Abstract
The zebrafish is a powerful model system for studying animal development, for modeling genetic diseases, and for large-scale in vivo functional genetics. Because of its ease of use and its high efficiency in targeted gene perturbation, CRISPR–Cas9 has recently gained prominence as the tool of choice for genetic manipulation in zebrafish. However, scaling up the technique for high-throughput in vivo functional genetics has been a challenge. We recently developed a method, Multiplexed Intermixed CRISPR Droplets (MIC-Drop), that makes large-scale CRISPR screening in zebrafish possible. Here, we outline the step-by-step protocol for performing functional genetic screens in zebrafish by using MIC-Drop. MIC-Drop uses multiplexed single-guide RNAs to generate biallelic mutations in injected zebrafish embryos, allowing genetic screens to be performed in F0 animals. Combining microfluidics and DNA barcoding enables simultaneous targeting of tens to hundreds of genes from a single injection needle, while also enabling retrospective and rapid identification of the genotype responsible for an observed phenotype. The primary target audiences for MIC-Drop are developmental biologists, zebrafish geneticists, and researchers interested in performing in vivo functional genetic screens in a vertebrate model system. MIC-Drop will also prove useful in the hands of chemical biologists seeking to identify targets of small molecules that cause phenotypic changes in zebrafish. By using MIC-Drop, a typical screen of 100 genes can be conducted within 2–3 weeks by a single user.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data pertaining to this paper are included in figures and tables. Raw data for Fig. 4 are provided as Source Data. Please direct reasonable material requests to R.T.P. Source data are provided with this paper.
References
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Adli, M. The CRISPR tool kit for genome editing and beyond. Nat. Commun. 9, 1911 (2018).
Doudna, J. A. & Charpentier, E. The new frontier of genome engineering with CRISPR-Cas9. Science 346, 1258096 (2014).
Sander, J. D. & Joung, J. K. CRISPR-Cas systems for editing, regulating and targeting genomes. Nat. Biotechnol. 32, 347–355 (2014).
Shalem, O., Sanjana, N. E. & Zhang, F. High-throughput functional genomics using CRISPR–Cas9. Nat. Rev. Genet. 16, 299–311 (2015).
Bock, C. et al. High-content CRISPR screening. Nat. Rev. Methods Prim. 2, 8 (2022).
Joung, J. et al. Genome-scale CRISPR-Cas9 knockout and transcriptional activation screening. Nat. Protoc. 12, 828–863 (2017).
Feldman, D. et al. Pooled genetic perturbation screens with image-based phenotypes. Nat. Protoc. 17, 476–512 (2022).
Jost, M. & Weissman, J. S. CRISPR approaches to small molecule target identification. ACS Chem. Biol. 13, 366–375 (2018).
Neggers, J. E. et al. Target identification of small molecules using large-scale CRISPR-Cas mutagenesis scanning of essential genes. Nat. Commun. 9, 502 (2018).
Kuhn, M., Santinha, A. J. & Platt, R. J. Moving from in vitro to in vivo CRISPR screens. Gene Genome Ed. 2, 100008 (2021).
Shin, U. et al. Large-scale generation and phenotypic characterization of zebrafish CRISPR mutants of DNA repair genes. DNA Repair 107, 103173 (2021).
Pei, W. et al. Guided genetic screen to identify genes essential in the regeneration of hair cells and other tissues. NPJ Regen. Med. 3, 11 (2018).
Varshney, G. K. et al. High-throughput gene targeting and phenotyping in zebrafish using CRISPR/Cas9. Genome Res. 25, 1030–1042 (2015).
Varshney, G. K. et al. A high-throughput functional genomics workflow based on CRISPR/Cas9-mediated targeted mutagenesis in zebrafish. Nat. Protoc. 11, 2357–2375 (2016).
Thyme, S. B. et al. Phenotypic landscape of schizophrenia-associated genes defines candidates and their shared functions. Cell 177, 478–491.e20 (2019).
Sun, Y. et al. Systematic genome editing of the genes on zebrafish Chromosome 1 by CRISPR/Cas9. Genome Res. 30, 118–126 (2019).
