Abstract
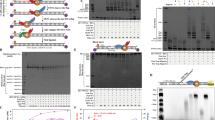
DNA replication introduces thousands of RNA primers into the lagging strand that need to be removed for replication to be completed. In Escherichia coli when the replicative DNA polymerase Pol IIIα terminates at a previously synthesized RNA primer, DNA Pol I takes over and continues DNA synthesis while displacing the downstream RNA primer. The displaced primer is subsequently excised by an endonuclease, followed by the sealing of the nick by a DNA ligase. Yet how the sequential actions of Pol IIIα, Pol I polymerase, Pol I endonuclease and DNA ligase are coordinated is poorly defined. Here we show that each enzymatic activity prepares the DNA substrate for the next activity, creating an efficient four-point molecular handover. The cryogenic-electron microscopy structure of Pol I bound to a DNA substrate with both an upstream and downstream primer reveals how it displaces the primer in a manner analogous to the monomeric helicases. Moreover, we find that in addition to its flap-directed nuclease activity, the endonuclease domain of Pol I also specifically cuts at the RNA–DNA junction, thus marking the end of the RNA primer and creating a 5′ end that is a suitable substrate for the ligase activity of LigA once all RNA has been removed.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Cryo-EM maps and atomic models have been deposited in the Electron Microscopy Database and Protein Data Bank, respectively, under accession codes EMD-17033, PDB 8OOY, EMD-17005 and PDB 8OO6. Source data are provided with this paper. Other requests should be addressed to M.H.L. (m.h.lamers@lumc.nl).
References
Mok, M. & Marians, K. J. The Escherichia coli preprimosome and DNA B helicase can form replication forks that move at the same rate. J. Biol. Chem. 262, 16644–16654 (1987).
McInerney, P., Johnson, A., Katz, F. & O’Donnell, M. Characterization of a triple DNA polymerase replisome. Mol. Cell 27, 527–538 (2007).
Yao, N. Y., Georgescu, R. E., Finkelstein, J. & O’Donnell, M. E. Single-molecule analysis reveals that the lagging strand increases replisome processivity but slows replication fork progression. Proc. Natl Acad. Sci. USA 106, 13236–13241 (2009).
Sakabe, K. & Okazaki, R. A unique property of the replicating region of chromosomal DNA. Biochim. Biophys. Acta. 129, 651–654 (1966).
Okazaki, R. Mechanism of DNA chain growth. J. Mol. Biol. 95, 63–70 (1967).
Blattner, F. R. et al. The complete genome sequence of Escherichia coli K-12. Science 277, 1453–1462 (1997).
Kitani, T., Yoda, K. Y., Ogawa, T. & Okazaki, T. Evidence that discontinuous DNA replication in Escherichia coli is primed by approximately 10 to 12 residues of RNA starting with a purine. J. Mol. Biol. 184, 45–52 (1985).
Wu, C. A., Zechner, E. L., Reems, J. A., McHenry, C. S. & Marians, K. J. Coordinated leading- and lagging-strand synthesis at the Escherichia coli DNA replication fork. V. Primase action regulates the cycle of Okazaki fragment synthesis. J. Biol. Chem. 267, 4074–4083 (1992).
Balakrishnan, L. & Bambara, R. A. Okazaki fragment metabolism. Cold Spring Harb. Perspect. Biol. 5, a010173 (2013).
LaDuca, R. J., Crute, J. J., McHenry, C. S. & Bambara, R. A. The β subunit of the Escherichia coli DNA polymerase III holoenzyme interacts functionally with the catalytic core in the absence of other subunits. J. Biol. Chem. 261, 7550–7557 (1986).
Stukenberg, P. T., Studwell-Vaughan, P. S. & O’Donnell, M. Mechanism of the sliding β-clamp of DNA polymerase III holoenzyme. J. Biol. Chem. 266, 11328–11334 (1991).
Georgescu, R. E. et al. Mechanism of polymerase collision release from sliding clamps on the lagging strand. EMBO J. 28, 2981–2991 (2009).
Lundquist, R. C. & Olivera, B. M. Transient generation of displaced single-stranded DNA during nick translation. Cell 31, 53–60 (1982).
Singh, K., Srivastava, A., Patel, S. S. & Modak, M. J. Participation of the fingers subdomain of Escherichia coli DNA polymerase I in the strand displacement synthesis of DNA. J. Biol. Chem. 282, 10594–10604 (2007).
Klett, R. P., Cerami, A. & Reich, E. Exonuclease VI, a new nuclease activity associated with E. coli DNA polymerase. Proc. Natl Acad. Sci. USA 60, 943–950 (1968).
