Abstract
The cancer-specific fusion oncoprotein SS18–SSX1 disturbs chromatin accessibility by hijacking the BAF complex from the promoters and enhancers to the Polycomb-repressed chromatin regions. This process relies on the selective recognition of H2AK119Ub nucleosomes by synovial sarcoma X breakpoint 1 (SSX1). However, the mechanism underlying the selective recognition of H2AK119Ub nucleosomes by SSX1 in the absence of ubiquitin (Ub)-binding capacity remains unknown. Here we report the cryo-EM structure of SSX1 bound to H2AK119Ub nucleosomes at 3.1-Å resolution. Combined in vitro biochemical and cellular assays revealed that the Ub recognition by SSX1 is unique and depends on a cryptic basic groove formed by H3 and the Ub motif on the H2AK119 site. Moreover, this unorthodox binding mode of SSX1 induces DNA unwrapping at the entry/exit sites. Together, our results describe a unique mode of site-specific ubiquitinated nucleosome recognition that underlies the specific hijacking of the BAF complex to Polycomb regions by SS18–SSX1 in synovial sarcoma.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMD, www.ebi.ac.uk/pdbe/emdb/) under the accession codes EMD-34954 (SSX1–H2AK119Ub NCP complex, 1:1), EMD-34956 (SSX1–unmodified NCP complex), EMD-34957 (H2AK119Ub NCP) and EMD-36747 (SX1-H2AK119Ub NCP complex, 2:1). The atomic models have been deposited in the Protein Data Bank (PDB, www.rcsb.org) under the accession codes 8HQY (SSX1–H2AK119Ub NCP complex) and 8HR1 (SSX1–unmodified NCP complex). All ChIP–seq data have been deposited in the GEO repository under the GEO accession number GSE236811. Source data are provided with this paper.
References
Klemm, S. L., Shipony, Z. & Greenleaf, W. J. Chromatin accessibility and the regulatory epigenome. Nat. Rev. Genet. 20, 207–220 (2019).
Kadoch, C. et al. Dynamics of BAF-Polycomb complex opposition on heterochromatin in normal and oncogenic states. Nat. Genet. 49, 213–222 (2017).
Clapier, C. R., Iwasa, J., Cairns, B. R. & Peterson, C. L. Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat. Rev. Mol. Cell Biol. 18, 407–422 (2017).
Kadoch, C. & Crabtree, G. R. Mammalian SWI/SNF chromatin remodeling complexes and cancer: mechanistic insights gained from human genomics. Sci. Adv. 1, e1500447 (2015).
Wang, H. et al. Role of histone H2A ubiquitination in Polycomb silencing. Nature 431, 873–878 (2004).
Kalb, R. et al. Histone H2A monoubiquitination promotes histone H3 methylation in Polycomb repression. Nat. Struct. Mol. Biol. 21, 569–571 (2014).
Cooper, S. et al. Jarid2 binds mono-ubiquitylated H2A lysine 119 to mediate crosstalk between Polycomb complexes PRC1 and PRC2. Nat. Commun. 7, 13661 (2016).
Tamburri, S. et al. Histone H2AK119 mono-ubiquitination is essential for Polycomb-mediated transcriptional repression. Mol. Cell 77, 840–856 e5 (2020).
Barbour, H., Daou, S., Hendzel, M. & Affar, E. B. Polycomb group-mediated histone H2A monoubiquitination in epigenome regulation and nuclear processes. Nat. Commun. 11, 5947 (2020).
Kasinath, V. et al. JARID2 and AEBP2 regulate PRC2 in the presence of H2AK119ub1 and other histone modifications. Science 371, eabc3393 (2021).
Wilson, M. D. et al. The structural basis of modified nucleosome recognition by 53BP1. Nature 536, 100–103 (2016).
Hu, Q. et al. Mechanisms of BRCA1-BARD1 nucleosome recognition and ubiquitylation. Nature 596, 438–443 (2021).
Dai, L. et al. Structural insight into BRCA1-BARD1 complex recruitment to damaged chromatin. Mol. Cell 81, 2765–2777.e6 (2021).
