Abstract
The PIWI-interacting RNA (piRNA) pathway is an adaptive defense system wherein piRNAs guide PIWI family Argonaute proteins to recognize and silence ever-evolving selfish genetic elements and ensure genome integrity. Driven by this intensive host–pathogen arms race, the piRNA pathway and its targeted transposons have coevolved rapidly in a species-specific manner, but how the piRNA pathway adapts specifically to target silencing in mammals remains elusive. Here, we show that mouse MILI and human HILI piRNA-induced silencing complexes (piRISCs) bind and cleave targets more efficiently than their invertebrate counterparts from the sponge Ephydatia fluviatilis. The inherent functional differences comport with structural features identified by cryo-EM studies of piRISCs. In the absence of target, MILI and HILI piRISCs adopt a wider nucleic-acid-binding channel and display an extended prearranged piRNA seed as compared with EfPiwi piRISC, consistent with their ability to capture targets more efficiently than EfPiwi piRISC. In the presence of target, the seed gate—which enforces seed–target fidelity in microRNA RISC—adopts a relaxed state in mammalian piRISC, revealing how MILI and HILI tolerate seed–target mismatches to broaden the target spectrum. A vertebrate-specific lysine distorts the piRNA seed, shifting the trajectory of the piRNA–target duplex out of the central cleft and toward the PAZ lobe. Functional analyses reveal that this lysine promotes target binding and cleavage. Our study therefore provides a molecular basis for the piRNA targeting mechanism in mice and humans, and suggests that mammalian piRNA machinery can achieve broad target silencing using a limited supply of piRNA species.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The cryo-EM density maps for HILI–piRNA, MILI–piRNA, MILI–piRNA–target (15 nt), EfPiwiN959K–piRNA and EfPiwiN959K–piRNA–target (16 nt) have been deposited in the Electron Microscopy Data Bank under accession codes EMD-33800, EMD-33817, EMD-33801, EMD-33809 and EMD-33797, respectively. Corresponding atomic models have been deposited in the Protein Data Bank under accession IDs 7YFX, 7YGN, 7YFY, 7YG6 and 7YFQ, respectively. The raw sequence reads for cleavage products by MILI, HILI and EfPiwi are available from NCBI under session PRJNA1018221. The following publicly available datasets are used in this study: piRNA reference GSE57420 and GSE122199, PDB 7KX7, 7KX9 and 5GUH. Source data are provided with this paper.
Code availability
The pipeline analyzing degradome sequencing data was deposited in GitHub (https://github.com/CMACH508/GuidewithTarget)54.
References
Bourque, G. et al. Ten things you should know about transposable elements. Genome Biol. 19, 199 (2018).
Khurana, J. S. et al. Adaptation to P element transposon invasion in Drosophila melanogaster. Cell 147, 1551–1563 (2011).
Aravin, A. A., Hannon, G. J. & Brennecke, J. The Piwi-piRNA pathway provides an adaptive defense in the transposon arms race. Science 318, 761–764 (2007).
Parhad, S. S. & Theurkauf, W. E. Rapid evolution and conserved function of the piRNA pathway. Open Biol. 9, 180181 (2019).
Malone, C. D. & Hannon, G. J. Small RNAs as guardians of the genome. Cell 136, 656–668 (2009).
Jangam, D., Feschotte, C. & Betran, E. Transposable element domestication as an adaptation to evolutionary conflicts. Trends Genet. 33, 817–831 (2017).
Levin, H. L. & Moran, J. V. Dynamic interactions between transposable elements and their hosts. Nat. Rev. Genet. 12, 615–627 (2011).
Sinzelle, L., Izsvak, Z. & Ivics, Z. Molecular domestication of transposable elements: from detrimental parasites to useful host genes. Cell. Mol. Life Sci. 66, 1073–1093 (2009).
Ozata, D. M., Gainetdinov, I., Zoch, A., O’Carroll, D. & Zamore, P. D. PIWI-interacting RNAs: small RNAs with big functions. Nat. Rev. Genet. 20, 89–108 (2019).
Aravin, A. et al. A novel class of small RNAs bind to MILI protein in mouse testes. Nature 442, 203–207 (2006).
Girard, A., Sachidanandam, R., Hannon, G. J. & Carmell, M. A. A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442, 199–202 (2006).
Grivna, S. T., Beyret, E., Wang, Z. & Lin, H. A novel class of small RNAs in mouse spermatogenic cells. Genes Dev. 20, 1709–1714 (2006).
