > 0.5) in each genotype. 0The peak frequency in mutants was lower than in control mice (Kruskal-Wallis test followed by Dunn’s comparisons, principal cells: H = 291.4, P = 0.0001, control vs. NR1 RH−/−: P = 0.0001, control vs. NR1 RH&HP−/−: p = 0.0001, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.03, control: n = 867, NR1 RH−/− : n = 731, NR1 RH&HP−/−: n = 464; interneurons: H = 59.15, P = 0.0001, control vs. NR1 RH−/−: P = 0.0001, control vs. NR1 RH&HP−/−: P = 0.0001, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.72, Control: n = 394, NR1 RH−/− : n = 836, NR1 RH&HP−/−: n = 282). Cells were used as experimental units. All tests were two-sided. ***P ≤ 0.001, **P ≤ 0.01, *P < 0.05.
Supplementary Figure 4 Examples of grid cells in control, NR1 RH–/– and NR1 RH&HP–/– mice.
Trajectories (black lines) with spike positions (red dots) and firing rate maps of grid cells with the highest grid score (top row of each set of images) and the lowest grid score -if available- (bottom row of each set of images) found in each mouse during the first and the second recording trial. The peak firing rate is displayed above each firing rate map. The color scale for firing rate maps is from 0 Hz to peak firing rate.
Supplementary Figure 5 Examples of border cells in control, NR1 RH–/– and NR1 RH&HP–/– mice.
Trajectories (black lines) with spike positions (red dots) and firing rate maps of border cells with the highest border score (top row of each set of images) and the lowest border score (bottom row of each set of images) found in each mouse during the first and the second recording trial. The peak firing rate is displayed above each firing rate map. The color scale for firing rate maps is from 0 Hz to peak firing rate.
Supplementary Figure 6 Examples of HD cells in control, NR1 RH–/– and NR1 RH&HP–/– mice.
Trajectories (black lines) with spike positions (red dots) and head-direction rate polar plots of HD cells with highest head direction score (top row of each set of images) and lowest head direction score (bottom row of each set of images) found in each mouse during the first and the second recording trial. The peak firing rate is displayed above each polar plot.
Supplementary Figure 7 Examples of place cells in control, NR1 RH–/– and NR1 RH&HP–/– mice.
For each hemisphere with more than three active (mean firing rate ≥ 0.25 Hz) pyramidal cells, the mean spatial information score was calculated and two representative cells (if available) with a spatial information score comparable to the corresponding mean are shown. For each cell, trajectories (black lines) with spike positions (red dots) and firing rate maps are depicted for the first and second recording trial. The color scale for firing rate maps is from 0 Hz to peak firing rate.
Supplementary Figure 8 L-maze performance does not improve across test trials.
a, Calculation of individual mean global vectors from several test trials following for the different L-maze conditions (Str., L-S and S-L). First, a trial vector connecting the start and the end of each trajectory was calculated (blue arrows). Next, the mean vector weighted by the length of each trial vector was computed (red arrows). The direction and length of the individual global vectors are depicted below each polar plot. The individual mean global vectors were subsequently used to calculate an average global vector for each genotype and condition. b, Global vector direction (circular mean ±s.e.m.) during the first 5 trials for each condition for each genotype (mice performed 4-5 sample trials of a given condition on a given day, and this was repeated twice following a pseudorandom sequence). Dashed lines indicate theoretical global vector directions. Performance did not improve across trials (One-way ANOVA: control: Str., F(4,71) = 0.87, P = 0.49, L-S, F(4,65) = 0.31, P = 0.89, S-L, F(4,61) = 0.52, P = 0.72; NR1 RH−/−: Str., F(4,96) = 0.39, P = 0.81, L-S, F(4,87) = 1.19, P = 0.32, S-L, F(4,73) = 0.50, P = 0.73; NR1 RH&HP−/−: Str., F(4,117) = 0.72, P = 0.58, L-S, F(4,112) = 0.02, P = 0.99, S-L, F(4,109) = 0.56, P = 0.69). Trials with a mean swimming speed lower than 5 cm/s and/or a vector length shorter than 20 cm were excluded from the analysis (hence, repeated measures analysis could not be performed). Similar results were found for the second 5 trials for each condition. These results are in agreement with the notion that path integration is an innate strategy, and hence the establishment and use of global vectors are invariant to learning. Mice were used as experimental units. Control: n = 17, NR1 RH−/−: n = 28, NR1 RH&HP−/−: n = 28. ns: not significant.
