Abstract
Intestinal stem cells (ISCs) of the fruit fly, Drosophila melanogaster, offer an excellent genetic model to explore homeostatic roles of ISCs in animal physiology. Among available genetic tools, the escargot (esg)-GAL4 driver, expressing the yeast transcription factor gene, GAL4, under control of the esg gene promoter, has contributed significantly to ISC studies. This driver facilitates activation of genes of interest in proximity to a GAL4-binding element, Upstream Activating Sequence, in ISCs and progenitor enteroblasts (EBs). While esg-GAL4 has been considered an ISC/EB-specific driver, recent studies have shown that esg-GAL4 is also active in other tissues, such as neurons and ovaries. Therefore, the ISC/EB specificity of esg-GAL4 is questionable. In this study, we reveal esg-GAL4 expression in the corpus allatum (CA), responsible for juvenile hormone (JH) production. When driving the oncogenic gene, RasV12, esg-GAL4 induces overgrowth in ISCs/EBs as reported, but also increases CA cell number and size. Consistent with this observation, animals alter expression of JH-response genes. Our data show that esg-GAL4-driven gene manipulation can systemically influence JH-mediated animal physiology, arguing for cautious use of esg-GAL4 as a “specific” ISC/EB driver to examine ISC/EB-mediated animal physiology.
Similar content being viewed by others
Introduction
Precise overexpression of genes in specific cell types and time windows is crucial to discover essential functions of those genes in multicellular organisms. Among model organisms, such gene expression manipulation techniques are best developed for the fruit fly, Drosophila melanogaster. In particular, the GAL4-UAS system is a powerful binary gene expression system in D. melanogaster for targeted genetic manipulation in a spatio-temporal specific manner to reveal gene functions1. This system utilizes the yeast transcription factor, GAL4, controlled by a tissue-specific enhancer/promoter sequence, in combination with a GAL4-biding element called Upstream Activating Sequence (UAS), inserted upstream of the gene of interest, either endogenously or exogenously. The impact of the GAL4-UAS system on D. melanogaster genetics research is immeasurable. However, despite its utility, a potential drawback of the GAL4-UAS system is the possibility of incomplete cell type- or tissue-specific expression patterns, complicating interpretation of results.
D. melanogaster escargot (esg)-GAL4, formally known as P{GawB}NP5130 (RRID:BDSC_93857)2, has widely been used as the fundamental GAL4 driver to manipulate genes “specifically” in intestinal stem cells (ISCs) and enteroblasts (EBs) (Fig. 1a). In D. melanogaster, ISCs regulate gut homeostasis by maintaining themselves and also by giving rise to other essential gut epithelial cells, including EBs, enteroendocrine cells, and enterocytes. Dysfunction of ISCs results in severe malfunctions of age-associated tissue integrity in the gut3. By virtue of convenient tools to analyze functions and roles of genes, D. melanogaster ISCs have served as a useful model system to study the homeostatic role of ISCs in gut physiology. Notably, the esg-GAL4 driver has facilitated overexpression of genes and various constructs to study fundamental roles of ISCs and EBs. For example, researchers have heavily used esg-GAL4 to generate ISC tumors by overexpressing oncogenic genes such as the gain-of-function transgenes, Ras and yokie (yki). esg-GAL4-driven ISC tumor animals have advanced our understanding of tumor-dependent impairment of systemic physiology, such as cachexia and the bloating phenotype. The crucial assumption for interpreting results of these studies as a phenotype originating from ISCs and EBs is that esg-GAL4 manipulates gene expression only in these cells in adults. However, recent studies reported that esg-GAL4 is expressed at least in some brain neurons4, ovaries5,6, and Malpighian tubules7, but not in muscles or fat bodies4. Therefore, the ISC/EB-specificity of esg-GAL4 has been questioned.