Kelly, R. B. Enzymatic synthesis of deoxyribonucleic acid. J. Biol. Chem. 53, 83–112 (1970).
Lyamichev, V., Brow, M. A. D. & Dahlberg, J. E. Structure-specific endonucleolytic cleavage of nucleic acids by eubacterial DNA polymerases. Science 260, 778–783 (1993).
Xu, Y. et al. Biochemical and mutational studies of the 5′-3′ exonuclease of DNA polymerase I of Escherichia coli. J. Mol. Biol. 268, 284–302 (1997).
Sugimoto, K., Okazaki, T. & Okazaki, R. Mechanism of DNA chain growth, II. Accumulation of newly synthesized short chains in E. coli infected with ligase-defective T4 phages. Proc. Natl Acad. Sci. USA 60, 1356–1362 (1968).
Pauling, C. & Hamm, L. Properties of a temperature-sensitive, radiation-sensitive mutant of Escherichia coli. II. DNA replication. Proc. Natl Acad. Sci. USA 64, 1195–1202 (1969).
Gao, Y. & Yang, W. Different mechanisms for translocation by monomeric and hexameric helicases. Curr. Opin. Struct. Biol. 61, 25–32 (2020).
Meir, A. & Greene, E. C. Srs2 and pif1 as model systems for understanding sf1a and sf1b helicase structure and function. Genes (Basel) 12, 1319 (2021).
Yuan, Q. & McHenry, C. S. Strand displacement by DNA polymerase III occurs through a τ-ψ-χ link to single-stranded DNA-binding protein coating the lagging strand template. J. Biol. Chem. 284, 31672–31679 (2009).
Schwartz, J. J. & Quake, S. R. Single molecule measurement of the ‘speed limit’ of DNA polymerase. Proc. Natl Acad. Sci. USA 106, 20294–20299 (2009).
O’Donnell, M. E. & Kornberg, A. Complete replication of templates by Escherichia coli DNA polymerase III holoenzyme. J. Biol. Chem. 260, 12884–12889 (1985).
Leu, F. P., Georgescu, R. & O’Donnell, M. Mechanism of the E. coli τ processivity switch during lagging-strand synthesis. Mol. Cell 11, 315–327 (2003).
Pascal, J. M. DNA and RNA ligases: structural variations and shared mechanisms. Curr. Opin. Struct. Biol. 18, 96–105 (2008).
Johnson, K. A. The kinetic and chemical mechanism of high-fidelity DNA polymerases. Biochim. Biophys. Acta - Proteins Proteom. 1804, 1041–1048 (2010).
Santoso, Y. et al. Conformational transitions in DNA polymerase I revealed by single-molecule FRET. Proc. Natl Acad. Sci. USA 107, 715–720 (2010).
Miller, B. R., Beese, L. S., Parish, C. A. & Wu, E. Y. The closing mechanism of DNA polymerase I at atomic resolution. Structure 23, 1609–1620 (2015).
Beese, L. S., Derbyshire, V. & Steitz, T. A. Structure of DNA polymerase I Klenow fragment bound to duplex DNA. Science https://doi.org/10.1142/9789811215865_0028 (1993).
Eom, S. H., Wang, J. & Steitz, T. A. Structure of Taq polymerase with DNA at the polymerase active site. Nature 382, 293–296 (1996).
Kiefer, J. R., Mao, C. & Beese, L. S. Visualizing DNA replication in a catalytically active Bacillus DNA polymerase crystal. Nature 391, 304–307 (1998).
Ghosh, S., Goldgur, Y. & Shuman, S. Mycobacterial DNA polymerase I: activities and crystal structures of the POL domain as apoenzyme and in complex with a DNA primer-template and of the full-length FEN/EXO–POL enzyme. Nucleic Acids Res. 48, 3165–3180 (2020).
Craggs, T. D. et al. Substrate conformational dynamics facilitate structure-specific recognition of gapped DNA by DNA polymerase. Nucleic Acids Res. 47, 10788–10800 (2019).
Kim, Y. et al. Crystal structure of Thermus aquaticus DNA polymerase. Nature 376, 288–292 (1995).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
AlMalki, F. A. et al. Direct observation of DNA threading in flap endonuclease complexes. Nat. Struct. Mol. Biol. 23, 640–646 (2016).
Tsutakawa, S. E. et al. Phosphate steering by Flap Endonuclease 1 promotes 5′-flap specificity and incision to prevent genome instability. Nat. Commun. 8, 15855 (2017).