Huang, C., Sloan, E. A. & Boerkoel, C. F. Chromatin remodeling and human disease. Curr. Opin. Genet. Dev. 13, 246–252 (2003).
Clark, J. et al. Identification of novel genes, SYT and SSX, involved in the t(X;18)(p11.2;q11.2) translocation found in human synovial sarcoma. Nat. Genet. 7, 502–508 (1994).
Ladanyi, M. et al. Impact of SYT-SSX fusion type on the clinical behavior of synovial sarcoma: a multi-institutional retrospective study of 243 patients. Cancer Res. 62, 135–140 (2002).
Kadoch, C. & Crabtree, G. R. Reversible disruption of mSWI/SNF (BAF) complexes by the SS18-SSX oncogenic fusion in synovial sarcoma. Cell 153, 71–85 (2013).
McBride, M. J. et al. The SS18-SSX fusion oncoprotein hijacks BAF complex targeting and function to drive synovial sarcoma. Cancer Cell 33, 1128–1141.e7 (2018).
Banito, A. et al. The SS18-SSX oncoprotein hijacks KDM2B-PRC1.1 to drive synovial sarcoma. Cancer Cell 33, 527–541.e8 (2018).
McBride, M. J. et al. The nucleosome acidic patch and H2A ubiquitination underlie mSWI/SNF recruitment in synovial sarcoma. Nat. Struct. Mol. Biol. 27, 836–845 (2020).
Chu, G.-C. et al. Cysteine-aminoethylation-assisted chemical ubiquitination of recombinant histones. J. Am. Chem. Soc. 141, 3654–3663 (2019).
Ai, H. et al. Examination of the deubiquitylation site selectivity of USP51 by using chemically synthesized ubiquitylated histones. ChemBioChem 20, 221–229 (2019).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Paul, S. Histone ‘acidic patch’: a hotspot in chromatin biology. Nucleus 64, 271–275 (2021).
McGinty, R. K. & Tan, S. Principles of nucleosome recognition by chromatin factors and enzymes. Curr. Opin. Struct. Biol. 71, 16–26 (2021).
McGinty, R. K. & Tan, S. Recognition of the nucleosome by chromatin factors and enzymes. Curr. Opin. Struct. Biol. 37, 54–61 (2016).
Nodelman, I. M. et al. Nucleosome recognition and DNA distortion by the Chd1 remodeler in a nucleotide-free state. Nat. Struct. Mol. Biol. 29, 121–129 (2022).
Yuan, J., Chen, K., Zhang, W. & Chen, Z. Structure of human chromatin-remodelling PBAF complex bound to a nucleosome. Nature 605, 166–171 (2022).
Luger, K., Mäder, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389, 251–260 (1997).
Dao, H. T., Dul, B. E., Dann, G. P., Liszczak, G. P. & Muir, T. W. A basic motif anchoring ISWI to nucleosome acidic patch regulates nucleosome spacing. Nat. Chem. Biol. 16, 134–142 (2020).
Xiao, X. et al. Histone H2A ubiquitination reinforces mechanical stability and asymmetry at the single-nucleosome level. J. Am. Chem. Soc. 142, 3340–3345 (2020).
Worden, E. J., Hoffmann, N. A., Hicks, C. W. & Wolberger, C. Mechanism of cross-talk between H2B ubiquitination and H3 methylation by Dot1L. Cell 176, 1490–1501.e12 (2019).
Ai, H. et al. H2B Lys34 ubiquitination induces nucleosome distortion to stimulate Dot1L activity. Nat. Chem. Biol. 18, 972–980 (2022).
McGinty, R. K., Henrici, R. C. & Tan, S. Crystal structure of the PRC1 ubiquitylation module bound to the nucleosome. Nature 514, 591–596 (2014).
Ai, H. et al. Chemical synthesis of post-translationally modified H2AX reveals redundancy in interplay between histone phosphorylation, ubiquitination, and methylation on the binding of 53BP1 with nucleosomes. J. Am. Chem. Soc. 144, 18329–18337 (2022).
Schreiber, S. L. The rise of molecular glues. Cell 184, 3–9 (2021).