Supplementary Figure 9 NR1 RH–/– and NR1 RH&HP–/– mice also show impaired path integration when mice (n = 3) with low % infection in the RH region (< 3%) are included in the control group Impaired performance of mutant mice can be observed in both the L-S and S-L conditions (L-S: vector direction, S-L: vector direction (although marginally) and length) even if the variability of the control group is increased by using data from low GFP-expressing mice.
a, Average global vector direction (circular mean ±s.e.m.). The upper panel show comparisons within genotypes (Watson-Williams F-tests, Control: F(2,57) = 28.54, P = 2.58 × 10-9, Str. vs. L-S: P = 7.57 × 10-4, Str. vs. S-L: P = 2.35 × 10-9, L-S. vs. S-L; P = 4.85 × 10-4;NR1 RH−/−: F(2,72) = 16.05, P = 1.72 × 10-6, Str. vs. L-S: P = 0.62, Str. vs. S-L: P = 2.02 × 10-6, L-S vs. S-L: P = 2.99 × 10-5; NR1 RH&HP−/−: F(2,81) = 16.28, P = 1.14 × 10-6, Str. vs. L-S: P = 0.86, Str. vs. S-L: P = 1.69 × 10-5, L-S vs. S-L: P = 1.6 × 10-7). The lower panel show comparisons across genotypes (Watson-Williams F-tests, Str.: F(2,70) = 0.81, P = 0.45; L-S: F(2,70) = 5.42, P = 0.005, control vs. NR1 RH−/−: P = 0.004, control vs. NR1 RH&HP−/−: P = 0.005, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.57; S-L: F(2,70) = 2.95: P = 0.058, control vs. NR1 RH−/−: P = 0.04, control vs. NR1 RH&HP−/−: P = 0.04, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.63). b, Global vector length (mean ±s.e.m.) for each genotype and condition. The upper and lower panels show comparisons within and across genotypes, respectively (Two-ways ANOVA with repeated measures, interaction F(4,140) = 2.75, P = 0.03, condition effect F(2,140) = 3.14, P = 0.04, genotype effect F(2,70) = 1.15, P = 0.36; upper panel: Tukey’s multiple comparisons, Control: Str. vs. L-S: P = 0.49, Str. vs. S-L: P = 0.99, L-S vs. S-L: P = 0.41; NR1 RH−/−: Str. vs. L-S: P = 0.61, Str. vs. S-L: P = 0.03, L-S vs. S-L: P = 0.03; NR1 RH&HP−/−−/−: Str. vs. L-S, P = 0.8, Str. vs. S-L, P = 0.01, L-S vs. S-L, P = 0.03; lower panel: Tukey’s multiple comparisons, Str.: control vs. NR1 RH−/−: P = 0.41, control vs. NR1 RH&HP−/−: P = 0.94, NR1 RH−/− vs. NR1 RH&HP−/−, P = 0.19; L-S: control vs. NR1 RH−/−, P = 0.39, control vs. NR1 RH&HP−/−, P = 0.74, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.79; S-L: control vs. NR1 RH−/−: P = 0.008, control vs. NR1 RH&HP−/−: P = 0.04, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.74). Control: n = 20, NR1 RH−/−: n = 25, NR1 RH&HP−/−: n = 28. Mice were used as experimental units. All tests were two-sided. ***P ≤ 0.001, **P ≤ 0.01, *P < 0.05, ns: not significant.
