Abstract
Resistant starch is a prebiotic accessed by gut bacteria with specialized amylases and starch-binding proteins. The human gut symbiont Ruminococcus bromii expresses Sas6 (Starch Adherence System member 6), which consists of two starch-specific carbohydrate-binding modules from family 26 (RbCBM26) and family 74 (RbCBM74). Here, we present the crystal structures of Sas6 and of RbCBM74 bound with a double helical dimer of maltodecaose. The RbCBM74 starch-binding groove complements the double helical α-glucan geometry of amylopectin, suggesting that this module selects this feature in starch granules. Isothermal titration calorimetry and native mass spectrometry demonstrate that RbCBM74 recognizes longer single and double helical α-glucans, while RbCBM26 binds short maltooligosaccharides. Bioinformatic analysis supports the conservation of the amylopectin-targeting platform in CBM74s from resistant-starch degrading bacteria. Our results suggest that RbCBM74 and RbCBM26 within Sas6 recognize discrete aspects of the starch granule, providing molecular insight into how this structure is accommodated by gut bacteria.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The X-ray structures and diffraction data reported in this paper have been deposited in the Protein Data Bank under accession codes 7UWU, 7UWVand 7UWW. The SAXS data are deposited in the small angle X-ray scattering database (SASDB) under accession code SASDPE2 (ref. 98). CBM74 sequences were pulled from the CAZy database (http://www.cazy.org; CAZy update, March 2022) and via BLAST against GenBank (https://www.ncbi.nlm.nih.gov/genbank) and/or UniProt (https://www.uniprot.org) databases in March 2022. Native MS data are publicly available in the Deep Blue Data Repository administered by the University of Michigan at https://doi.org/10.7302/5fmh-8f87. The HDX–MS data are publicly available in the Zenodo database under accession number 8371163 (https://zenodo.org/record/8371163). Source data and Supplementary Data files are provided with this paper. All other relevant data supporting the key findings of this study are available within the article, its Supplementary Information or from the corresponding authors upon reasonable request. Source data are provided with this paper.
Code availability
SAXS data collection was performed using the Python-based BioCon software developed at and for the BioCAT beamline, available at https://github.com/biocatiit/beamline-control-user/tree/master/biocon. The UniDec software is available at https://github.com/michaelmarty/UniDec/. Source data are provided with this paper.
References
Salminen, S., Isolauri, E. & Onnela, T. Gut flora in normal and disordered states. Chemotherapy 41, 5–15 (1995).
Gibson, G. R. & Roberfroid, M. B. Dietary modulation of the human colonic microbiota: introducing the concept of prebiotics. J. Nutr. 125, 1401–1412 (1995).
Cummings, J. H. & Macfarlane, G. T. The control and consequences of bacterial fermentation in the human colon. J. Appl. Bacteriol. 70, 443–459 (1991).
Backhed, F. et al. Host–bacterial mutualism in the human intestine. Science 307, 1915–1920 (2005).
Wu, X. et al. Effects of the intestinal microbial metabolite butyrate on the development of colorectal cancer. J. Cancer 9, 2510–2517 (2018).
Zaman, S. A. & Sarbini, S. R. The potential of resistant starch as a prebiotic. Crit. Rev. Biotechnol. 36, 578–584 (2016).
Bertoft, E. Understanding starch structure: recent progress. Agronomy 7, 56 (2017).
Pérez, S. & Bertoft, E. The molecular structures of starch components and their contribution to the architecture of starch granules: a comprehensive review. Starch 62, 389–420 (2010).
Ze, X. et al. Ruminococcus bromii is a keystone species for the degradation of resistant starch in the human colon. ISME J. 6, 1535–1543 (2012).
Jung, D. H. et al. Bifidobacterium adolescentis P2P3, a human gut bacterium having strong non-gelatinized resistant starch-degrading activity. J. Microbiol. Biotechnol. 29, 1904–1915 (2019).
Teichmann, J. & Cockburn, D. W. In vitro fermentation reveals changes in butyrate production dependent on resistant starch source and microbiome composition. Front. Microbiol. 12, 640253 (2021).
Duranti, S. et al. Genomic characterization and transcriptional studies of the starch-utilizing strain Bifidobacterium adolescentis 22L. Appl. Environ. Microbiol. 80, 6080–6090 (2014).
