Abstract
The SMALL ACIDIC PROTEIN (SMAP) gene is evolutionarily indispensable for organisms. There are two copies of the SMAP gene in the Arabidopsis thaliana genome, namely, SMAP1 and SMAP2. The function of SMAP2 is similar to that of SMAP1, and both can mediate 2,4-D responses in the root of Arabidopsis. This study cloned the AtSMAP2 genetic promoter sequence. Two promoter fragments of different lengths were designed according to the distribution of their cis-acting elements, and the corresponding β- glucuronidase (GUS) expression vector was constructed. The expression activity of promoters of two lengths, 1993 bp and 997 bp, was studied by the genetic transformation in Arabidopsis. The prediction results of cis-acting elements in the promoter show that there are many hormone response elements in 997 bp, such as three abscisic acid response elements ABRE, gibberellin response elements P-box and GARE-motif and auxin response element AuxRR-core. Through GUS histochemical staining and qRT‒PCR analysis, it was found that the higher promoter activity of PAtSMAP2-997, compared to PAtSMAP2-1993, drove the expression of GUS genes at higher levels in Arabidopsis, especially in the root system. The results provide an important basis for subsequent studies on the regulation of AtSMAP2 gene expression and biological functions.
Similar content being viewed by others
Introduction
During the growth and development of plants, genes are expressed in different growth and development stages and in different tissues and the promoter of the gene itself plays a key role in this process. A promoter is a segment of DNA sequence located upstream of the gene that provides recognition and binding by RNA polymerase. Transgenic technology is often used in the functional study of specific genes or in molecular breeding techniques that enable recipient organisms to acquire specific traits. The accurate expression of the promoter of the target gene is particularly important in this process, as sometimes even the same target gene is expressed differently depending on its genetic characteristics. According to expression characteristics, promoters can be categorized as constitutive promoters, inducible promoters, and tissue-specific promoters. The selection of an appropriate promoter is important for the foreign gene to perform the desired function. In most transgenic engineering, constitutive promoters are often used to drive the expression of exogenous genes due to their independence from temporal and spatial constraints and the induction of substances, as well as their advantages of being efficient and more stable, such as the cauliflower virus (CaMV) 35S promoter1 and ubiquitin 1 (Ubi) promoter2,3. Inducible promoters do not transcribe or have low transcriptional activity under normal circumstances but can effectively regulate the expression of downstream genes when induced by specific biological factors or physical and chemical substances. At the same time, this type of promoter can reduce the damage caused by the expression of foreign genes in transgenic plants, such as the rbcs promoter4. Tissue-specific promoters have the characteristics of driving the selective expression of target genes in specific cells, tissues, or developmental stages and are commonly used to regulate plant growth and development5, such as cotton fiber-specific GhPRP56 and tomato fruit-specific E8 promoter7. Several studies have demonstrated that promoters play a key role in gene expression and regulation in plants. Promoters mainly regulate the initiation time, tissue specificity, and expression intensity of gene expression8,9. It can also combine with corresponding transcription factors and other co-factors to precisely control the efficiency of gene expression, respond to various biotic and abiotic stresses, and initiate various life activities10,11. Therefore, the study of gene promoters is of great significance for understanding the expression characteristics of genes.
The phytohormone auxin regulates plant development by inducing cellular responses and altering gene expression. 2,4-dichlorophenoxyacetic acid (2,4-D) is a functional analog of the plant endogenous auxin indole-3-acetic acid (IAA), which is often used as a source of auxin and can induce auxin-related responses, such as suppression of root growth and induction of lateral root growth12. IAA is unstable under blue light and UV light13, and 2,4-D has greater stability in growth media, so in various physiological experiments, compared with IAA, 2,4-D is more suitable as an exogenous source of auxin. The difference between IAA and 2,4-D is in mode of transport and metabolism; 2,4-D is excreted and broken down more slowly than IAA and therefore accumulates more in the cells14,15,16,17.