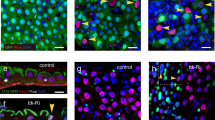
The transcription factor encoding gene, esg, was expressed in the CA. (a) (Left) GFP (green) driven by esg-GAL4 with tubP-GAL80ts (esgts >) was expressed in a subpopulation of adult midgut cells. (Right) magnified view of the area enclosed by the white square in the left figure. Blue is the DAPI signal. (b) esg > GFP, mCD8::GFP expression in whole bodies of adult males and females. Arrowheads and arrows indicate the CA and midguts, respectively. (c) esgts > GFP (green) was expressed not only in some midgut cells, but also in the CA (arrowhead). This sample was derived from a female. The CA was labeled with anti-JHAMT antibody (magenta). Two inset images correspond to a region marked with a dashed line surrounding the brain, CA, and proventriculus. Both esgts > GFP and anti-JHAMT immunoreactive signals were observed in the CA. (d) GFP (green) driven by esg-GAL4 labeled the CA in both adult males (upper) and females (lower). The CA was labeled with anti-JHAMT antibody (magenta). (e) GFP (green) driven by esg-GAL4 labeled the CA and a part of the prothoracic gland (PG) in wandering 3rd-instar (WL3) larva. The CA was labeled with anti-JHAMT antibody (magenta) and the PG was labeled with anti-Shroud (Sro) antibody (blue). (f) (Left) esg-knock in-GFP (green) was expressed in a subpopulation of adult midgut cells. (Right) A magnified view of the area encircled with a white line in the left figure. Blue is the DAPI signal. (g) esg-knock in-GFP (esg-GFP, green) was expressed in the CA in both males (upper) and females (lower). The CA was labeled with anti-JHAMT antibody (magenta). (h) esg-knock in-GFP (green) was expressed in the CA in WL3 larvae. The CA was labeled with anti-JHAMT antibody (magenta) and the PG was labeled with anti-Sro antibody (blue).
In this study, we report that esg-GAL4 is also expressed in the insect endocrine organ, the corpus allatum (CA), which is essential for synthesizing insect juvenile hormone (JH). Our data show that esg-GAL4-driven gene manipulation can systemically influence JH-mediated animal physiology, arguing for cautious use of esg-GAL4 as a “specific” ISC/EB driver to examine ISC/EB-mediated animal physiology.
Result
Esg-GAL4 is expressed in the endocrine corpus allatum
We conducted experiments using the esg-GAL4 driver combined with tubulin promoter-driven temperature-sensitive GAL80 (tubP-GAL80ts). Hereafter, esg-GAL4; tubP-GAL80ts is designated “esgts-GAL4” or “esgts > ”. This strain has widely been used for adult stage-specific gene manipulation in ISCs and EBs5,8,9,10. In all experimental conditions in this study, we reared esgts > flies at a permissive temperature (21 °C) during development, such that esg-GAL4 activity is suppressed by GAL80 right before eclosion. Then, after eclosion, we subjected these flies to a restrictive temperature (29 °C) to activate esg-GAL4 only in the adult stage.
We realized by chance that esg-GAL4 was active in the tissue located at the most anterior part of the thorax in both males and females (Fig. 1b). More precisely, esgts-GAL4-positive tissue was observed between the brain and proventriculus (Fig. 1c). This tissue was co-immunostained with an antibody against juvenile hormone acid O-methyltransferase (JHAMT), the essential enzyme that synthesizes JH in the CA11,12. This result strongly indicates that esgts-GAL4-positive tissue between the brain and proventriculus is the CA. We also confirmed that esgts-GAL4 was expressed in the CA of both male and female adult flies (Fig. 1d). Moreover, esgts-GAL4 was expressed in the ring gland, particularly in the CA of wandering 3rd-instar larvae, as well as of adults (Fig. 1e). These results suggest that esgts-GAL4 labels the CA in both male and female larvae and adults.
To confirm whether the esg gene itself is expressed in the CA, we used the esg-knock-in-GFP (esg-GFP) line13. As with esgts > GFP expression, esg-GFP was expressed not only in a certain cells in the midgut, which seem to be ISCs/EBs (Fig. 1f)13, but also in the CA of both adult males and females (Fig. 1g). Furthermore, esg-GFP was expressed in the CA of wandering 3rd-instar larvae, while considerable expression of esg-GFP was also detected in other ring gland cells (Fig. 1h). These results suggest that esg is endogenously expressed in the CA.
RNAi of JH-biosynthetic enzyme by esg-GAL4 also impairs oogenesis
Next, we explored the possibility that esgts-GAL4-driven transgenic RNAi suppresses gene expression in the CA. To examine this point, we conducted an RNAi experiment with esgts-GAL4 to target jhamt, which is expressed explicitly in the CA12. Immunostaining signals of anti-JHAMT antibody were drastically decreased by jhamt RNAi, compared to controls (Fig. 2a,b; Supplementary Table S1).
jhamt-RNAi driven by esg-GAL4 inhibited jhamt in the CA and reduced the number of eggs in the ovary. (a) Immunostaining signal of anti-JHAMT (magenta) in the CA was drastically decreased by jhamt-RNAi, driven by esg-GAL4. Upper panels are controls (esgts > GFP) and lower panels are jhamt-RNAi (esgts > GFP, jhamt-IRKK) flies. The CA is encircled with a dashed line. (b) Quantification of anti-JHAMT antibody immunostaining signals normalized by the GFP signal between control and jhamt-RNAi flies. (c) Ovaries of control (upper: esgts > GFP) and jhamt-RNAi (lower: esgts > GFP, jhamt-IRKK) virgin females. (d) Numbers of mature eggs in virgin females were significantly decreased by adult-specific jhamt-RNAi driven by esg-GAL4. For each experimental condition, 20 adult females were used. The Wilcoxon rank sum test was used for these data. *P < 0.05, **P < 0.01, ***P < 0.001.