Belenguer, A. et al. Two routes of metabolic cross-feeding between Bifidobacterium adolescentis and butyrate-producing anaerobes from the human gut. Appl. Environ. Microbiol. 72, 3593–3599 (2006).
Venkataraman, A. et al. Variable responses of human microbiomes to dietary supplementation with resistant starch. Microbiome 4, 33 (2016).
Baxter, N. T. et al. Dynamics of human gut microbiota and short-chain fatty acids in response to dietary interventions with three fermentable fibers. mBio 10, e02566–18 (2019).
Ze, X. et al. Unique organization of extracellular amylases into amylosomes in the resistant starch-utilizing human colonic firmicutes bacterium Ruminococcus bromii. mBio 6, e01058–15 (2015).
Smith, S. P. & Bayer, E. A. Insights into cellulosome assembly and dynamics: from dissection to reconstruction of the supramolecular enzyme complex. Curr. Opin. Struct. Biol. 23, 686–694 (2013).
Bayer, E. A., Morag, E. & Lamed, R. The cellulosome—a treasure-trove for biotechnology. Trends Biotechnol. 12, 379–386 (1994).
Mukhopadhya, I. et al. Sporulation capability and amylosome conservation among diverse human colonic and rumen isolates of the keystone starch-degrader Ruminococcus bromii. Environ. Microbiol. 20, 324–336 (2018).
Janecek, S. et al. Starch-binding domains as CBM families—history, occurrence, structure, function and evolution. Biotechnol. Adv. 37, 107451 (2019).
Valk, V. et al. Carbohydrate-binding module 74 is a novel starch-binding domain associated with large and multidomain alpha-amylase enzymes. FEBS J. 283, 2354–2368 (2016).
Dobranowski, P. A. & Stintzi, A. Resistant starch, microbiome, and precision modulation. Gut Microbes 13, 1926842 (2021).
Ravi, A. et al. Hybrid metagenome assemblies link carbohydrate structure with function in the human gut microbiome. Commun. Biol. 5, 932 (2022).
Xu, J. et al. Metatranscriptomic analysis of colonic microbiota’s functional response to different dietary fibers in growing pigs. Anim. Microbiome 3, 45 (2021).
Zhang, H. et al. dbCAN2: a meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 46, W95–W101 (2018).
Lombard, V. et al. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495 (2014).
Blum, M. et al. The InterPro protein families and domains database: 20 years on. Nucleic Acids Res. 49, D344–D354 (2021).
Cerqueira, F. M. et al. Sas20 is a highly flexible starch-binding protein in the Ruminococcus bromii cell-surface amylosome. J. Biol. Chem. 298, 101896 (2022).
Fontes, C. M. & Gilbert, H. J. Cellulosomes: highly efficient nanomachines designed to deconstruct plant cell wall complex carbohydrates. Annu. Rev. Biochem. 79, 655–681 (2010).
Matsui, M., Kakuta, M. & Misaki, A. Comparison of the unit-chain distributions of glycogens from different biological sources, revealed by anion exchange chromatography. Biosci. Biotechnol. Biochem. 57, 623–627 (1993).
Brewer, M. K. & Gentry, M. S. Brain glycogen structure and its associated proteins: past, present and future. Adv. Neurobiol. 23, 17–81 (2019).
Singh, R. S., Saini, G. K. & Kennedy, J. F. Pullulan: microbial sources, production and applications. Carbohydr. Polym. 73, 515–531 (2008).
Khalikova, E., Susi, P. & Korpela, T. Microbial dextran-hydrolyzing enzymes: fundamentals and applications. Microbiol Mol. Biol. Rev. 69, 306–325 (2005).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Valk, V., Rachel, M. V. D. K. & Dijkhuizen, L. The evolutionary origin and possible functional roles of FNIII domains in two Microbacterium aurum B8.A granular starch degrading enzymes, and in other carbohydrate acting enzymes. Amylase 1, 1–11 (2017).
Holm, L. Using Dali for protein structure comparison. Methods Mol. Biol. 2112, 29–42 (2020).
Notenboom, V. et al. Crystal structures of the family 9 carbohydrate-binding module from Thermotoga maritima xylanase 10A in native and ligand-bound forms. Biochemistry 40, 6248–6256 (2001).