Keatinge, M. et al. CRISPR gRNA phenotypic screening in zebrafish reveals pro-regenerative genes in spinal cord injury. PLOS Genet. 17, e1009515 (2021).
Parvez, S. et al. MIC-Drop: a platform for large-scale in vivo CRISPR screens. Science 373, 1146–1151 (2021).
Patton, E. E. & Zon, L. I. The art and design of genetic screens: zebrafish. Nat. Rev. Genet. 2, 956–966 (2001).
Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 496, 498–503 (2013).
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. & Schilling, T. F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995).
Driever, W. et al. A genetic screen for mutations affecting embryogenesis in zebrafish. Development 123, 37–46 (1996).
Haffter, P. et al. The identification of genes with unique and essential functions in the development of the zebrafish, Danio rerio. Development 123, 1–36 (1996).
Eisen, J. S. Zebrafish make a big splash. Cell 87, 969–977 (1996).
Poss, K. D., Wilson, L. G. & Keating, M. T. Heart regeneration in zebrafish. Science 298, 2188–2190 (2002).
Gemberling, M., Bailey, T. J., Hyde, D. R. & Poss, K. D. The zebrafish as a model for complex tissue regeneration. Trends Genet. 29, 611–620 (2013).
Marques, I. J., Lupi, E. & Mercader, N. Model systems for regeneration: zebrafish. Development 146, dev167692 (2019).
Meshalkina, D. A. et al. Understanding zebrafish cognition. Behav. Process. 141, 229–241 (2017).
Gerlai, R. Evolutionary conservation, translational relevance and cognitive function: the future of zebrafish in behavioral neuroscience. Neurosci. Biobehav. Rev. 116, 426–435 (2020).
Kalueff, A. V. et al. Towards a comprehensive catalog of zebrafish behavior 1.0 and beyond. Zebrafish 10, 70–86 (2013).
Geng, Y. & Peterson, R. T. The zebrafish subcortical social brain as a model for studying social behavior disorders. Dis. Model. Mech. 12, dmm039446 (2019).
Bahl, A. & Engert, F. Neural circuits for evidence accumulation and decision making in larval zebrafish. Nat. Neurosci. 23, 94–102 (2020).
Nelson, J. C. & Granato, M. Zebrafish behavior as a gateway to nervous system assembly and plasticity. Development 149, dev177998 (2022).
McConnell, A. M., Noonan, H. R. & Zon, L. I. Reeling in the zebrafish cancer models. Annu. Rev. Cancer Biol. 5, 331–350 (2021).
White, R., Rose, K. & Zon, L. Zebrafish cancer: the state of the art and the path forward. Nat. Rev. Cancer 13, 624–636 (2013).
Zhang, T. & Peterson, R. T. Modeling lysosomal storage diseases in the zebrafish. Front. Mol. Biosci. 7, 82 (2020).
Campbell, P. D. & Granato, M. Zebrafish as a tool to study schizophrenia-associated copy number variants. Dis. Model. Mech. 13, dmm043877 (2020).
Grone, B. P. & Baraban, S. C. Animal models in epilepsy research: legacies and new directions. Nat. Neurosci. 18, 339–343 (2015).
Basu, S. & Sachidanandan, C. Zebrafish: a multifaceted tool for chemical biologists. Chem. Rev. 113, 7952–7980 (2013).
Hwang, W. Y. et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat. Biotechnol. 31, 227–229 (2013).
Burger, A. et al. Maximizing mutagenesis with solubilized CRISPR-Cas9 ribonucleoprotein complexes. Development 143, 2025–2037 (2016).
Gagnon, J. A. et al. Efficient mutagenesis by Cas9 protein-mediated oligonucleotide insertion and large-scale assessment of single-guide RNAs. PLoS One 9, e98186 (2014).
Shah, A. N., Davey, C. F., Whitebirch, A. C., Miller, A. C. & Moens, C. B. Rapid reverse genetic screening using CRISPR in zebrafish. Nat. Methods 12, 535–540 (2015).