Supplementary Figure 10 NR1 RH&HP–/– but not NR1 RH–/– mice are impaired in the win–shift task.
a, Schematics of the win-shift task on a T-maze (rewarded alternation). The task consisted of a sample and a choice trial. During the sample run, one arm was blocked and the other contained food reward at its end. During the choice run, the mouse was rewarded for choosing the previously unvisited arm. b, Probability to alternate (mean ±s.e.m.) during choice trials for each genotype. One-way ANOVA followed by Tukey’s multiple comparisons, F(2,50) = 12.12, P = 0.0001, control vs. NR1 RH−/−: P = 0.48, control vs. NR1 RH&HP−/−: P = 0.0001, NR1 RH−/− vs. NR1 RH&HP−/−: P = 0.001. Control: n = 14, NR1 RH−/− : n = 19, NR1 RH&HP−/−: n = 20. c, Relationship between the probability to alternate and % infection in the RH and HP regions in mutant mice. Pearson correlation, RH: r = -0.29, P = 0.08, HP: r = -0.51, P = 0.001, n = 39. Mice were used as experimental units. All tests were two-sided. **P ≤ 0.01, ns: not significant.
Supplementary Figure 11 NR1 RH–/– and NR1 RH&HP–/– mice show normal performance in a beaconing task.
a, Schematics of the beaconing task. Each mouse swam 6-8 times per condition, i.e., 0°, 30° and 60°, following a pseudorandom sequence. b, Polar plots depicting individual mean global vectors (blue arrows) and average global vectors (red arrows) for each genotype and condition. c Mean global vector direction (circular mean ±s.e.m.) for each genotype and condition. The three groups of mice exhibited significantly different global vector directions during 0°, 30° and 60° trials (Watson-Williams F-test followed by pairwise comparisons, control: F(2,36) = 118.61, P = 1 × 10-12, 0° vs. 30°: P = 4.49 × 10-7, 0° vs. 60°: P = 1 × 10-12, 30° vs. 60°: P = 6.16 × 10-9; NR1 RH−/−: F(2,30) = 84.35, P = 1 × 10-12, 0° vs. 30°: P = 1.1 × 10-5, 0° vs. 60°: P = 2.26 × 10-10, 30° vs. 60°: P = 3.14 × 10-8; NR1 RH&HP−/−: F(2,27) = 169.86, P = 1 × 10-12, 0° vs. 30°: P = 1.52 × 10-5, 0° vs. 60°: P = 3.2 × 10-11, 30° vs. 60°: P = 1.35 × 10-10). For each condition, vector direction was similar across genotypes (Watson-Williams F-test, 0°: F(2,31) = 1.74, P = 0.19, 30°: F(2,31) = 0.67, P = 0.52; 60°: F(2,31) = 0.89, P = 0.42). d, Mean global vector length (mean ± SEM) for each genotype and condition. No differences were found in vector length within and across genotypes (Two-ways ANOVA with repeated measures, genotype effect: F(2,31) = 1.75, P = 0.19, condition effect: F(2,62) = 2.14, P = 0.13, interaction: F(4,62) = 0.67, P = 0.61). Control: n = 13, NR1 RH−/−: n = 11, NR1 RH&HP−/−: n = 10. Mice were used as experimental units. All tests were two-sided. ***P ≤ 0.001; ns: not significant.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Gil, M., Ancau, M., Schlesiger, M.I. et al. Impaired path integration in mice with disrupted grid cell firing. Nat Neurosci 21, 81–91 (2018). https://doi.org/10.1038/s41593-017-0039-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-017-0039-3
This article is cited by
-
Hippocampal firing fields anchored to a moving object predict homing direction during path-integration-based behavior
Nature Communications (2023)
-
Grid cell modeling with mapping representation of self-motion for path integration
Neural Computing and Applications (2023)
-
Place cells dynamically refine grid cell activities to reduce error accumulation during path integration in a continuous attractor model
Scientific Reports (2022)
-
Disruption of the grid cell network in a mouse model of early Alzheimer’s disease
Nature Communications (2022)
-
The genie in the bottle-magnified calcium signaling in dorsolateral prefrontal cortex
Molecular Psychiatry (2021)