Milles, L. F. et al. Calcium stabilizes the strongest protein fold. Nat. Commun. 9, 4764 (2018).
Zheng, H. et al. CheckMyMetal: a macromolecular metal-binding validation tool. Acta Crystallogr. Struct. Biol. 73, 223–233 (2017).
Strynadka, N. C. J. & James, M. N. G. Towards an understanding of the effects of calcium on protein structure and function. Curr. Opin. Struct. Biol. 1, 905–914 (1991).
Boraston, A. B. et al. A structural and functional analysis of α-glucan recognition by family 25 and 26 carbohydrate-binding modules reveals a conserved mode of starch recognition. J. Biol. Chem. 281, 587–598 (2006).
Holm, L. DALI and the persistence of protein shape. Protein Sci. 29, 128–140 (2020).
Cockburn, D. W. et al. Novel carbohydrate binding modules in the surface anchored α-amylase of Eubacterium rectale provide a molecular rationale for the range of starches used by this organism in the human gut. Mol. Microbiol. 107, 249–264 (2018).
Rodriguez-Sanoja, R. et al. A single residue mutation abolishes attachment of the CBM26 starch-binding domain from Lactobacillus amylovorus α-amylase. J. Ind. Microbiol. Biotechnol. 36, 341–346 (2009).
Guillen, D. et al. Alpha-amylase starch binding domains: cooperative effects of binding to starch granules of multiple tandemly arranged domains. Appl. Environ. Microbiol. 73, 3833–3837 (2007).
Abbott, D. W. & Boraston, A. B. Quantitative approaches to the analysis of carbohydrate-binding module function. Methods Enzymol. 510, 211–231 (2012).
Imberty, A. et al. The double-helical nature of the crystalline part of A-starch. J. Mol. Biol. 201, 365–378 (1988).
Imberty, A. & Perez, S. A revisit to the three-dimensional structure of B-type starch. Biopolymers 27, 1205–1221 (1988).
O’Sullivan, A. C. & Perez, S. The relationship between internal chain length of amylopectin and crystallinity in starch. Biopolymers 50, 381–390 (1999).
Gessler, K. et al. V-Amylose at atomic resolution: X-ray structure of a cycloamylose with 26 glucose residues (cyclomaltohexaicosaose). Proc. Natl Acad. Sci. USA 96, 4246–4251 (1999).
Ashkenazy, H. et al. ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res. 44, W344–W350 (2016).
Ashkenazy, H. et al. ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res. 38, W529–W533 (2010).
Celniker, G. et al. ConSurf: using evolutionary data to raise testable hypotheses about protein function. Isr. J. Chem. 53, 199–206 (2013).
Soper, M. T. et al. Amyloid-β/neuropeptide interactions assessed by ion mobility-mass spectrometry. Phys. Chem. Chem. Phys. 15, 8952–8961 (2013).
Hizukuri, S. Relationship between the distribution of the chain length of amylopectin and the crystalline structure of starch granules. Carbohydr. Res. 141, 295–306 (1985).
Cerqueira, F. M. et al. Starch digestion by gut bacteria: crowdsourcing for carbs. Trends Microbiol. 28, 95–108 (2020).
Jung, D. H. et al. The presence of resistant starch-degrading amylases in Bifidobacterium adolescentis of the human gut. Int. J. Biol. Macromol. 161, 389–397 (2020).
Rees, D. A. & Welsh, E. J. Secondary and tertiary structure of polysaccharides in solutions and gels. Angew. Chem. 16, 214–224 (1977).
Tauzin, A. S. et al. Molecular dissection of xyloglucan recognition in a prominent human gut symbiont. mBio 7, e02134–15 (2016).
Hillmann, G. Measurement by end-point determination on paper, in Methods of enzymatic analysis 2nd edn (ed. Bergmeyer H. U.) (Academic Press, 1974).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Turnbull, W. B. & Daranas, A. H. On the value of c: can low affinity systems be studied by isothermal titration calorimetry? J. Am. Chem. Soc. 125, 14859–14866 (2003).
Vonrhein, C. et al. Data processing and analysis with the autoPROC toolbox. Acta Crystallogr. 67, 293–302 (2011).
Kabsch, W. Xds. Acta Crystallogr. Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. Biol. Crystallogr. 69, 1204–1214 (2013).