In many insects, including D. melanogaster, JH promotes ovarian development by accumulating yolk components such as yolk protein and vitellogenin14,15. In D. melanogaster, a previous study reported that loss of jhamt activity results in smaller ovaries and reduced egg numbers16. Therefore, we observed ovary morphology and counted numbers of mature eggs in adult females expressing esgts > jhamt RNAi. We found that esgts > jhamt RNAi flies had smaller ovaries than controls (Fig. 2c). Consistent with this observation, the number of mature eggs was significantly decreased in RNAi flies (Fig. 2d; Supplementary Table S1). These results suggest that esgts-GAL4-driven RNAi suppresses gene expression in the CA and influences JH-mediated biological events such as oogenesis.
Oncogenic Ras V12 expression by esg-GAL4 causes CA hypertrophy and abnormal expression of JH-responsive genes
In some recent studies, esgts-GAL4 and UAS-RasV12 have been utilized to induce ISC/EB tumors to investigate cell turnover in the midgut and tumor-mediated systemic physiology8,17,18. Since esg-GAL4 is also expressed in the CA, esgts > RasV12 might affect both ISC/EB and CA cells. Notably, esgts > RasV12 resulted not only in abnormal expansion of the esgts-GAL4-driven GFP-positive area in the midgut (Fig. 3a)17, but also increased CA size and cell number (Fig. 3b–d; Supplementary Table S2).
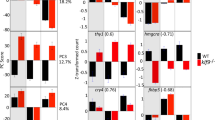
RasV12 driven by esg-GAL4 caused hypertrophy of the CA. (a) Ectopic expression of RasV12 induced overproliferation of esg-GAL4 positive midgut cells (GFP: green). (b) The CA of control (esgts > GFP) adult virgin females (upper) and RasV12 overexpression (esgts > GFP, RasV12) adult virgin females (lower). The same laser power and software gain were used to image intestinal or CA fluorescence, regardless of genotype. (c) The CA was enlarged by adult-specific RasV12 overexpression. (d) Numbers of CA cells were increased by adult-specific RasV12 overexpression. (e–g) Expression levels of JH-responsive genes (e: Kr-h1, f: Jon25Bii, g: Obp99b) were significantly changed by adult-specific RasV12 overexpression. N means sample size. The Wilcoxon rank sum test was used for (c,d). Student’s t-test was used for (e–g). *P < 0.05, **P < 0.01, ***P < 0.001. (h) Aug21 > GFP, mCD8::GFP (green) was expressed in the CA (arrowhead), but not in midgut cells. This sample was derived from a female. The CA was labeled with anti-JHAMT antibody (magenta). Blue is the DAPI signal.
Considering morphological abnormalities in the CA, it seemed possible that esgts > RasV12 expression enhances JH biosynthesis in the CA. Therefore, we next performed quantitative PCR on three JH-responsive genes, Krüppel-homolog 1 (Kr-h1), Jonah 25Bii (Jon25Bii), Odorant-binding protein 99b (Obp99b), to estimate the amount of JH in RasV12 overexpressors and controls19. Previous studies have shown that expression levels of Kr-h1 and Jon25Bii correlate positively with the amount of JH in the body, while Obp99b correlates negatively19,20. Our qPCR results showed that expression levels of Kr-h1 and Jon25Bii were increased by RasV12, while Obp99b was decreased (Fig. 3e–g; Supplementary Table S2). These results strongly suggest that esgts > RasV12 leads to abnormalities in CA cells and increased JH biosynthesis.
The CA-driver Aug21-GAL4 is not active in ISCs/EBs
A previous study has suggested that JH secreted from the CA is received by ISCs and/or EBs, enhancing ISC proliferation and leading to post-mating gut remodeling5. To overexpress genes to inhibit JH biosynthesis, the previous study utilized Aug21-GAL4, which is one of the most widely used CA-GAL4 drivers21. Since esg-GAL4 has high activity not only in ISCs/EBs, but also in the CA, we also verified whether Aug21-GAL4 is active in ISCs/EBs, as well as in the CA. We found that Aug21-GAL4-driven GFP signals, as well as anti-JHAMT immunoreactive signals, were present in the CA, but not in ISCs, EBs, or other types of midgut cells (Fig. 3h). This observation suggests that Aug21-GAL4-mediated manipulation of gene expression does not directly affect gene expression in ISCs and EBs.