Aravin, A. A., Sachidanandam, R., Girard, A., Fejes-Toth, K. & Hannon, G. J. Developmentally regulated piRNA clusters implicate MILI in transposon control. Science 316, 744–747 (2007).
Deniz, O., Frost, J. M. & Branco, M. R. Regulation of transposable elements by DNA modifications. Nat. Rev. Genet. 20, 417–431 (2019).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Kofler, R. piRNA clusters need a minimum size to control transposable element invasions. Genome Biol. Evol. 12, 736–749 (2020).
Waterston, R. H. et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Brennecke, J. et al. Discrete small RNA-generating loci as master regulators of transposon activity in. Cell 128, 1089–1103 (2007).
Yu, T. et al. The piRNA response to retroviral invasion of the koala genome. Cell 179, 632–643 (2019).
Rosenkranz, D., Zischler, H. & Gebert, D. piRNAclusterDB 2.0: update and expansion of the piRNA cluster database. Nucleic Acids Res. 50, D259–D264 (2022).
Matsumoto, N. et al. Crystal structure of silkworm PIWI-clade Argonaute Siwi bound to piRNA. Cell 167, 484–497 (2016).
Yamaguchi, S. et al. Crystal structure of Drosophila Piwi. Nat. Commun. 11, 858 (2020).
Anzelon, T. A. et al. Structural basis for piRNA targeting. Nature 597, 285–289 (2021).
Flores-Jasso, C. F., Salomon, W. E. & Zamore, P. D. Rapid and specific purification of Argonaute-small RNA complexes from crude cell lysates. RNA 19, 271–279 (2013).
Schirle, N. T. & MacRae, I. J. The crystal structure of human Argonaute2. Science 336, 1037–1040 (2012).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Lewis, B. P., Burge, C. B. & Bartel, D. P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Schirle, N. T., Sheu-Gruttadauria, J. & MacRae, I. J. Structural basis for microRNA targeting. Science 346, 608–613 (2014).
Wang, Y. et al. Structure of an argonaute silencing complex with a seed-containing guide DNA and target RNA duplex. Nature 456, 921–926 (2008).
Wang, Y. et al. Nucleation, propagation and cleavage of target RNAs in Ago silencing complexes. Nature 461, 754–761 (2009).
Arif, A. et al. GTSF1 accelerates target RNA cleavage by PIWI-clade Argonaute proteins. Nature 608, 618–625 (2022).
Wee, L. M., Flores-Jasso, C. F., Salomon, W. E. & Zamore, P. D. Argonaute divides its RNA guide into domains with distinct functions and RNA-binding properties. Cell 151, 1055–1067 (2012).
Gainetdinov, I. et al. Relaxed targeting rules help PIWI proteins silence transposons. Nature 619, 394–402 (2023).
Wang, W. et al. The initial uridine of primary piRNAs does not create the tenth adenine that is the hallmark of secondary piRNAs. Mol. Cell 56, 708–716 (2014).
Wang, W. et al. Slicing and binding by Ago3 or Aub trigger Piwi-bound piRNA production by distinct mechanisms. Mol. Cell 59, 819–830 (2015).
Klum, S. M., Chandradoss, S. D., Schirle, N. T., Joo, C. & MacRae, I. J. Helix-7 in Argonaute2 shapes the microRNA seed region for rapid target recognition. EMBO J. 37, 75–88 (2018).
Dai, S. et al. A family of C. elegans VASA homologs control Argonaute pathway specificity and promote transgenerational silencing. Cell Rep. 40, 111265 (2022).
De Fazio, S. et al. The endonuclease activity of Mili fuels piRNA amplification that silences LINE1 elements. Nature 480, 259–263 (2011).
Dowling, M. et al. In vivo PIWI slicing in mouse testes deviates from rules established in vitro. RNA 29, 308–316 (2023).
Vourekas, A. et al. Mili and Miwi target RNA repertoire reveals piRNA biogenesis and function of Miwi in spermiogenesis. Nat. Struct. Mol. Biol. 19, 773–781 (2012).
Gou, L. T. et al. Pachytene piRNAs instruct massive mRNA elimination during late spermiogenesis. Cell Res 24, 680–700 (2014).
Goh, W. S. et al. piRNA-directed cleavage of meiotic transcripts regulates spermatogenesis. Genes Dev. 29, 1032–1044 (2015).
Wu, P. H. et al. The evolutionarily conserved piRNA-producing locus pi6 is required for male mouse fertility. Nat. Genet. 52, 728–739 (2020).
Choi, H., Wang, Z. & Dean, J. Sperm acrosome overgrowth and infertility in mice lacking chromosome 18 pachytene piRNA. PLoS Genet. 17, e1009485 (2021).