El Omari, K. et al. Pushing the limits of sulfur SAD phasing: de novo structure solution of the N-terminal domain of the ectodomain of HCV E1. Acta Crystallogr. Biol. Crystallogr. 708, 2197–2203 (2014).
Terwilliger, T. C. et al. Decision-making in structure solution using Bayesian estimates of map quality: the PHENIX AutoSol wizard. Acta Crystallogr. 65, 582–601 (2009).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. Biol. Crystallogr. 60, 2126–2132 (2004).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. Biol. Crystallogr. 68, 352–367 (2012).
Zheng, H. et al. Validation of metal-binding sites in macromolecular structures with the CheckMyMetal web server. Nat. Protoc. 9, 156–170 (2014).
Agirre, J. et al. Privateer: software for the conformational validation of carbohydrate structures. Nat. Struct. Mol. Biol. 22, 833–834 (2015).
Kirby, N. et al. Improved radiation dose efficiency in solution SAXS using a sheath flow sample environment. Acta Crystallogr. Struct. Biol. 72, 1254–1266 (2016).
Hopkins, J. B., Gillilan, R. E. & Skou, S. BioXTAS RAW: improvements to a free open-source program for small-angle X-ray scattering data reduction and analysis. J. Appl. Crystallogr. 50, 1545–1553 (2017).
Rambo, R. P. & Tainer, J. A. Accurate assessment of mass, models and resolution by small-angle scattering. Nature 496, 477–481 (2013).
Piiadov, V. et al. SAXSMoW 2.0: online calculator of the molecular weight of proteins in dilute solution from experimental SAXS data measured on a relative scale. Protein Sci. 28, 454–463 (2019).
Franke, D., Jeffries, C. M. & Svergun, D. I. Machine learning methods for X-ray scattering data analysis from biomacromolecular solutions. Biophys. J. 114, 2485–2492 (2018).
Svergun, D. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Crystallogr. 25, 495–503 (1992).
Manalastas-Cantos, K. et al. ATSAS 3.0: expanded functionality and new tools for small-angle scattering data analysis. J. Appl. Crystallogr. 54, 343–355 (2021).
Schneidman-Duhovny, D. et al. FoXS, FoXSDock and MultiFoXS: single-state and multi-state structural modeling of proteins and their complexes based on SAXS profiles. Nucleic Acids Res. 44, W424–W429 (2016).
Murphy, R. D. et al. The Toxoplasma glucan phosphatase TgLaforin utilizes a distinct functional mechanism that can be exploited by therapeutic inhibitors. J. Biol. Chem. 298, 102089 (2022).
van de Waterbeemd, M. et al. High-fidelity mass analysis unveils heterogeneity in intact ribosomal particles. Nat. Methods 14, 283–286 (2017).
Marty, M. T. et al. Bayesian deconvolution of mass and ion mobility spectra: from binary interactions to polydisperse ensembles. Anal. Chem. 87, 4370–4376 (2015).
Gulbakan, B. et al. Native electrospray ionization mass spectrometry reveals multiple facets of aptamer–ligand interactions: from mechanism to binding constants. J. Am. Chem. Soc. 140, 7486–7497 (2018).
Wang, W., Kitova, E. N. & Klassen, J. S. Influence of solution and gas phase processes on protein–carbohydrate binding affinities determined by nanoelectrospray Fourier transform ion cyclotron resonance mass spectrometry. Anal. Chem. 75, 4945–4955 (2003).
Báez Bolivar, E. G. et al. Submicron emitters enable reliable quantification of weak protein–glycan interactions by ESI–MS. Anal. Chem. 93, 4231–4239 (2021).
Loos, M. et al. Accelerated isotope fine structure calculation using pruned transition trees. Anal. Chem. 87, 5738–5744 (2015).
Drula, E. et al. The carbohydrate-active enzyme database: functions and literature. Nucleic Acids Res. 50, D571–D577 (2022).
Valk, V. et al. Degradation of granular starch by the bacterium Microbacterium aurum Strain B8.A involves a modular α-amylase enzyme system with FNIII and CBM25 domains. Appl. Environ. Microbiol. 81, 6610–6620 (2015).
Altschul, S. F. et al. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Sayers, E. W. et al. GenBank. Nucleic Acids Res. 49, D92–D96 (2021).