Studies have shown that auxin signaling is mainly accomplished through the degradation of the AUXIN/IAA (AUX/IAA) repressor proteins, which is dependent on the ubiquitin‒proteasome degradation pathway18. The degradation of AUX/IAA is directly regulated by SCFTIR1/AFB complex, consisting of CULLIN 1 (CUL1), S PHASE KINASE-ASSOCIATED PROTEIN 1 (SKP1), RING-BOX PROTEIN 1 (RBX1) and a substrate-recognizing F-box protein, TRANSPORT INHIBITOR RESPONSE 1 (TIR1) or AUXIN SIGNALING F-BOXES (AFBs)19. Auxin binds to TIR1/AFB to promote SCFTIR1/AFB with AUX/IAA proteins and the ubiquitinated AUX/IAA protein is degraded by the 26S proteasome. The AUX/IAA protein and auxin response factors (ARFs) form heterodimers and inhibit the transcriptional activity of ARFs20. ARFs can specifically bind to the auxin response element (AuxRE) "TGTCTC" in the promoter region of auxin-responsive genes to activate or inhibit gene expression21. Therefore, the degradation of AUX/IAA proteins leads to ARFs release, thereby regulating the expression of downstream genes and activating a series of auxin responses.
To better understand the mechanisms of auxin signaling, mutants with altered auxin responses have been screened using a variety of genetic approaches; however, due to the absolute necessity of auxin for embryogenesis, seedling mutants treated with auxin screening have certain limitations. Another approach is to screen Arabidopsis mutants for abnormal responses to compounds that alter or antagonize the auxin response. Therefore, studies have reported that using an inhibitor of auxin signaling events, p-chlorophenoxyisobutyric acid (PCIB), to screen Arabidopsis mutants, and finally obtained an anti-auxin resistant (aar) mutant, this mutant can still grow longer roots on PCIB-containing medium22,23. In one of the mutants, aar1-1, in terms of root elongation, lateral root formation, seed germination in the presence of abscisic acid, and degradation of AUX/IAA proteins, the response to endogenous auxin IAA is the same as that of the wild type but shows specific resistance to the synthetic auxin 2,4-D, while having no effect on 2,4-D transport or metabolism24. Molecular characterization of this mutant showed that the SMALL ACIDIC PROTEIN1 (SMAP1) gene confers 2,4-D and PCIB resistance and a longer hypocotyl phenotype23,24. The SMAP1 protein consists of 62 amino acids with a size of 6.9 kDa and an isoelectric point of 3.4. Although the SMAP protein has no known functional domains, it has a highly conserved 18 amino acid sequence rich in phenylalanine and aspartic acid at its C-terminal region24. Studies of aar1 mutants and the SMAP1 gene show that the SMAP1 protein functions upstream of AUX/IAA protein degradation in auxin signaling and physically interacts with the constitutive photomorphogenic9 signalosome (CSN) through its F/D region24,25. CSN is a key regulator of cullin-RING ubiquitin ligases, the largest family of E3 ubiquitin ligases, including SCFTIR1/AFB, the major upstream pathway that regulates auxin signaling26. This F/D region is present in the genomes of various plants and animals, which means that SMAP genes are evolutionarily indispensable to organisms24,27.
There is another copy of the SMAP gene in the Arabidopsis genome, SMALL ACIDIC PROTEIN 2 (SMAP2). The SMAP2 protein consists of 72 amino acids with a size of 9.4 kDa and an isoelectric point of 3.5. SMAP1 and SMAP2 share 43.5% identity in the entire amino acid sequence and 83.3% identity in their highly conserved C-terminal F/D-rich domains27. While there are two SMAP genes in the Arabidopsis genome, a BLAST search revealed only one SMAP gene in other dicots, such as populus, grape, and tomato27. RNA expression analysis showed that SMAP1 mRNA was expressed in the whole Arabidopsis plant, while SMAP2 mRNA was only expressed in siliques and anthers, with a high level of expression in anthers and a small amount in green siliques27. By overexpressing the SMAP1 and SMAP2 genes, respectively, in aar1 mutants, the function of SMAP2 is similar to that of SMAP1, and both can mediate the 2,4-D response in Arabidopsis roots27.