Xu, Y., Grindley, N. D. F. & Joyce, C. M. Coordination between the polymerase and 5′-nuclease components of DNA polymerase I of Escherichia coli. J. Biol. Chem. 275, 20949–20955 (2000).
Pauszek, R. F. III, Lamichhane, R., Singh, A. R. & Millar, D. P. Single-molecule view of coordination in a multi-functional DNA polymerase. eLife 10, e62046 (2021).
Ohtani, N., Tomita, M. & Itaya, M. Junction ribonuclease: a ribonuclease HII orthologue from Thermus thermophilus HB8 prefers the RNA-DNA junction to the RNA/DNA heteroduplex. Biochem. J. 412, 517–526 (2008).
Rychlik, M. P. et al. Crystal structures of RNase H2 in complex with nucleic acid reveal the mechanism of RNA-DNA junction recognition and cleavage. Mol. Cell 40, 658–670 (2010).
Sriskanda, V. A second NAD+-dependent DNA ligase (LigB) in Escherichia coli. Nucleic Acids Res. 29, 4930–4934 (2001).
Olivera, B. M. Enzymic joining of polynucleotides. J. Mol. Biol. 46, 481–492 (1968).
Stukenberg, P. T., Turner, J. & O’Donnell, M. An explanation for lagging strand replication: polymerase hopping among DNA sliding clamps. Cell 78, 877–887 (1994).
López De Saro, F. J., Georgescu, R. E., Goodman, M. F. & O’Donnell, M. Competitive processivity-clamp usage by DNA polymerases during DNA replication and repair. EMBO J. 22, 6408–6418 (2003).
López de Saro, F. J. & O’Donnell, M. Interaction of the β sliding clamp with MutS, ligase, and DNA polymerase I. Proc. Natl Acad. Sci. USA 98, 8376–8380 (2001).
Bhardwaj, A., Ghose, D., Gopal Thakur, K. & Dutta, D. Escherichia coli β-clamp slows down DNA polymerase I dependent nick translation while accelerating ligation. PLoS ONE 13, e0199559 (2018).
Kukshal, V. et al. M. tuberculosis sliding β-clamp does not interact directly with the NAD+-dependent DNA ligase. PLoS ONE 7, e35702 (2012).
Levin, D. S., Bai, W., Yao, N., O’Donnell, M. & Tomkinson, A. E. An interaction between DNA ligase I and proliferating cell nuclear antigen: implications for Okazaki fragment synthesis and joining. Proc. Natl Acad. Sci. USA 94, 12863–12868 (1997).
Zhang, P. et al. Direct interaction of proliferating cell nuclear antigen with the p125 catalytic subunit of mammalian DNA polymerase δ. J. Biol. Chem. 274, 26647–26653 (1999).
Gomes, X. V. & Burgers, P. M. J. Two modes of FEN1 binding to PCNA regulated by DNA. EMBO J. 19, 3811–3821 (2000).
Xie, B. et al. Reconstitution and characterization of the human DNA polymerase delta four-subunit holoenzyme. Biochemistry 41, 13133–13142 (2002).
Georgescu, R. E. et al. Structure of a sliding clamp on DNA. Cell 132, 43–54 (2008).
Fernandez-Leiro, R., Conrad, J., Scheres, S. H. W. & Lamers, M. H. cryo-EM structures of the E. coli replicative DNA polymerase reveal its dynamic interactions with the DNA sliding clamp, exonuclease and τ. eLife https://doi.org/10.7554/eLife.11134 (2015).
Fernandez-Leiro, R. et al. Self-correcting mismatches during high-fidelity DNA replication. Nat. Struct. Mol. Biol. 24, 140–143 (2017).
Toste Rêgo, A., Holding, A. N., Kent, H. & Lamers, M. H. Architecture of the Pol III-clamp-exonuclease complex reveals key roles of the exonuclease subunit in processive DNA synthesis and repair. EMBO J. 32, 1334–1343 (2013).
Garg, P., Stith, C. M., Sabouri, N., Johansson, E. & Burgers, P. M. Idling by DNA polymerase δ maintains a ligatable nick during lagging-strand DNA replication. Genes Dev. 18, 2764–2773 (2004).
Rossi, M. L., Purohit, V., Brandt, P. D. & Bambara, R. A. Lagging strand replication proteins in genome stability and DNA repair. Chem. Rev. 106, 453–473 (2006).
Stodola, J. L. & Burgers, P. M. Resolving individual steps of Okazaki-fragment maturation at a millisecond timescale. Nat. Struct. Mol. Biol. 23, 402–408 (2016).
Ganai, R. A. & Johansson, E. DNA replication—a matter of fidelity. Mol. Cell 62, 745–755 (2016).