UniProt Consortium. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 49, D480–D489 (2021).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Whelan, S. & Goldman, N. A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol. Biol. Evol. 18, 691–699 (2001).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–791 (1985).
Kumar, S. et al. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23, 127–128 (2007).
Kikhney, A. G. et al. SASBDB: towards an automatically curated and validated repository for biological scattering data. Protein Sci. 29, 66–75 (2020).
Acknowledgements
This work is primarily supported by a Ruth L. Kirschstein National Research Service Award Individual Predoctoral Fellowship (F31–F31AT011282 to A.L.P.) from the National Center for Complementary and Integrative Health (NCCIH) and a Research Program Project grant (P01-HL149633 to N.M.K.) from the National Heart, Lung and Blood Institute (NHLBI) of the National Institutes of Health (NIH). Next-generation Native MS technologies were supported by the National Institute of General Medical Sciences (NIGMS) of the NIH (R01-GM095832 to B.T.R.). HDX–MS acquisition was supported by the National Science Foundation (NSF) (DBI 2018007 to C.W.V.K.). The structural biology approaches used resources of the Advanced Photon Source; a U.S. Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under contract no. DE-AC02-06CH11357. The Biophysics Collaborative Access Team is supported by P30-GM138395 from NIGMS-NIH. Use of the Pilatus3 X 1M detector was provided by Grant 1S10OD018090-01 from NIGMS-NIH. Use of the LS-CAT Sector 21 was supported by the Michigan Economic Development Corporation and the Michigan Technology Tri-Corridor (grant 085P1000817). S.J. and F.M. thank the Slovak Grant Agency VEGA for the financial support by grant no. 2/0146/21. In collaboration with this research, we acknowledge support from the University of Michigan Biomedical Research Core Facilities Light Microscopy Core. For the native MS work, we would like to acknowledge the Biological Mass Spectrometry facility at the University of Michigan Department of Chemistry. The content is solely the responsibility of the authors and does not necessarily represent the official views of VEGA, the National Science Foundation or the NIH.
Author information
Authors and Affiliations
Contributions
N.M.K. and A.L.P. conceived the study. A.L.P., F.M.C., R.C.V-V., K.M.A., F.M., T.C., Z.W., J.H., C.W.V.K, S.J., B.T.R. and N.M.K. curated the data. A.L.P., F.M.C., R.C.V-V., K.M.A., F.M., T.C., Z.W., J.H., C.W.V.K., S.J., B.T.R. and N.M.K. conducted formal analysis and data interpretation. N.M.K., A.L.P., C.W.V.K., S.J., B.T.R., Z.W. and J.H. were involved with funding acquisition. A.L.P., F.M.C., R.C.V-V., K.M.A., F.M., T.C., Z.W., J.H., C.W.V.K. and S.J. performed the investigations. A.L.P., F.M.C., R.C.V-V., K.M.A., F.M., T.C., Z.W., J.H., C.W.V.K., S.J., B.T.R. and N.M.K. created the methodology. A.L.P., F.M.C., R.C.V-V., C.W.V.K., S.J. and N.M.K. wrote the original draft of the manuscript. A.L.P., F.M.C., R.C.V-V., F.M., J.H., C.W.V.K., S.J., B.T.R. and N.M.K. were involved with writing, review and editing. N.M.K., J.H., C.W.V.K., S.J. and B.T.R. supervised the study. A.L.P., F.M.C., R.C.V-V., T.C. and S.J. visualized the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Stephen Withers and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Katarzyna Ciazynska, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Small Angle X-Ray Scattering indicates that Sas6 remains mostly compact in solution with minor extension beyond that of the crystal structure.