However, there are few studies on the other functions of the SMAP2 gene and its expression regulation mechanism, and studies on its promoter activity have not been reported. In this study, the promoter sequence of the AtSMAP2 gene was cloned, and two promoter fragments of different lengths were designed according to the distribution of its cis-acting elements, and the corresponding β-glucuronidase (GUS) expression vector was constructed. The expression activities of the two lengths of promoter, which were 1993 bp and 997 bp respectively, were investigated by genetic transformation in Arabidopsis. Through GUS histochemical staining and qRT‒PCR analysis, it was found that the higher promoter activity of PAtSMAP2-997, compared to PAtSMAP2-1993, drove the expression of GUS genes at higher levels in Arabidopsis, especially in the root system.
Results
Sequence analysis of the AtSMAP2 promoter
The 1998 bp promoter sequence upstream of the transcriptional start codon ATG was obtained from the AtSMAP2 gene (AT3G24280) sequence on The Arabidopsis Information Resource (TAIR) (https://www.arabidopsis.org/). The cis-acting elements of the 1998 bp promoter sequence upstream of ATG (the translation initiation site ATG is numbered as + 1) were predicted by searching the PLACE and PlantCARE databases (Fig. 1). Using these two types of analysis software, four types of cis-acting elements were found to be present in this sequence, namely major cis-regulatory elements, phytohormone response elements, light response elements, and environmental stress regulation-related elements. The core sequences and predicted functions of the various cis-acting elements are listed in Table 1. The prediction results showed that multiple hormone signal response elements were concentrated in the 997 bp fragment, including three abscisic acid response elements, two gibberellin response elements, and one auxin response element. It is speculated that AtSMAP2 gene expression may be related to the hormone response.
Sequence analysis of full-length promoter of AtSMAP2 gene. The translation initiation site ‘ATG’ (marked in underline) of AtSMAP2 was numbered as + 1. The cis-acting elements of the 1998 bp full-length promoter sequence upstream of ATG were predicted by searching the PLACE and PlantCARE databases. The different cis-acting elements were marked in different colours. Among these cis-acting elements, CAAT-box is the major cis-regulatory elements; TCA-element, CGTCA-motif, TGACG-motif, ABRE, P-box, GARE-motif, AuxRR-core belong to the phytohormone response elements; TC-rich repeats, ARE, circadian belong to the environmental stress regulation-related elements; TCT-motif, chs-CMA1a, AE-box, Box 4, ATCT-motif, GATA-motif, MRE belong to the light response elements. The bold sequences in red are the forward primer sequences used to clone the 1993 bp and 997 bp promoter fragments and the common reverse primer sequences, respectively.
Generation of GUS-expressing transgenic Arabidopsis driven by different AtSMAP2 promoter fragments
Based on the results obtained from cis-acting element prediction of the promoter sequence upstream of ATG of the AtSMAP2 gene using PLACE and PlantCARE, a 1993 bp promoter fragment containing most of the cis-acting elements upstream of this gene and a 997 bp promoter fragment upstream of this gene containing multiple hormone response elements, namely 3 abscisic acid response elements, 2 gibberellin response elements, and 1 auxin response element, were amplified by two specific forward PCR primers and one common reverse primer (Fig. 2). To determine the expression activity of the 1993 bp and 997 bp promoters, the PCR product was fused with the GUS reporter gene and used for Agrobacterium-mediated Arabidopsis genetic transformation.