Gou, L. T. et al. Ubiquitination-deficient mutations in human Piwi cause male infertility by impairing histone-to-protamine exchange during spermiogenesis. Cell 169, 1090–1104 (2017).
Wu, X. et al. The biogenesis and functions of piRNAs in human diseases. Mol. Ther. Nucleic Acids 21, 108–120 (2020).
Lei, J. & Frank, J. Automated acquisition of cryo-electron micrographs for single particle reconstruction on an FEI Tecnai electron microscope. J. Struct. Biol. 150, 69–80 (2005).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr D. Biol. Crystallogr 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D. Biol. Crystallogr 66, 213–221 (2010).
Schrödinger, L., & DeLano, W. PyMOL http://www.pymol.org/pymol (2020).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Lu, W. P. & Fei, L. A logarithmic approximation to initial rates of enzyme reactions. Anal. Biochem. 316, 58–65 (2003).
Zeng, L. GuidewithTarget (Degradome Sequencing Bioinformatics Pipeline). GitHub https://github.com/CMACH508/GuidewithTarget (2023).
Piuco, R. & Galante, P. A. piRNAdb: a piwi-interacting RNA database. Preprint at BioRxiv https://doi.org/10.1101/2021.09.21.461238 (2021).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinf. 10, 421 (2009).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Acknowledgements
We thank C. C. Mello, P. D. Zamore, D. Li and H. Gao for insightful suggestions; H. Yu (Westlake University) for critical proofreading of the manuscript; I. Gainetdinov and P. D. Zamore for guidance in analyzing Degradome Sequencing data, members of Shen and Wu laboratories for discussions; the Cryo-EM Facility of Westlake University for providing support on cryo-EM data collection; the Key Laboratory of Zhejiang Province for Aptamers and Theranostics for their technical support in the SPR assay and the Mass Spectrometry and Metabolomics Core Facility of Westlake University for protein analysis. This work was supported by Westlake Education Foundation, Zhejiang Provincial Foundation of China (2021R01013) and National Natural Science Foundation of China (NSFC32070628) to E.-Z.S., National Natural Science Foundation of China (32271261), Zhejiang Provincial Natural Science Foundation of China (LR22C050003), Westlake University (1011103860222B1) and Institutional Startup Grant from the Westlake Education Foundation (101486021901) to J.W. and the National Natural Science Foundation of China (NO. 92068103) and Westlake Laboratory of Life Science to C.-Q.S.
Author information
Authors and Affiliations
Contributions
E.-Z.S. conceptualized the study. Zhiqing Li, Zhenzhen Li, Y.Z., L. Zhou, Q.X., L.L., L. Zeng, J.X., H.N., J.Z., Q.Y., D.L., M.G., Y.H., S.T., Z.Z., C.-Q.S., J.W. and E.-Z.S. performed the investigations. E.-Z.S. wrote the original draft. Z.Z., C.-Q.S., J.W. and E.-Z.S. reviewed and edited the manuscript.C.-Q.S., J.W. and E.-Z.S. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Claus Kuhn and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Sara Osman was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Retrotransposon and piRNA cluster in invertebrates and mammals.
a,b, Ratio of retrotransposon (a) and piRNA cluster (b) of invertebrate and mammalian genomes.
Extended Data Fig. 2 PIWI family tree and preparation of Piwi-piRNA complexes.
a, Phylogram of the PIWI proteins shows that MILI, HILI, and EfPiwi proteins belong to the same branch. Branch lengths are shown above the line. b,c, Size-exclusion chromatography profile of the purified MILI-piRNA complex (arrow) (b) and HILI-piRNA complex (arrow) (c) using a modified oligo capture approach. d,e, SDS-PAGE analysis of the purified MILI-piRNA complex (d) and HILI-piRNA complex (e). Gels were stained with Coomassie blue (representative of protein purifications performed at least ten times independently). f,g, Guide RNA-mediated cleavage assay with MILI (f) or HILI (g). MILI or HILI loaded with or without synthetic guide RNAs were incubated with 5′-FAM-labeled fully matched (FM) target RNAs or non-complementary (NC) target RNAs for 1 h at 37 °C. Products were resolved on a denaturing urea-PAGE gel alongside synthetic 22-nt marker and 1-nt RNA ladder. Similar results are seen in three experiments. h, piRNA and target RNA sequences used in f, g. The fluorescein amidite (FAM) modification in the target strand is highlighted in green, and the cleavage site is indicated with a red arrow. The cleaved 5′ fragment with FAM label is 24 nucleotides.