Jin, Y. H. et al. The 3′→5′ exonuclease of DNA polymerase δ can substitute for the 5′ flap endonuclease Rad27/Fen1 in processing Okazaki fragments and preventing genome instability. Proc. Natl Acad. Sci. USA 98, 5122–5127 (2001).
Grasby, J. A., Finger, L. D., Tsutakawa, S. E., Atack, J. M. & Tainer, J. A. Unpairing and gating: sequence-independent substrate recognition by FEN superfamily nucleases. Trends Biochem. Sci. 37, 74–84 (2012).
Stith, C. M., Sterling, J., Resnick, M. A., Gordenin, D. A. & Burgers, P. M. Flexibility of eukaryotic Okazaki fragment maturation through regulated strand displacement synthesis. J. Biol. Chem. 283, 34129–34140 (2008).
Matsumoto, Y., Brooks, R. C., Sverzhinsky, A., Pascal, J. M. & Tomkinson, A. E. Dynamic DNA-bound PCNA complexes co-ordinate Okazaki fragment synthesis, processing and ligation. J. Mol. Biol. 432, 166698 (2020).
Notomi, T. et al. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 28, E63 (2000).
Thompson, D. & Lei, Y. Mini review: recent progress in RT-LAMP enabled COVID-19 detection. Sens. Actuators Rep. 2, 100017 (2020).
Kashir, J. & Yaqinuddin, A. Loop mediated isothermal amplification (LAMP) assays as a rapid diagnostic for COVID-19. Med. Hypotheses 141, 109786 (2020).
Mautner, L. et al. Rapid point-of-care detection of SARS-CoV-2 using reverse transcription loop-mediated isothermal amplification (RT-LAMP). Virol. J. 17, 160 (2020).
Mackay, I. M., Arden, K. E. & Nitsche, A. Real-time PCR in virology. Nucleic Acids Res. 30, 1292–1305 (2002).
Rabe, B. A. & Cepko, C. SARS-CoV-2 detection using isothermal amplification and a rapid, inexpensive protocol for sample inactivation and purification. Proc. Natl Acad. Sci. USA 117, 24450–24458 (2020).
Luna-Vargas, M. P. A. et al. Enabling high-throughput ligation-independent cloning and protein expression for the family of ubiquitin specific proteases. J. Struct. Biol. 175, 113–119 (2011).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Venkata Subramaniya, S. R. M., Terashi, G. & Kihara, D. Super resolution cryo-EM maps with 3D deep generative networks. Biophys. J. 120, 283a (2021).
Zi Tan, Y. et al. Addressing preferred specimen orientation in single-particle cryo-EM through tilting. Nat. Methods 14, 793–796 (2017).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011).
Nicholls, R. A., Tykac, M., Kovalevskiy, O. & Murshudov, G. N. Current approaches for the fitting and refinement of atomic models into cryo-em maps using CCP-EM. Acta Crystallogr. D 74, 492–505 (2018).
Nicholls, R. A., Fischer, M., McNicholas, S. & Murshudov, G. N. Conformation-independent structural comparison of macromolecules with ProSMART. Acta Crystallogr. D 70, 2487–2499 (2014).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Dalrymple, B. P., Kongsuwan, K., Wijffels, G., Dixon, N. E. & Jennings, P. A. A universal protein-protein interaction motif in the eubacterial DNA replication and repair systems. Proc. Natl Acad. Sci. USA 98, 11627–11632 (2001).
de Castro, E. et al. ScanProsite: detection of PROSITE signature matches and ProRule-associated functional and structural residues in proteins. Nucleic Acids Res. 34, W362–W365 (2006).
Chaiet, L. & Wolf, F. J. The properties of streptavidin, a biotin-binding protein produced by Streptomycetes. Arch. Biochem. Biophys. 106, 1–5 (1964).
Acknowledgements
We thank the staff of the LUMC EM facility and the Netherlands Center for Electron Nanoscopy (NeCEN) for help with data collection and data processing. This work has been supported by an LUMC Research Fellowship to M.H.L. Access to NeCEN was supported by the Netherlands Electron Microscopy Infrastructure, project no. 184.034.014 of the National Roadmap for Large-Scale Research Infrastructure of the Dutch Research Council (NWO). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
M.H.L. and M.M.B. conceived the overall experimental design. M.M.B. prepared samples, performed biochemical assays and analyzed data. A.B. collected and processed cryo-EM data. M.M.B. and M.H.L. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Nicholas Dixon, Mike O’Donnell and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Sara Osman and Dimitris Typas were the primary editors on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cryo-EM data collection and data processing details.