T1 generation transgenic Arabidopsis seeds were screened on an MS medium containing 50 mg/L hygromycin. After extracting DNA from a small number of leaves of the T1 generation resistant plants, PCR detection was performed with promoter-GUS-specific primers, and the bands were amplified to the same size as the positive control target band, indicating that the two AtSMAP2 promoter fragments were successfully integrated into the Col-0 genome (Fig. 3). At least 20 individual lines with a single transgene insertion were selected in the T2 generation based on the 3:1 segregation ratio for hygromycin resistance. Through the evaluation of the segregation ratio, the T3 transgenes homozygotes were confirmed from a single generation of T2 plants and used for further histochemical staining and qRT‒PCR analysis. A number of independent transgenic lines were obtained, and at least 5 typical GUS-positive independent T3 lines were analyzed by histochemical staining. Histochemical staining of transgenic Arabidopsis for GUS revealed that the two promoter fragments always had different activities, while the GUS expression was consistent under the same promoter fragment.
PCR analysis of transgenic A. thaliana events showing two promoter-GUS fragments. (a) PCR identification of PAtSMAP2-997-GUS strain. (b) PCR identification of PAtSMAP2-1993-GUS strain. M: DNA marker (100-2000 bp); lane 1: untransformed negative control, lane 2: pMDC162 plasmid DNA, lane 3 to11: transgenic plants of A. thaliana.
Expression patterns of two AtSMAP2 promoter fragments in different growth stages and tissues
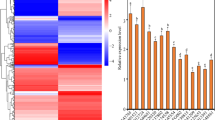
To understand the expression activities of the two promoters, GUS staining was performed at the seedling stage of the transgenic positive lines, and staining was performed every 24 h (Fig. 4a). The 24 h staining results showed that for the PAtSMAP2-997-GUS strain, the dyeing effect was more obvious, and for the PAtSMAP2-1993-GUS strain, the dyeing effect was not obvious. The staining results of seedlings on 2 days of germination showed that the roots and leaves of the PAtSMAP2-997-GUS line were stained, and the leaves and root tips of the PAtSMAP2-1993-GUS strain were also stained, but their expression activity was lower. The staining results of seedlings on the 3rd and 4th day of germination showed that both the PAtSMAP2-997-GUS and PAtSMAP2-1993-GUS strains were stained throughout, but the PAtSMAP2-1993-GUS strain had lower expression activity in leaves, hypocotyls, and roots than the PAtSMAP2-997-GUS strain. The staining results from the 5th day to the 7th day of germination showed that unlike the PAtSMAP2-1993-GUS line, the entire PAtSMAP2-997-GUS strain was stained and its root hairs were also stained, in addition to significantly stronger staining of its primary roots.
Detection of GUS expression driven by different AtSMAP2 promoter fragments in transgenic Arabidopsis plants at the seedling stage. (a) GUS histochemical assay in transgenic Arabidopsis seedlings. Bars = 1 mm. (b) qRT-PCR detection of GUS expression patterns in transgenic Arabidopsis plants at the seedling stage. Data were presented as the mean ± SE of three separate measurements for each independent line.
GUS staining was also performed in 20-day-old transgenic Arabidopsis lines and mature organs of transgenic Arabidopsis. The leaves, flowers, and roots of mature transgenic Arabidopsis lines were stained and observed. Histochemical staining analysis showed that (Fig. 5a), consistent with the staining results at the seedling stage, root staining was stronger in both the 20-day-old and mature stage of the PAtSMAP2-997-GUS strain than in the PAtSMAP2-1993-GUS strain. The staining results at the 20-day-old plants showed that in the PAtSMAP2-997-GUS strain, the degree of staining of leaves was not uniform, and the first pair of leaf primordia and hypocotyl showed a stronger degree of staining than the PAtSMAP2-1993-GUS strain, while the second pair of leaf primordia was lighter stained than those of the PAtSMAP2-1993-GUS strain. There was no significant difference in the degree of staining of leaves at the mature stage of the two lines, and the degree of staining of flowers of the PAtSMAP2-997-GUS line was stronger than that of the PAtSMAP2-1993-GUS line.