Dong, G., Ding, Y., He, S. & Sheng, C. Molecular glues for targeted protein degradation: from serendipity to rational discovery. J. Med. Chem. 64, 10606–10620 (2021).
Dyer, P. N. et al. Reconstitution of nucleosome core particles from recombinant histones and DNA. Methods Enzymol. 375, 23–44 (2004).
Ai, H. et al. Synthetic E2-Ub-nucleosome conjugates for studying nucleosome ubiquitination. Chem 9, 1221–1240 (2023).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Acknowledgements
We thank the National Key R&D Program of China (grant no. 2022YFC3401500) for financial support. This study was supported by the National Natural Science Foundation of China (grant nos. 22137005, 92253302, 22227810 for L. Liu, and grant no. 22277073 for M.P.), the Shanghai Rising-Star Program (grant no. 22QA1404900), the Shanghai Pilot Program for Basic Research – Shanghai Jiao Tong University (grant no. 21TQ1400224) and the China Postdoctoral Science Foundation (grant nos. 2022TQ0170 and 2022M720075 for H.A.). We also acknowledge the XPLORER prize. H.A. thanks the National Facility for Translational Medicine (Shanghai) for funding. We acknowledge the Tsinghua University Branch of the China National Center for Protein Sciences (Beijing) for cryo-EM screening and data collection in 200-kV Arctica Tecnai microscopy and 300-kV Titan Krios microscopy. We thank the laboratory of S. Ding (School of Pharmaceutical Sciences, Tsinghua University, Beijing, China) for supporting our cellular experiments.
Author information
Authors and Affiliations
Contributions
H.A., Z.T., M.P. and L. Liu proposed the idea, designed the experiments and analyzed the results. Z.T., H.A., Z.X. and Q.S. cloned the plasmids, expressed the proteins (SSX1 and histones) and reconstituted the nucleosomes. H.A. and Z.T. prepared the cryo-EM samples, collected the cryo-EM data and solved the structures of the H2AK119Ub nucleosomes, SSX1–unmodified nucleosome complex and SSX1–H2AK119Ub nucleosome complex. Z.T. and Z.X. performed the pull-down and AlphaLISA experiments. Z.T., H.A. and Z.X. conducted the REAA assay. Z.X. synthesized the histone H2AK119Ub and H3 mutants. G.-C.C. synthesized the molecules for CAACU chemical ligation. K.H. and Q.X. conducted cellular experiments (immunofluorescence and ChIP–seq). Z.T., H.A., M.P. and L. Liu wrote and revised the manuscript. H.A., Z.T., Z.X., M.P., L. Liu, Z.D., M.S., Y.D., L. Liang and J.-B.L. read and analyzed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Dimitris Typas, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 SSX1-H2AK119Ub nucleosome complex purification and 2D, 3D classification.
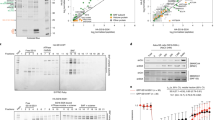
a. Schematic representation of the fusion oncoprotein SS18-SSX1 and GST-SSX1 construct. Arrows indicate the fusion site. b. Size exclusion chromatography (SEC) of SSX1-H2AK119Ub nucleosome complex. The peak fraction was analysed by SDS-PAGE and visualized with Coomassie blue-staining. c. SEC of SSX1-H2AK119Ub nucleosome complex after glutaraldehyde cross-linking. d. Selected 2D class averages generated from the particles used to reconstruct the SSX1-H2AK119Ub complex. e. The cryo-EM density map of SSX1-H2AK119Ub nucleosome at the level of 0.0045 in ChimeraX. The additional density of GST was visualized (as marked in red orange line) in d, e. f. SDS-PAGE and Native PAGE of SSX1-H2AK119Ub nucleosome complexes after glutaraldehyde cross-linking. These gels are representative of three independent experiments. The original gels are provided as source data.
Extended Data Fig. 2 Structure determination of the SSX1-H2AK119Ub nucleosome complex.
a. Representative motion-corrected micrograph from data set. b. 2D classification averages of SSX1-H2AK119Ub complex. c. Flow chart of data processing steps.