Jao, L.-E., Wente, S. R. & Chen, W. Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system. Proc. Natl Acad. Sci. USA 110, 13904–13909 (2013).
Wu, R. S. et al. A rapid method for directed gene knockout for screening in G0 zebrafish. Dev. Cell 46, 112–125.e4 (2018).
Kroll, F. et al. A simple and effective F0 knockout method for rapid screening of behaviour and other complex phenotypes. eLife 10, e59683 (2021).
Hoshijima, K. et al. Highly efficient CRISPR-Cas9-based methods for generating deletion mutations and F0 embryos that lack gene function in zebrafish. Dev. Cell 51, 645–657.e4 (2019).
Long, L. et al. Regulation of transcriptionally active genes via the catalytically inactive Cas9 in C. elegans and D. rerio. Cell Res. 25, 638–641 (2015).
Dong, X. et al. Zebrafish Znfl1 proteins control the expression of hoxb1b gene in the posterior neuroectoderm by acting upstream of pou5f3 and sall4 genes. J. Biol. Chem. 292, 13045–13055 (2017).
Ghanta, K. S., Ishidate, T. & Mello, C. C. Microinjection for precision genome editing in Caenorhabditis elegans. STAR Protoc. 2, 100748 (2021).
Stepicheva, N. A. & Song, J. L. High throughput microinjections of sea urchin zygotes. J. Vis. Exp. 2014, e50841 (2014).
Peng, P. et al. CRISPR-Cas9 mediated genome editing in Drosophila. Bio. Protoc. 9, e3141 (2019).
Nakayama, T. et al. Cas9-based genome editing in Xenopus tropicalis. Methods Enzymol. 546, 355–375 (2014).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013).
Schubert, S., Keddig, N., Hanel, R. & Kammann, U. Microinjection into zebrafish embryos (Danio rerio) - a useful tool in aquatic toxicity testing? Environ. Sci. Eur. 26, 32 (2014).
McKee, R. A. & Wingert, R. A. Nephrotoxin microinjection in zebrafish to model acute kidney injury. J. Vis. Exp. 2016, 10.3791/54241 (2016).
Amsterdam, A. et al. A large-scale insertional mutagenesis screen in zebrafish. Genes Dev. 13, 2713–2724 (1999).
Gaiano, N. et al. Insertional mutagenesis and rapid cloning of essential genes in zebrafish. Nature 383, 829–832 (1996).
Amsterdam, A. et al. Identification of 315 genes essential for early zebrafish development. Proc. Natl Acad. Sci. USA 101, 12792–12797 (2004).
Golling, G. et al. Insertional mutagenesis in zebrafish rapidly identifies genes essential for early vertebrate development. Nat. Genet. 31, 135–140 (2002).
Fuentes, R., Letelier, J., Tajer, B., Valdivia, L. E. & Mullins, M. C. Fishing forward and reverse: advances in zebrafish phenomics. Mech. Dev. 154, 296–308 (2018).
Sander, J. D. et al. Targeted gene disruption in somatic zebrafish cells using engineered TALENs. Nat. Biotechnol. 29, 697–698 (2011).
Huang, P. et al. Heritable gene targeting in zebrafish using customized TALENs. Nat. Biotechnol. 29, 699–700 (2011).
Doyon, Y. et al. Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases. Nat. Biotechnol. 26, 702–708 (2008).
Foley, J. E. et al. Rapid mutation of endogenous zebrafish genes using zinc finger nucleases made by Oligomerized Pool ENgineering (OPEN). PLoS One 4, e4348 (2009).
Bradford, Y. M. et al. Zebrafish information network, the knowledgebase for Danio rerio research. Genetics 220, iyac016 (2022).
Mullins, M. C., Acedo, J. N., Priya, R., Solnica-Krezel, L. & Wilson, S. W. The zebrafish issue: 25 years on. Development 148, dev200343 (2021).
Jin, X. et al. In vivo Perturb-Seq reveals neuronal and glial abnormalities associated with autism risk genes. Science 370, eaaz6063 (2020).
Trubiroha, A. et al. A rapid CRISPR/Cas-based mutagenesis assay in zebrafish for identification of genes involved in thyroid morphogenesis and function. Sci. Rep. 8, 5647 (2018).