a, Representative micrograph from the 11,000 micrographs collected. b, 2D class averages from full dataset. c, Fourier Shell Correlation between half-maps from final refinement. Green line: unmasked. Blue line: masked. Red line: phase randomized. Black line: corrected. Bottom (4th) panel shows the Fourier Shell Correlation of the model-to-map of the three structures (green line: full-length structure, blue line: closed structure, red line: open structure), d, Schematic representation of main data processing procedures. See Methods section for more details. e, Final map obtained after applying SuperEM code to Relion postprocessed map from dataset 1 and colored by local resolution. Color bar below the map shows the resolution range of the cryo-EM map in Å. f, Detail of model fitted to the final cryo-EM map from dataset 1. g, Orientational distribution of final set of refined particles from dataset 1. h, Anisotropy analysis of open and closed structure. Red line shows the half map FSC, with green lines representing the spread of the directional resolution defined by plus and minus one standard deviation from the mean. Blue bars show the histogram of one hundred directional resolutions evenly sampled of the 3D FSC. i, overlay of cryo-EM pol I structure with five X-ray crystallography structures of Pol I (pdb codes: 1l3t, 2bdp, 3tar, 6dsy, 7k50). Cryo-EM structure of open Pol I in cyan, with template DNA in black, extended primer in green, and displaced primer in red. All previously determined structures are shown in gray.
Extended Data Fig. 2 Positional modelling of the endonuclease domain in Pol I.
a, E. coli Pol I with predicted position of the endonuclease domain on top of the fingers domain, based on the position of the single-stranded flap (see also main Fig. 3a). b, Thermus aquaticus Pol I with its endonuclease domain adjacent to the 3′-5′ exonuclease domain33. c, Mycobacterium smegmatis Pol I with its endonuclease domain adjacent to the thumb domain31. d, Superposition of three Pol I endonuclease domains from T. aquaticus, M. smegmatis, and E. coli, (modelled using AlphaFold34). Red star marks the endonuclease active site. e, Superposition of E. coli Pol I endonuclease domain and T5 endonuclease bound to its substrate DNA35. The square marks the enlarged area shown below. Site of incision is marked by the two nucleotides in light blue. Two catalytic aspartates are shown in sticks. f, Similar comparison with human FEN136. Structures of T5 endonuclease and FEN1 were used to model the position of Pol I endonuclease in main Fig. 3b.
Extended Data Fig. 3 Pol I does not progress past a C in the template strand in absence of dGTP.
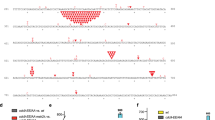
a, Pol I activity in absence of LigA using the templates shown in panel b, using only the three nucleotides dATP, dCTP, and dTTP. Experimental conditions are the same as in Fig. 5b. b, Substrates used for experiment in panel a.
Extended Data Fig. 4 The location of β-binding motifs of Pol I, LigA and Pol IIIα.
a, Model of Pol I and β-clamp. The β-clamp is shown in green surface, with binding pocket highlighted in yellow. The predicted β-binding motif of Pol I is shown in magenta sticks and is located in a helix of the thumb domain that interacts with the minor groove of the DNA. b, Model of LigA and β-clamp shown in two views rotated by 180°. The two predicted β-binding motifs are shown in magenta sticks. Motif LigA-1 is located in a helix that precedes a loop that interacts with the DNA. Motif LigA-2 is located on a strand that is part of the OB-domain that also interacts with the DNA. c, Cryo-EM structure of Pol IIIα bound to the β-clamp, exonuclease ε, and DNA52. The Pol IIIα β-binding motif is shown in magenta sticks and located on a loop at the end of the fingers domain and interacts with the binding pocket of the β-clamp. d, Alignment of β-binding motifs from Pol I, LigA and Pol IIIα that are highlighted in magenta in panels a-c.
Supplementary information
Supplementary Information
Supplementary Tables 1–3.
Supplementary Video 1
Video showing a morph from closed to open cryo-EM map of Pol I.
Supplementary Video 2
Model of the single base pair translocation of the DNA in Pol I and the resulting displacement of a single nucleotide in the displaced strand.
Source data
Source Data Fig. 1
Unprocessed gels.
Source Data Fig. 3
Unprocessed gels.
Source Data Fig. 4
Unprocessed gels.
Source Data Fig. 5
Unprocessed gels.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Unprocessed gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Botto, M.M., Borsellini, A. & Lamers, M.H. A four-point molecular handover during Okazaki maturation. Nat Struct Mol Biol 30, 1505–1515 (2023). https://doi.org/10.1038/s41594-023-01071-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-01071-y