Extended Data Fig. 3 Cryo-EM validation and example density for the structure of SSX1-H2AK119Ub nucleosome complex.
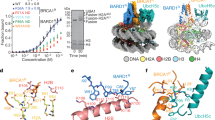
a. Top, the final 3.1 Å reconstruction of the SSX1-H2AK119Ub nucleosome complex (1:1 complex) contoured at a level of 0.015 is coloured according to local resolution with cyan and crimson representing the highest and lowest resolution, respectively. Bottom, euler angle distribution of all the particles used in the final reconstruction 3D reconstruction. b. Top, the final 3.6 Å reconstruction of the SSX1-H2AK119Ub nucleosome complex (2:1 complex) contoured at a level of 0.009 is coloured according to local resolution with cyan and crimson representing the highest and lowest resolution, respectively. Bottom, Euler angle distribution of all the particles used in the final reconstruction 3D reconstruction. c. FSC curve calculated between two independent half-maps from refinements in RELION reported at 0.143 FSC cutoff. d. Fourier shell correlation (FSC) curves between model and the Cryo-EM map. Resolution at FSC 0.5 is indicated. e. Representative regions of the cryo-EM density map for the H2A, H2B, H3, H4, SSX1, Ub, and DNA. Models are presented in sticks and densities in surfaces. f. Representative regions of the cryo-EM density map for the H2A/H2B acidic patch. g. Representative regions of the cryo-EM density map for H3-UbH2AK119 basic groove. h. Representative regions of the cryo-EM density map for H2A hydrophobic pocket.
Extended Data Fig. 4 Interactions between SSX1 and the nucleosomal H2A/H2B surface.
a. Close-up views of the contacts between SSX1 and the H2A–H2B acidic patch. The histone octamer is presented as surface electrostatic potential, positively charged surface coloured in blue and negatively charged surface in red. b, SSX1 (residues W164, R167) interact with H2B (residues Q47, E113). c. SSX1 (residues L168, R169) interact with the main chain of H2A α2 helix and H2A (residues E61, D90, E92). The cryo-EM density map of SSX1, H2A, H2B in b, c are presented in mesh. d. Structural comparisons of the H2A hydrophobic pocket bound by SSX1, Chd1 (PDB:7tn2), SMARCA4 of PBAF complex (PDB:7vdv). e. Structure-based sequence alignment between SSX1 (residues 155–188), CHD1 (residues 115–147) and SMARCA4 (residues 1364–1390). Conserved residues are underlined in yellow. f. Multiple-sequence alignment of human SSX1 (1–9). Highly conserved basic motif (blue) and acidic motif (red), and tyrosine (yellow) are indicated. Residues involved in the H2A/H2B acidic patch, H2A α2/3 hydrophobic pocket, and H3-UbH2AK119 basic groove binding are marked.
Extended Data Fig. 5 Structure determination of the SSX1-unmodified nucleosome.
a. Flow chart of data processing steps. b. 2D classification averages of SSX1-unmodified nucleosome complex. c. The final 3.0 Å reconstruction of the SSX1-unmodified nucleosome complex is coloured according to local resolution with cyan and crimson representing the highest and lowest resolution, respectively. d. Euler angle distribution of all the particles used in the final reconstruction 3D reconstruction. e. FSC curve calculated between two independent half-maps from refinements in RELION reported at 0.143 FSC cut off.
Extended Data Fig. 6 Mutations in crucial SSX1 interactions with H2AK119Ub nucleosomes noticeably affect the cellular localization and genomic distribution of SS18-SSX1 BAF complexes.
a. Immunofluorescence analysis of Flag-SS18 and Flag-SS18-SSX variants (WT, W164A, R167A, L168A, R169A, Y177A, S181A-D182K-E184R, S181A-D182K-E184R EDDE deletion, 7 acidic amino acids deletion) relative to H2AK119Ub in 293T cells. Arrowheads indicate Barr bodies. Scale bars, 20 μm. b. Distribution figures depict profiles for genome-wide localization of H2AUb, Flag-SS18-SSX, Flag-SS18, and Flag-SS18-SSX variants. c. ChIP-seq density heatmaps reflecting the characteristic retargeting patterns of Flag-SS18-SSX1 compared to Flag-SS18 and Flag-SS18-SSX1 (W164A, R167A, L168A, R169A, Y177A, 7 acidic amino acids deletion, S181A-D182K-E184R, S181A-D182K-E184R EDDE deletion).