Extended Data Fig. 3 EMSA of piRISC binding to target RNAs and kinetics of Ago2-miRNA target binding.
a, Representative PAGE of the protein-guide RNA complexes binding to another fully matched (FM) target RNA. Non-complementary (NC) target is a negative control. Similar results are seen in two experiments. b, The guide and target RNAs used in this experiment. c, Equilibrium dissociation constant (KD) of g2-g8 complementary target for protein-guide RNA complexes. Data are mean ± s.d. from independent triplicates. d, SPR traces showing binding kinetics of g2-g8 complementary target for Ago2-miRNA binary complex. e, SPR sensorgrams of non-complementary target RNA binding to Ago2-miRNA complex.
Extended Data Fig. 4 Cryo-EM analysis of MILI-piRNA, MILI-piRNA-target (15 nt).
a,b, Data processing workflow of cryo-EM analysis of MILI-piRNA (a) and MILI-piRNA-target (15 nt) (b). Representative motion-corrected dose-weighted micrographs were selected out of 4,602 and 8,107 micrographs for the two datasets, respectively. Scale bar of the micrographs: 50 nm. Box size of 2D class averages: 209 Å. Local resolution, angular distribution, gold-standard FSC curves of the final maps (resolution cut-off at FSC = 0.143), and model vs. map curves of the final maps against the final models (resolution cut-off at FSC = 0.5) were presented. Please note that an open state was also observed during 2D and 3D classifications in the MILI-piRNA-target (15 nt) dataset.
Extended Data Fig. 5 Cryo-EM analysis of HILI-piRNA.
Data processing workflow for HILI-piRNA cryo-EM analysis. Representative motion-corrected dose-weighted micrograph was selected out of 2,136 micrographs. Scale bar of the micrograph: 50 nm. Box size of 2D class averages: 209 Å. Local resolution, angular distribution, gold-standard FSC curves of the final maps (resolution cut-off at FSC = 0.143), and model vs. map curves of the final map against the final model (resolution cut-off at FSC = 0.5) were presented. The density of the bound piRNA in the selected 3D class (with a dotted box) was indicated by a red arrow.
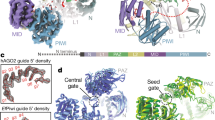
Extended Data Fig. 6 Structural comparison of individual domains of MILI, HILI, EfPiwi and Siwi, and electron density map as well as structural basis for piRNA targeting in the non-seed region.
a, Superposition of the individual domains of MILI (pink), HILI (orange), EfPiwi (PDB: 7KX7) (aquamarine), and Siwi (PDB: 5GUH) (chartreuse). b, The density map (grey mesh) for the entire bound piRNA in MILI–piRISC is shown in the left panel. A close-up view of the EM density map (grey mesh) for the piRNA 5′ segment is shown in the right panel. Modeled guide RNAs are shown as red sticks. c, The density map (grey mesh) for the entire bound piRNA in HILI-piRISC is shown in the left panel. Two close-up views of the EM density map (grey mesh) for the piRNA 5′ segment and 3′ half are shown in the middle and right panels. Modeled guide RNAs are shown as red sticks. d,e, Non-seed interactions between piRNA-target RNA duplex and EfPiwi protein (d) and MILI protein (e). f, Relative target-binding ratios for MILI, HILI, and EfPiwi. The relative ratio here is mismatched target binding % over fully matched target binding %. Data are from independent duplicates.
Extended Data Fig. 7 Effect of GTSF1 on slicer activity of PIWI proteins and biochemical determination of target binding, target cleavage and mismatch tolerance of Siwi.
a, Coomassie blue-stained SDS-PAGE of purified recombinant Flag-tagged GTSF1 proteins from different species (representative of protein purifications performed three times independently). b, piRISC target cleavage assays in the presence of increasing amounts of GTSF1. Em, Ephydatia muelleri; Bm, Bombyx mori; Mm, Mus musculus; Hs, Homo sapiens. Note: The Ephydatia fluviatilis genome sequence is unavailable, so EfGFST1 remains unidentified. GTSF1 from the related sponge Ephydatia muelleri was therefore incubated with EfPiwi. Similar results are seen in three experiments. c, Representative native PAGE analysis of protein-guide RNA complexes binding to fully matched (FM) target RNA. NC, non-complementary (NC) target negative control. Similar results are seen in three experiments. d, Representative in vitro cleavage reactions in the presence of equimolar substrate (target RNA) and enzyme (Piwi-piRNA complexes). Similar results are seen in three experiments. e, Guide-target-pairing schematic for targets with 3-nt mismatch regions to guide RNA used in (f, g). f, Representative urea-PAGE showing the ability of EfPiwi, Siwi, MILI, and HILI to cleave targets with three consecutive mismatches between g2 and g22. Similar results are seen in two experiments. g, Quantification of the in vitro cleavage assays (right) showing changes in cleavage activity for three consecutive mismatches between g2 and g22.