Expression patterns of AtSMAP2 promoter fragments in different developmental stages and tissues. (a) GUS histochemical assay in 20-day-old seedlings and different mature organs of transgenic Arabidopsis. Bars = 1 mm. (b) qRT-PCR detection of GUS expression patterns in 20-day-old seedlings and different mature organs of transgenic Arabidopsis. Data were presented as the mean ± SE of three separate measurements for each independent line.
To validate the results of GUS staining, a highly sensitive qRT‒PCR analysis of GUS gene transcription level was further used to verify the activity of the two AtSMAP2 promoters with different fragment lengths. The qRT‒PCR results were consistent with the GUS histochemical staining results. From the first day of germination to the seventh day of germination, the promoter activity of PAtSMAP2-997 was always stronger than that of PAtSMAP2-1993 (Fig. 4b). Comparing different tissues, it was found that the difference between the two promoters was most pronounced in the roots at the 20-day-old and mature stage of Arabidopsis thaliana, with PAtSMAP2-997 promoter activity consistently stronger than PAtSMAP2-1993 activity in roots (Fig. 5b).
To sum up the results, the higher promoter activity of PAtSMAP2-997, compared with PAtSMAP2-1993, drives the expression of the GUS gene at a higher level in Arabidopsis thaliana, especially in the root system where the expression difference is more obvious, suggesting that there are DNA sequence elements in the range of -1 bp ~ -997 bp to ensure the expression of AtSMAP2 gene at a higher level, while there may be sequences that inhibit its expression in the range of − 997 ~ − 1993 bp.
Expression activity of the two AtSMAP2 promoter fragments under different concentrations of 2,4-D and IAA
Both PAtSMAP2-997-GUS and PAtSMAP2-1993-GUS lines were stained after treatment with exogenous hormone 2,4-D and IAA. After treatment with the exogenous hormone IAA, the depth of GUS staining in roots and leaves of the PAtSMAP2-997-GUS strains did not change significantly as a result of the increase in its concentration (Fig. 6a). The PAtSMAP2-1993-GUS strain showed a significant decrease in the depth of staining in roots and leaves at 50 nM IAA treatment, whereas no significant change in the depth of staining occurred under 100 nM and 150 nM IAA treatment conditions (Fig. 6a). Notably, the PAtSMAP2-997-GUS strain always stained darker than the PAtSMAP2-1993-GUS strain under the three concentrations of IAA treatment (Fig. 6a). The PAtSMAP2-997-GUS strain showed enhanced expression activity in the roots and leaves at all three concentrations of 2,4-D treatment compared to the control group (Fig. 6a). The PAtSMAP2-1993-GUS strain was also stained under all three concentrations of 2,4-D treatment, but its expression activity in roots and leaves did not differ from that of its control strain without the hormone (Fig. 6a). After 2,4-D treatment, the expression activity of the PAtSMAP2-1993-GUS strain remained significantly lower than that of the PAtSMAP2-997-GUS strain (Fig. 6a).
Detection of GUS expression driven by different AtSMAP2 promoter fragments in transgenic Arabidopsis treated with different concentrations of the exogenous hormone 2,4-D and IAA. (a) GUS histochemical analysis of transgenic Arabidopsis after treatment with 2,4-D and IAA. Bars = 1 mm. (b) qRT-PCR detection of GUS expression in transgenic Arabidopsis seedlings treated with 2,4-D and IAA. Data were presented as the mean ± SE of three separate measurements for each independent line. Different lower-case letters above the bars indicated significantly different at p < 0.05.