Extended Data Fig. 7 Structural comparison of the H2AK119Ub nucleosome and SSX1-H2AK119Ub nucleosome complex (1:1 complex), SSX1-H2AK119Ub nucleosome complex (2:1 complex).
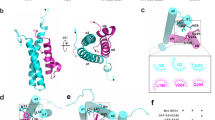
a-c. Top and middle, two views of the three Cryo-EM maps. Bottom, cartoon representation of SSX1 binding to H2AK119Ub nucleosome-induced DNA unwrapping. Table, the degree of DNA unwrapping was correlated with the density of SSX1 and the density of Ub.
Extended Data Fig. 8 Structural comparison of the unmodified nucleosome and SSX1-H2AK119Ub nucleosome complex (1:1 complex), SSX1-H2AK119Ub nucleosome complex (2:1 complex).
a. Left, Overall view of the superimposed structures of the unmodified nucleosome and 1:1 complex. Right, close-up view of the contact between exit DNA and H3 α-N helix. b. Left, overall view of the superimposed structures of the unmodified nucleosome and 2:1 complex. Top right, close-up view of the contact between exit DNA and H3 α-N helix. Bottom right, close-up view of the contact between entry DNA and H3 α-N helix.
Extended Data Fig. 9 Restriction enzyme accessibility assay (REAA) experiments on different nucleosomes with and without SSX1.
The DNA unwrapping caused by SSX1 binding on the 601 H2AK119Ub nucleosome (a), the 25N25 H2AK119Ub nucleosome(b) and 25N25 H1 H2AK119Ub nucleosome and (c) was analysed by REAA. d. The DNA unwrapping on the 601 H3R49E/R52E H2AK119Ub nucleosome and 601 H3R49E/R52E H2AK119Ub nucleosome and was analysed by REAA. The above gels are representative of three independent experiments. The original gels are provided as source data.
Extended Data Fig. 10 Structural comparison of JARID2, AEBP2, and SSX1 bound to the H2AK119Ub nucleosome.
a. Close-up view of the contact between JARID2 and Ub. Ub is shown in a crimson surface. JARID2 and nucleosomes are shown in cartoon. b. Close-up view of the contact between AEBP2 and Ub. Ub is shown in media spring orange surface. c. Close-up view of the contact between SSX1 and H3-UbH2AK119 basic groove. Ub is shown in yellow surface. d. Structural comparison of JARID2, SSX1, AEBP2 bound to the H2AK119Ub nucleosome. Histone octamer is shown in light grey surface. e. Comparison of the position of Ub in the structure of AEBP2 bound to H2AK119Ub nucleosomes and the structure of JARID2 bound to H2AK119Ub nucleosomes. f. Comparison of the position of Ub in the structure of AEBP2 bound to H2AK119Ub nucleosomes and the structure of SSX1 bound to H2AK119Ub nucleosomes.
Supplementary information
Supplementary Information
Supplementary materials and methods, Figs. 1–18 and uncropped scans of gels.
Source data
Source Data Fig. 1
Uncropped gels for Fig. 1.
Source Data Fig. 2
Uncropped gels for Fig. 2.
Source Data Fig. 5
Uncropped gels for Fig. 5.
Source Data Extended Data Fig. 1
Uncropped gels for Extended Fig. 1.
Source Data Extended Data Fig. 9
Uncropped gels for Extended Fig. 9.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tong, Z., Ai, H., Xu, Z. et al. Synovial sarcoma X breakpoint 1 protein uses a cryptic groove to selectively recognize H2AK119Ub nucleosomes. Nat Struct Mol Biol 31, 300–310 (2024). https://doi.org/10.1038/s41594-023-01141-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-01141-1
This article is cited by
-
Recent advances in chemical protein synthesis: method developments and biological applications
Science China Chemistry (2024)