Expression of other ISC/EB drivers in the CA
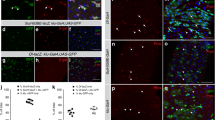
Lastly, we examined whether other GAL4 and LexA drivers used for gene manipulation in ISCs/EBs are also active in the CA. Beside esg-GAL4, many previous studies have used esg-LexA as an alternate binary gene expression driver active in ISCs/EBs22,23,24,25. We tested two esg-LexA drivers available from a stock center and found that both esg-LexA drivers were active in ISCs/EBs, but not in the adult or larval CA (Fig. 4a,b). It has also been reported that Delta-GAL4 and Su(H)-GAL4 are active in ISCs and EBs, respectively26. We found that the GFP signals driven by these GAL4 drivers were not present in the CA (Fig. 4c,d). These results suggest that the phenotypes resulting from genetic manipulation using these drivers are unlikely to be due to JH actions.
Expression of other ISC/EB drivers in the CA. GFP (green) driven by several drivers is shown. The CA was labeled with anti-JHAMT antibody (magenta). (a,b) GFP expression driven by two independent esg-LexA drivers in adult midguts (top), adult CA (middle), and ring glands (RG), containing the CA and PG, from wandering 3rd-instar (WL3) larvae (bottom). The PG was labeled with anti-Shroud (Sro) antibody (blue). (c,d) GFP expression driven by Delta-GAL4 (c) and Su(H)-GAL4 (d) in adult CA. (e,f) GFP expression driven by I-KCKT-GAL4 (e) and ISC-KCKTts-GAL4 (f) in adult midguts (top) and CA (bottom).
Another recent study reported new genetic tools, intestinal-kickout (I-KCKT)-GAL4 and ISC-KCKTts-GAL4, which are based in an intersectional method that restricts GAL4 expression to ISCs and/or EBs27 (Fig. 4e,f). The study has confirmed that I-KCKT-GAL4 and ISC-KCKTts-GAL4 are active in ISCs/EBs more specifically than conventional ISC/EB GAL4 drivers, such as esg-GAL4, Delta-GAL4, and Su(H)-GAL4, all of which are active in many types of cells other than ISCs/EBs4,27. We found that neither I-KCKT-GAL4 nor ISC-KCKTts-GAL4 labeled the CA, suggesting that these KCKT-based GAL4 lines do not affect CA gene expression.
Discussion
In this study, we found that esg-GAL4, which is widely used to label midgut ISC/EB2, was also expressed in the CA. esgts-GAL4 manipulates and influences gene expression in the CA, as esgts-GAL4 > jhamt RNAi decreases mature egg formation, which is a typical phenotype of reduced JH titer. Whereas esg-GAL4 is active in some ovarian cells5,6, it is reported by FlyBase (https://flybase.org/reports/FBgn0028841.html) that jhamt expression is not detected in the ovary. Therefore, the mature egg number phenotype by esgts-GAL4 > jhamt RNAi is due to the knock-down of jhamt expression in the CA but not the ovary. Moreover, esgts > RasV12 caused CA hypertrophy and influenced JH-responsive gene expression (Fig. 3b–g), suggesting that esgts > RasV12 increases JH biosynthesis.
Our data suggest that esg itself is expressed in the CA in both males and females of both larvae and adults. Enrichment of esg expression in the larval ring gland and the adult CA were suggested by a previous microarray analysis28 and a recent single-cell RNA-seq analysis29, respectively. esg encodes a Snail-type transcription factor that contributes to cell cycle regulation, cell differentiation, and cell–cell adhesion in many cell types in D. melanogaster30,31,32,33. However, functions of Esg that regulate differentiation and morphogenesis of the CA have not been studied. Thus, additional studies are needed to clarify how Esg is involved in CA cell regulation, especially whether it regulates JH biosynthesis.