Extended Data Fig. 8 Sequence alignment of vertebrate Ago3-like PIWI proteins.
The secondary structure of MmMILI is indicated above the sequences, color coded by domains as indicated in Fig. 2. Key residues are marked by triangles. Species abbreviations: Mm, Mus musculus (mouse); Hs, Homo sapiens (human); Mm, Macaca mulatta (macaque); Ec, Equus caballus (horse); Dr, Danio rerio (zebrafish); Xt, Xenopus tropicalis (frog); Ef, Ephydatia fluviatilis (sponge); Hv, Hydra vulgaris (hydra); Bm, Bombyx mori (silkworm); Dm, Drosophila melanogaster (fruit fly); Ba, Bicyclus anynana (wood nymph); So, Sitophilus oryzae (rice weevil).
Extended Data Fig. 9 Cryo-EM analysis of EfPiwiN959K-piRNA, EfPiwiN959K-piRNA-target (16 nt).
a,b, Data processing workflow of cryo-EM analysis of EfPiwiN959K-piRNA (a) and EfPiwiN959K-piRNA-target (16 nt) (b). Representative motion-corrected dose-weighted micrographs were selected out of 2,279 and 3,983 micrographs for the two datasets, respectively. Scale bar of the micrographs: 50 nm. Box size of 2D class averages: 209 Å. Local resolution, angular distribution, gold-standard FSC curves of the final maps (resolution cut-off at FSC = 0.143), and model vs. map curves of the final maps against the final models (resolution cut-off at FSC = 0.5) were presented.
Extended Data Fig. 10 Target binding and cleavage data by EfPiwi, MILI, HILI and Ago2 in vitro and analysis of Drosophila Ago3-catalyzed splicing of transcripts in vivo.
a, Guide-target pairing schematic for targets with increasing complementarity for target binding analysis in (b). b, Equilibrium dissociation constants (KD) of Argonaute-guide complexes binding target RNAs with increasing complementarity. Data points from biological duplicates are shown. c, Guide-target pairing schematic for targets with increasing complementarity for cleavage analysis in (d, e). d, Quantification of reaction products shown as fraction of target cleaved. Data are from two independent experiments. e, Representative urea-PAGE showing EfPiwi-, MILI-, HILI-, and Ago2-mediated cleavage of targets with increasing guide complementarity. Similar results are seen in two experiments. f, Schema of data preparation and preprocessing to identify Ago3-mediated cleavage products in Drosophila ovaries. Ago3-bound piRNAs (Ago3_IP.fa) and non-associated piRNA (Ago3_bg.fa) were identified and compared to potential 3′ cleavage products whose abundance decreased in mutant ago3 degradome sequencing libraries (mutago3; SRR1926158) compared to wild-type ago3 degradome sequencing libraries (wtago3; SRR1926188). See Methods for details. g, piRNA queries (Ago3_IP.fa/Ago3_bg.fa) were aligned to candidate targets whose cleaved 5′ ends are complementary to g2–g10 of a piRNA. Targets with extended pairing were identified from g2–g25. For each pairing arrangement g2–gX, the fraction of cleaved targets was calculated as the fraction of cleaved targets explained by Ago3-associated piRNAs minus the fraction explained by Ago3-non-associated piRNAs. h, Plot of the fraction of cleaved targets in fly ovaries for contiguous pairing from nucleotide g2–gX, as indicated. Ago3-non-associated piRNA were sampled 10 times.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2, Figs. 1–3 and Methods.
Source data
Source Data Fig. 1
Unprocessed gel images.
Source Data Fig. 3
Unprocessed gel images.
Source Data Fig. 4
Unprocessed gel images.
Source Data Fig. 5
Unprocessed gel images.
Source Data Extended Data Fig. 2
Unprocessed gel images.
Source Data Extended Data Fig. 6
Unprocessed gel images.
Source Data Extended Data Fig. 7
Unprocessed gel images.
Source Data Extended Data Fig. 10
Unprocessed gel images.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Z., Li, Z., Zhang, Y. et al. Mammalian PIWI–piRNA–target complexes reveal features for broad and efficient target silencing. Nat Struct Mol Biol (2024). https://doi.org/10.1038/s41594-024-01287-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41594-024-01287-6