Klatt Shaw, D. & Mokalled, M. H. Efficient CRISPR/Cas9 mutagenesis for neurobehavioral screening in adult zebrafish. G3 (Bethesda) 11, jkab089 (2021).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Moreno-Mateos, M. A. et al. CRISPRscan: designing highly efficient sgRNAs for CRISPR-Cas9 targeting in vivo. Nat. Methods 12, 982–988 (2015).
Concordet, J.-P. & Haeussler, M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 46, W242–W245 (2018).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Quick, R. E., Buck, L. D., Parab, S., Tolbert, Z. R. & Matsuoka, R. L. Highly efficient synthetic CRISPR RNA/Cas9-based mutagenesis for rapid cardiovascular phenotypic screening in F0 zebrafish. Front. Cell Dev. Biol. 9, 735598 (2021).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Košir, A. B. et al. Droplet volume variability as a critical factor for accuracy of absolute quantification using droplet digital PCR. Anal. Bioanal. Chem. 409, 6689–6697 (2017).
Acknowledgements
We are grateful to the Centralized Zebrafish Animal Resource (CZAR) for providing animal husbandry, maintenance and microinjection equipment. We acknowledge support from the NIH (5R01GM134069-02 to R.T.P.), an American Heart Association postdoctoral fellowship to S.P. and a T32 training grant (T32HG008962) to Z.J.B.
Author information
Authors and Affiliations
Contributions
S.P., Z.J.B. and R.T.P. contributed to developing the protocol and writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
A patent covering the MIC-Drop platform, on which S.P. and R.T.P are listed as inventors, is pending.
Peer review
Peer review information
Nature Protocols thanks Thomas Becker, Marcus Keatinge, Adam Miller, Andy Willaert and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Parvez, S. et al. Science 373, 1146–1151 (2021); https://doi.org/10.1126/science.abi8870
Supplementary information
Supplementary Information
Supplementary Figs. 1–3
Supplementary Table 1
sgRNA target sequences and primers used in the protocol. sgRNA target/spacer sequences of control scrambled sgRNAs, sgRNAs targeting rx3 and tbx16 genes and primers used for barcode generation and barcode amplification.
Droplet generation using a QX200 droplet generator. Droplets targeting up to eight genes are generated during a single run of the droplet generator. A single run takes ~2 min.
Transfer of MIC-Drops into a microloader tip. Colored droplets (used as proxies for droplets targeting different genes) being loaded into a microloader tip.
Trimming of the microinjection needle for droplet injection. After transferring droplets to a microinjection needle, the needle is trimmed to the desired width. Excess oil is cleared out until the first droplet reaches the tip of the injection needle.
Injection of MIC-Drops in zebrafish embryos. The first 10–30 droplets are separated by a larger volume of 3% (wt/vol) FS-HFE and require several pushes of the foot pedal (5–10 pushes) to clear out the excess oil.
Injection of MIC-Drops in zebrafish embryos. After the first 10–30 droplets have been injected, the remaining droplets are closer together and require only two or three pushes of the foot pedal to clear out the excess oil.
Injection of droplets containing phenol red. In the previous videos, food coloring is used in the droplets for better visualization. Food coloring may be toxic to developing embryos and should not be used in experiments.
Video illustrating accidental injection of an oil droplet in an embryo. Although injection of a small volume of oil is non-consequential to embryo development, it is best to avoid injecting oil.
Clearing a clogged needle. Occasionally, the injection needle can get clogged. The needle can be unclogged by pressing the ‘Clear’ button or trimming the needle.
Source data
Source Data Fig. 4d
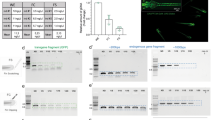
Uncropped gel for Fig. 4d
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Parvez, S., Brandt, Z.J. & Peterson, R.T. Large-scale F0 CRISPR screens in vivo using MIC-Drop. Nat Protoc 18, 1841–1865 (2023). https://doi.org/10.1038/s41596-023-00821-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00821-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.