In D. melanogaster, one of the reported functions of JH is that this hormone directly acts on ISCs and EBs through nuclear JH receptors, Methoprene-tolerant (Met) and Germ cell expressed (Gce), to regulate gut remodeling in mated or aged females5,8. Interestingly, previous studies reported that esgts > jhamt RNAi reduces numbers of ISCs and EBs. This phenotype is also observed in esgts > Met or gce RNAi animals. Since these studies use esgts-GAL4 as the ISC/EB- “specific” GAL4 driver, these papers propose that JH is biosynthesized in ISCs and EBs outside the CA, and cell-autonomously regulates maintenance of ISCs and EBs during aging8. However, our data strongly indicate that esg-GAL4 is also expressed in the CA. More importantly, esgts > jhamt RNAi causes a decrease in JHAMT protein in the CA (Fig. 2a), which implies decreased JH biosynthesis in the CA, hence the systemic decrease in JH titer. We emphasize that although these previous studies did not examine jhamt expression in ISCs and EBs, they carefully evaluated functions of JHAMT in ISCs and EBs with additional experiments in which they utilize other GAL4 drivers to knock down jhamt in ISCs and EBs via Delta-GAL4 and Su(H)-GAL4, respectively8. Therefore, we argue that JH biosynthesis most likely occurs in ISCs and EBs, although we cannot detect obvious anti-JHAMT reactivities in the gut region (Fig. 1c, 3h).
Nevertheless, a previous study using Aug21-GAL4, the well-known CA-GAL4 driver21, revealed that Aug21-GAL4-mediated inhibition of JH biosynthesis in the CA suppresses ISC proliferation5. Aug21-GAL4 is not active in ISCs/EBs (Fig. 3h), suggesting that the Aug21-GAL4-driven ISC phenotype cannot be explained by JH biosynthesis in ISCs/EBs. Taken together, we cannot rule out the possibility that JH is also supplied from the CA for maintenance of ISCs and EBs. In addition, ISC/EB increased by JH derived from the CA may further enhance JH production in the gut, implying that CA-ISC/EB interactions are more complex than previously thought. In the future, it will be important to investigate how much CA and ISC/EB each contribute to the circulating JH levels.
In the last decade, esg-GAL4 has widely been used to generate ISC tumors by overexpressing oncogenic genes such as the gain-of-function transgenes, Ras, Raf, and yki17,34. In particular, very recently, a number of studies have utilized esg-GAL4-driven oncogenic gene models to study how ISC tumors impact gut homeostasis, as well as systemic physiology35. However, based on our results, when interpreting these esg-GAL4-driven phenotypes, we should consider not only effects of ISC/EB tumors, but also effects of JH biosynthesis abnormalities caused by CA hypertrophy. For example, some intestinal cells receive JH from the CA through Met and Gce, influencing gut remodeling5,36. Beside RasV12, recent studies have shown that esgts > yki3SA leads to severe cachexia and bloating phenotypes, mediated by abnormal hormone secretion from organs such as Malpighian tubules and midgut9,23,37,38. Considering the systemic nature of these ISC/EB tumor-associated phenotypes, it may be necessary to consider the function of JH, which has major impacts on insect physiology. Based on observations in this and a previous study27, I-KCKT-GAL4 and ISC-KCKTts-GAL4 seem to be the current best choice of GAL4 drivers to manipulate gene expression specifically in ISCs and EBs without influencing the CA. However, even I-KCKT-GAL4 is active in Malpighian tubules27. Generally speaking, it will be important to examine phenotypes using more than just one GAL4 driver.
Methods
Drosophila strains and maintenance
D. melanogaster was raised on a standard yeast-corn meal-glucose fly medium (0.275 g agar, 5.0 g glucose, 4.5 g cornmeal, 2.0 g yeast extract, 150 μL propionic acid, and 175 μL 10% butyl p-hydroxybenzoate (in 70% ethanol) in 50 mL water) at 25 ºC under a 12:12 h light/dark cycle.
Throughout this study, we used esgts-GAL4 flies (a gift from Fumiaki Obata, RIKEN Center for Biosystems Dynamics Research) that carried both esg-GAL4 (RRID:BDSC_93857)27 and tub-GAL80ts. The following transgenic strains were also used: Aug21-GAL421 (BDSC #30137), Delta-GAL426 (a gift from Yuichiro Nakajima, University of Tokyo), esg-GFP (BDSC #78333), esg-LexABL66630 22 (BDSC #66630, a gift from Sa Kan Yoo, RIKEN Center for Biosystems Dynamics Research), esg-LexABL6663222 (BDSC #66632, a gift from Sa Kan Yoo, RIKEN Center for Biosystems Dynamics Research), I-KCKT-GAL4.p6527 (BDSC #91526), ISC-KCKT-GAL4ts27 (BDSC #91411), LexAop-myrGFP (BDSC #32209), Su(H)-GAL426 (a gifts from Yuichiro Nakajima, University of Tokyo), UAS-GFP; UAS-mCD8::GFP39 (a gift from Kei Ito, University of Cologne, Germany), UAS-jhamt-IRKK (VDRC #103958), and UAS-RasV12 (BDSC #4847). For adult-specific GAL4 activation, flies carrying esgts-GAL4 were reared at 21 °C from embryos to newly eclosed adults. 0–12 h after eclosion, flies were moved to 29 °C. To visualize esgts > GFP, wandering 3rd-inster larvae were used. Larvae were reared at 25 °C until the middle 3rd-larval instar and transferred to 29 °C for 24 h before dissection. Heterozygous controls were obtained by crossing w1118 with strains of GAL4 drivers.