a. Total subtracted scattering intensity (left y axis) and Rg (right y axis) as a function of time for the SEC-SAXS elution. The elution resolved several peaks, including a single strong monodisperse peak as indicated by the constant radius of gyration (Rg). b. Guinier fit analysis with normalized residual shown in the bottom panel. Rg and I(0) values of 29.44 ± 0.04Å and 0.04 ± 3.65 × 10−5 were obtained and the fit and normalized fit residuals confirmed this peak was monodisperse. The molecular weight of Sas6T from the SAXS data was calculated to be 61.0 kDa (theoretical 68.9 kDa) indicating it is primarily monomeric in solution. c. P(r) versus r normalized by I(0). The Dmax from the P(r) function for Sas6T is 90Å. The overall shape of the P(r) function for Sas6T, calculated by indirect Fourier transform (IFT) using GNOM, has a relatively Gaussian shape that is characteristic of a globular compact particle with the main peak at r = ∼30 Å. There is a small peak at r = 55Å which suggests there are two structurally separate motifs, possibly RbCBM26 and RbCBM74. d. Dimensionless Kratky plot; y = 3/e and x = \(\sqrt{3}\) as dashed gray lines to indicate where a globular protein would peak. The small plateau in the mid to high q region, around qRg = 5 in the dimensionless Kratky plot indicates some extension or disorder in the system. These results suggest the presence of two separate modules with flexibility between them, likely corresponding to the two CBMs. e. FoXS and f. MultiFoXS fits (black) to the Sas6T SAXS data (red) with normalized residual shown in the bottom panel. The FoXS fit had a χ2= 2.46 and showed systematic deviations in the normalized fit residual suggesting significant differences between the lowest energy conformation of Sas6T in the crystal structure and the structure of Sas6T in solution. For MultiFoXS we assigned the linkers between the domains (residues 130-137 and 572-583) as flexible. MultiFoXS gave a best fit with a 1-state solution with a χ2= 0.96 and calculated Rg of 29.2Å which corroborates the Guinier Rg calculation. g. Topology map of BIgA and BIgB domains illustrating the Greek key motif in BIgA and showing the loops that hydrogen bond with one another. h. A surface area analysis of the BIg domains using PISA in CCP4 gives a buried surface area of 353.9Å34. Residues providing hydrogen bonding are represented by stick side chains and the hydrogen bonds are shown by dashed yellow lines.
Extended Data Fig. 2 RbCBM74 is a singular globular domain, most similar to TmCBM9.
a. Structure of RbCBM74 (PDB 7uww) colored from N-terminus (blue) to C-terminus (red). b. Short β-strands leading into and out of RbCBM74 domain are colored in red and blue. c. Overlay of TmCBM9 (gold) (PDB 1i82-A) and RbCBM74 (blue). The DALI server calculated an RMSD of 3.2Å and sequence identity of 17%. d. Close-up view of TmCBM9 binding site showing the two TmCBM9 Trp residues involved in binding cellobiose (gold) and W373 of RbCBM74 (blue) which lies in the same region but is occluded from the surface by a loop containing residues 374-384. e. Zoomed in view of calciums coordinated in the RbCBM74 domain with side chains shown in sticks, main chain shown in lines and Ca2+ ions by yellow spheres. Atomic distances are shown in Å and residues are labeled. Residues are colored by element with oxygen shown in red.
Extended Data Fig. 3 RbCBM26 shares a conserved binding site with other CBM26.
The top structural homologs of RbCBM26 from DALI36,42 are the CBM25 from Bacillus halodurans C-125 (BhCBM25) from α-amylase G-6 (PDB ID: 2C3V-A, Z-score: 12.4, RMSD 1.9Å, identity: 16%) and CBM26 (BhCBM26) from the same enzyme (PDB ID: 6B3P-B, Z-score: 12.1, RMSD 1.9Å, identity: 20%)41. Another top DALI result is ErCBM26b of Amy13K from Eubacterium rectale (PDB ID 2C3H-B, Z-score: 10.8, RMSD 1.7Å, identity: 19%)43. a. Sequence alignment of RbCBM26 (RBL236_00020), ErCBM26 (ERE_20420), BhCBM26 (BH0413), and LaCBM26 (Q48502). Conserved binding site residues are indicated by a red arrow while variable residues are indicated by a blue arrow and provide hydrogen bonding. b. Overlay of RbCBM26 (green) with BhCBM26 (PDB 2c3h, orange), and ErCBM26 (PDB 6b3p, purple). c. Overlay of unliganded RbCBM26 (blue) and ACX-bound RbCBM26 (green) showing that loop 1 does not move upon ligand binding. b-strands are numbered for reference.
Extended Data Fig. 4 Representative ITC graphs of Sas6 domains.
Sas6T, RbCBM26, and BIg-RbCBM74-BIg binding to a. potato amylopectin, b. maltodecaose (G10), and c. α-cyclodextrin (ACX). Note that exothermic heat release is denoted with an upward peak on this machine.