Immunohistochemistry
Tissues were dissected in PBS and fixed in 4% paraformaldehyde in PBS for 30–60 min at 25–27 °C. Fixed samples were rinsed thrice in PBS, washed for 15 min with PBS containing 0.3% Triton X-100 (Nacalai Tesque #35501-15) (PBT), and treated with a blocking solution (2% bovine serum albumin in PBT; Sigma #A9647) for 1 h at 25–27 °C or overnight at 4 °C. Samples were incubated with a primary antibody in blocking solution overnight at 4 °C. Primary antibodies used were as follows: chicken anti-GFP antibody (Abcam #ab13970, 1:2,000), guinea pig anti-JHAMT antibody (1:2,000)40, rabbit anti-JHAMT antibody (1:1,000)12, guinea pig anti-Shroud (Sro) antibody (1:400)41. Samples were rinsed thrice with PBS and then washed for 15 min with PBT, followed by incubation with fluorophore (Alexa Fluor 488, 555, and 633)-conjugated secondary antibodies (Thermo Fisher Scientific #A32931, #A21435, #A32732, and #A21105; 1:200), in blocking solution for 2 h at RT or overnight at 4 °C. Nuclear stains used in this study were 4ʹ,6-diamidino-2-phenylindole (DAPI; Sigma-Aldrich #D9542; final concentration 1 μg/ml). For DAPI staining, after incubation with secondary antibodies, samples were washed and then incubated with DAPI for 1 h. After another round of washing, all samples were mounted on glass slides using FluorSave reagent (Merck Millipore, #345789). All fluorescence images were captured using Nikon SMZ25 dissection microscope with the imaging software NIS-Elements version 5.42.01 (for Fig. 1b) and Carl Zeiss LSM700 or LSM900 confocal microscopy with the imaging software Zen version 8,1,0,484 (for all other figures). Quantification of immunostaining signals was conducted using ImageJ software version 1.53q42.
Counting mature egg numbers in ovaries
Virgin females carrying esgts-GAL4 were reared at 21 °C from embryos to newly eclosed adults. 0–12 h after eclosion, flies were transferred to 29 °C and reared for 4 days. In each vial, fewer than 10 females were reared. Ovaries of virgin females were dissected in PBS. Numbers of mature eggs (stage-14 oocytes)43 in the ovaries were counted under a stereomicroscope (Leica MZ10F).
Reverse transcription-quantitative PCR (RT-qPCR)
Total RNA was extracted from whole bodies of 4-day-old adult virgin female flies. RNA was reverse-transcribed using ReverTra Ace qPCR RT Master Mix with gDNA Remover (TOYOBO #FSQ-301). Synthesized cDNA samples were used as templates for quantitative PCR using THUNDERBIRD SYBR qPCR Mix (TOYOBO #QPS-201) on a Thermal Cycler Dice Real Time System (Takara Bio #TP870). The amount of target RNA was normalized to the endogenous control ribosomal protein 49 gene (rp49) and the relative change was calculated. Expression levels of each gene were compared using the ΔΔCt method. The following primers were used for this analysis: rp49 F (5ʹ-CGGATCGATATGCTAAGCTGT-3ʹ), rp49 R (5ʹ-GCGCTTGTTCGATCCGTA-3ʹ), kr-h1 F (5ʹ-TCACACATCAAGAAGCCAACT-3ʹ), kr-h1 R (5ʹ-GCTGGTTGGCGGAATAGTAA-3ʹ), obp99b F (5ʹ-AGCACGGATTCGATGTCCACAAGA-3ʹ), obp99b R (5ʹ-TTGGAGTTCATGAAGCACATGCCG-3ʹ), jon25Bii F (5ʹ-CAGGCTCAGTACACCCACAC-3ʹ), jon25Bii R (5ʹ-TGGTGTTGTAGTCCGAGTGC-3ʹ).
Statistical analysis
All experiments were performed independently at least twice. Sample sizes were chosen based on the number of independent experiments required for statistical significance and technical feasibility. Experiments were not randomized, and investigators were not blinded. All statistical analyses were performed using “R”, software version 4.0.3. Details of statistical analyses are described in figure legends.
Data availability
All numerical data are available in Supplementary Tables S1 and S2. All other data are available upon request to R.N.
References
Brand, A. H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 (1993).
Goto, S. & Hayashi, S. Proximal to distal cell communication in the Drosophila leg provides a basis for an intercalary mechanism of limb patterning. Development 126, 3407–3413 (1999).
Jasper, H. Intestinal stem cell aging: Origins and interventions. Annu. Rev. Physiol. 82, 203–226 (2020).
Weaver, L. N., Ma, T. & Drummond-Barbosa, D. Analysis of Gal4 expression patterns in adult Drosophila females. G3 10, 4147–4158 (2020).
Reiff, T. et al. Endocrine remodelling of the adult intestine sustains reproduction in Drosophila. Elife 4, e06930 (2015).
Hudry, B., Khadayate, S. & Miguel-Aliaga, I. The sexual identity of adult intestinal stem cells controls organ size and plasticity. Nature 530, 344–348 (2016).
Wang, C. & Spradling, A. C. An abundant quiescent stem cell population in Drosophila Malpighian tubules protects principal cells from kidney stones. Elife 9, e54096 (2020).
Rahman, M. M., Franch-Marro, X., Maestro, J. L., Martin, D. & Casali, A. Local juvenile hormone activity regulates gut homeostasis and tumor growth in adult Drosophila. Sci. Rep. 7, 11677 (2017).
Song, W. et al. Tumor-derived ligands trigger tumor growth and host wasting via differential MEK activation. Dev. Cell 48, 277-286.e6 (2019).
Micchelli, C. A. & Perrimon, N. Evidence that stem cells reside in the adult Drosophila midgut epithelium. Nature 439, 475–479 (2006).
Shinoda, T. & Itoyama, K. Juvenile hormone acid methyltransferase: A key regulatory enzyme for insect metamorphosis. Proc. Natl. Acad. Sci. USA 100, 11986–11991 (2003).
Niwa, R. et al. Juvenile hormone acid O-methyltransferase in Drosophila melanogaster. Insect Biochem. Mol. Biol. 38, 714–720 (2008).
He, L., Si, G., Huang, J., Samuel, A. D. T. & Perrimon, N. Mechanical regulation of stem-cell differentiation by the stretch-activated Piezo channel. Nature 555, 103–106 (2018).
Riddiford, L. M. Rhodnius, golden oil, and met: A history of juvenile hormone research. Front. Cell Dev. Biol. 8, 679 (2020).
John, H. P. & Weiser, K. Vitellogenesis induced by juvenile hormone in the female sterile mutant apterous-four in Drosophila melanogaster. Nat. New Biol. 244, 284–285 (1973).
Wen, D. et al. Methyl farnesoate plays a dual role in regulating Drosophila metamorphosis. PLoS Genet. 11, e1005038 (2015).
Lee, J., Cabrera, A. J. H., Nguyen, C. M. T. & Kwon, Y. V. Dissemination of RasV12-transformed cells requires the mechanosensitive channel Piezo. Nat. Commun. 11, 3568 (2020).
Jin, Y. et al. EGFR/Ras signaling controls Drosophila intestinal stem cell proliferation via capicua-regulated genes. PLoS Genet. 11, e1005634 (2015).
Yamamoto, R., Bai, H., Dolezal, A. G., Amdam, G. & Tatar, M. Juvenile hormone regulation of Drosophila aging. BMC Biol. 11, 85 (2013).
Andreatta, G., Kyriacou, C. P., Flatt, T. & Costa, R. Aminergic signaling controls ovarian dormancy in Drosophila. Sci. Rep. 8, 2030 (2018).
Siegmund, T. & Korge, G. Innervation of the ring gland of Drosophila melanogaster. J. Comp. Neurol. 431, 481–491 (2001).
Kockel, L. et al. A Drosophila LexA enhancer-trap resource for developmental biology and neuroendocrine research. G3 6, 3017–3026 (2016).
Chen, Y. et al. Renal NF-κB activation impairs uric acid homeostasis to promote tumor-associated mortality independent of wasting. Immunity 55, 1594-1608.e6 (2022).
Okada, M. et al. Oncogenic stress-induced Netrin is a humoral signaling molecule that reprograms systemic metabolism in Drosophila. EMBO J. 42, e111383 (2023).
Xu, W., Li, G., Chen, Y., Ye, X. & Song, W. A novel antidiuretic hormone governs tumour-induced renal dysfunction. Nature 624, 425–432 (2023).
Zeng, X., Chauhan, C. & Hou, S. X. Characterization of midgut stem cell- and enteroblast-specific Gal4 lines in Drosophila. Genesis 48, 607–611 (2010).