Extended Data Fig. 5 RbCBM74 selects a double helical ligand geometry.
a. Overlay of RbCBM74 from Sas6T structure (PDB 7uww) in blue with RbCBM74 from BIg-RbCBM74-BIg co-crystal structure (PDB 7uwv) in deep teal. b. Loop from G374-G382 demonstrating that the unliganded loop (blue) occludes W373 but moves to allow access to W373 in the ligand-bound structure (deep teal). c. An extended view of the geometry of the G10 ligand. Intramolecular hydrogen bonds (3.6Å cutoff for ideal geometry and 3.2Å with minimal acceptable geometry) within and between G10 chains are shown in slate. Φ (O5-C1-O4′-C4′) and ψ (C1-O4′-C4′-C5′) angles of the Glc linkages in the G10 double helix ligand are labeled with G10A in magenta and G10B in grey. d. The geometry of the G10 ligand more closely resembles that of double helical B starch (cyan)48 than single helical cycloamylose (yellow, 1c58)50. Models were manually aligned in PyMOL to compare the angles, pitch, and period of the helical turns.
Extended Data Fig. 6 HDX-MS analysis of RbCBM74.
a. Heatmap of exchange dynamics of BIg-RbCBM74-BIg. All values are the average of three replicates. b. Representative differential uptake for peptides that both showed no significant difference (upper panels) and those which showed significant differential decreased deuteration (lower panels) in the G10 bound BIg-RbCBM74-BIg. Data points are represented by the mean +/− standard deviation.c. Heatmap of the differential exchange dynamics of BIg-RbCBM74-BIg in the absence and presence of G10. Blue represents lower exchange (protection) in the G10 bound form and red higher exchange in the G10 bound form. All values are the average of three replicates.
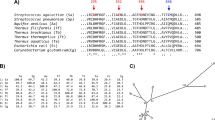
Extended Data Fig. 7 Phylogenetic tree of the 99 CBM74 family members.
a. A maximum-likelihood tree covering 99 sequences with emphasis on the two experimentally characterized CBM74s, Sas6 from Ruminococcus bromii (No. 28, blue cluster) and the subfamily GH13_32 α-amylase from Microbacterium aurum (No. 52; cyan cluster)35. The bootstrap values higher than 70% are shown. For details concerning all 99 CBM74 sequences, see Supplementary Table 1.
Extended Data Fig. 8 Representative ITC graphs of RbCBM74 mutations.
BIg-RbCBM74-BIg, H289A, F236A, and W373A mutations binding to a. maltodecaose (G10), and b. potato amylopectin (PAP). Note that exothermic heat release is denoted with an upward peak on this machine.
Extended Data Fig. 9 Mass spectra of Sas6 constructs at different ligand concentrations (0–300µM) and a fixed protein concentration of 5µM.
Charge states for unbound protein are annotated with an orange dashed line. Peaks corresponding to different bound states are observed after each charge state of the unbound protein. Spectra of a. BIg-RbCBM74-BIg or b. Sas6T in equilibrium with G10. Spectra of c. BIg-RbCBM74-BIg or d. Sas6T in equilibrium with G14.
Supplementary information
Supplementary Information
Supplementary Tables 1–4, Supplementary Figs. 1–5. Legends are with each table or figure.
Source data
Source Data Fig. 1
Raw microscopy images and unprocessed gels and blots.
Source Data Fig. 2
Raw data used to generate Fig. 2f.
Source Data Fig. 2
Unprocessed gels for Fig. 2g.
Source Data Fig. 4
All 99 sequences annotated by dbCAN with data used to generate Fig. 4b.
Source Data Fig. 5
Raw data used to generate Fig. 5a.
Source Data Fig. 5
Uncropped gel for Fig. 5c.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Table 1
Phi-psi angle calculations.
Source Data Extended Data Fig. 9
Statistical Source data for relative intensities in Fig. ED9.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Photenhauer, A.L., Villafuerte-Vega, R.C., Cerqueira, F.M. et al. The Ruminococcus bromii amylosome protein Sas6 binds single and double helical α-glucan structures in starch. Nat Struct Mol Biol 31, 255–265 (2024). https://doi.org/10.1038/s41594-023-01166-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-01166-6