Buddika, K., Xu, J., Ariyapala, I. S. & Sokol, N. S. I-KCKT allows dissection-free RNA profiling of adult Drosophila intestinal progenitor cells. Development 148, dev196568 (2021).
Ou, Q. et al. The insect prothoracic gland as a model for steroid hormone biosynthesis and regulation. Cell Rep. 16, 247–262 (2016).
Zhu, H., Ludington, W. B. & Spradling, A. C. Cellular and molecular organization of the Drosophila foregut. Proc. Natl. Acad. Sci. USA 121, e2318760121 (2024).
Voog, J. et al. Escargot restricts niche cell to stem cell conversion in the Drosophila testis. Cell Rep. 7, 722–734 (2014).
Fuse, N., Hirose, S. & Hayashi, S. Diploidy of Drosophila imaginal cells is maintained by a transcriptional repressor encoded by escargot. Genes Dev. 8, 2270–2281 (1994).
Antonello, Z. A., Reiff, T., Ballesta-Illan, E. & Dominguez, M. Robust intestinal homeostasis relies on cellular plasticity in enteroblasts mediated by miR-8-Escargot switch. EMBO J. 34, 2025–2041 (2015).
Tanaka-Matakatsu, M., Uemura, T., Oda, H., Takeichi, M. & Hayashi, S. Cadherin-mediated cell adhesion and cell motility in Drosophila trachea regulated by the transcription factor Escargot. Development 122, 3697–3705 (1996).
Markstein, M. et al. Systematic screen of chemotherapeutics in Drosophila stem cell tumors. Proc. Natl. Acad. Sci. USA 111, 4530–4535 (2014).
Gong, S., Zhang, Y., Tian, A. & Deng, W.-M. Tumor models in various Drosophila tissues. WIREs Mech. Dis. 13, e1525 (2021).
Zipper, L., Jassmann, D., Burgmer, S., Görlich, B. & Reiff, T. Ecdysone steroid hormone remote controls intestinal stem cell fate decisions via the PPARγ-homolog Eip75B in Drosophila. Elife 9, e55795 (2020).
Kwon, Y. et al. Systemic organ wasting induced by localized expression of the secreted insulin/IGF antagonist ImpL2. Dev. Cell 33, 36–46 (2015).
Ding, G. et al. Coordination of tumor growth and host wasting by tumor-derived Upd3. Cell Rep. 36, 109553 (2021).
Ito, K. et al. The organization of extrinsic neurons and their implications in the functional roles of the mushroom bodies in Drosophila melanogaster Meigen. Learn. Mem. 5, 52–77 (1998).
Mizuno, Y. et al. A population of neurons that produce hugin and express the diuretic hormone 44 receptor gene projects to the corpora allata in Drosophila melanogaster. Dev. Growth Differ. 63, 249–261 (2021).
Shimada-Niwa, Y. & Niwa, R. Serotonergic neurons respond to nutrients and regulate the timing of steroid hormone biosynthesis in Drosophila. Nat. Commun. 5, 5778 (2014).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
King, R. C. The meiotic behavior of the Drosophila oocyte. Int. Rev. Cytol. 28, 125–168 (1970).
Acknowledgements
We thank Yuto Yoshinari for critical input during the initial stage of this work. We also thank Kei Ito, Fumiaki Obata, Sa Kan Yoo, Yuichiro Nakajima, Bloomington Stock Center, Vienna Drosophila Resource Center for fly strains. We are also grateful to Yuichiro Nakajima and Morihiro Okada for helpful discussions, and Steve D. Aird for editing the manuscript. This work was supported by the Japan Society of the Promotion of Science KAKENHI (21J20365 to Y.K. and 23KJ0252 to Y.M.) and by the Japan Science and Technology Agency grant SPRING JPMJSP2124. Y.K., Y.M., and T.K. received fellowships from the JSPS.
Author information
Authors and Affiliations
Contributions
T.K. and R.N. designed the research. Y.K., Y.M., and T.K. conceived of the experiment. Y.K. and Y.M. acquired the data. All authors analyzed the data. Y.K., Y.M., and R.N. wrote the manuscript and prepared figures. All authors reviewed the manuscript. Y.K., Y.M., and T.K. contributed equally to this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kurogi, Y., Mizuno, Y., Kamiyama, T. et al. The intestinal stem cell/enteroblast-GAL4 driver, escargot-GAL4, also manipulates gene expression in the juvenile hormone-synthesizing organ of Drosophila melanogaster. Sci Rep 14, 9631 (2024). https://doi.org/10.1038/s41598-024-60269-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-60269-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.