The qRT-PCR results were consistent with the GUS histochemical staining results that the activity of the PAtSMAP2-997 promoter was always significantly higher than that of the PAtSMAP2-1993 promoter, regardless of the addition of IAA or 2,4-D (Fig. 6b). The expression activity of the PAtSMAP2-997 promoter was not significantly different from that of the control after the addition of the exogenous hormone IAA, whereas the expression activity of the PAtSMAP2-1993 promoter was significantly lower than that of the control under the 50 nM IAA treatment condition (Fig. 6b). The expression activity of PAtSMAP2-997 initiated by the addition of both 50 nM and 100 nM exogenous hormone 2,4-D was significantly higher than that of the control, whereas the PAtSMAP2-1993 promoter was not significantly different from that of the control at any of the three 2,4-D concentrations (Fig. 6b). In summary, in Arabidopsis, the SMAP2 gene promoter showed different responses to IAA and 2,4-D, also, the activity of the PAtSMAP2-997 promoter was consistently stronger than that of the PAtSMAP2-1993 promoter in this process.
Expression activity of the two AtSMAP2 promoter fragments under different concentrations of ABA
The addition of three concentrations of exogenous ABA, 25 µM, 50 µM and 100 µM, respectively, resulted in reduced GUS expression activity in the leaves of the PAtSMAP2-997-GUS strain compared to its control, as well as in its roots at 25 µM and 50 µM (Fig. 7a). Observation of GUS expression activity in the PAtSMAP2-1993-GUS strain revealed that it exhibited reduced GUS expression activity in both leaves and roots compared to its control at 50 µM and 100 µM ABA (Fig. 7a). The qRT-PCR results showed that the three concentrations of ABA treatment exhibited a significant attenuation of GUS expression activity in both PAtSMAP2-997-GUS and PAtSMAP2-1993-GUS lines compared to their controls (Fig. 7b). Therefore, it was hypothesized that the expression activity of the SMAP2 gene promoter would be regulated by ABA, which inhibits its promoter expression activity. Finally, it is worth noting that the activity of the PAtSMAP2-997 promoter remained consistently stronger than that of the PAtSMAP2-1993 promoter when exogenous ABA was added (Fig. 7b).
Detection of GUS expression driven by different AtSMAP2 promoter fragments in transgenic Arabidopsis treated with different concentrations of the exogenous hormone ABA. (a) GUS histochemical analysis of transgenic Arabidopsis after treatment with ABA. Bars = 1 mm. (b) qRT-PCR detection of GUS expression in transgenic Arabidopsis seedlings treated with ABA. Data were presented as the mean ± SE of three separate measurements for each independent line. Different lower-case letters above the bars indicated significantly different at p < 0.05.
Expression activity of the two AtSMAP2 promoter fragments under different concentrations of MeJA
After treating Arabidopsis seedlings with three concentrations of exogenous MeJA, 1 µM, 25 µM, and 50 µM, respectively, seedlings of the PAtSMAP2-997-GUS strain were observed, and none of them showed changes in GUS expression activity in the whole plant compared to its control (Fig. 8a). Seedlings of the PAtSMAP2-1993-GUS strain were observed and found to exhibit stronger GUS expression activity than the corresponding controls when all three concentrations of MeJA were added (Fig. 8a). The qRT-PCR results showed that the GUS expression in the PAtSMAP2-997-GUS strain, after the addition of the three concentrations of MeJA, was not significantly different from that of its control, whereas the GUS expression in the PAtSMAP2-1993-GUS strain showed significant up-regulation in the three different concentrations of the treatment compared to its control (Fig. 8b). Overall, the PAtSMAP2-1993 promoter showed sensitivity to exogenous MeJA, but the expression activity of the PAtSMAP2-997 promoter was still stronger than that of the PAtSMAP2-1993 promoter.
Detection of GUS expression driven by different AtSMAP2 promoter fragments in transgenic Arabidopsis treated with different concentrations of MeJA. (a) GUS histochemical analysis of transgenic Arabidopsis after treatment with MeJA. Bars = 1 mm. (b) qRT-PCR detection of GUS expression in transgenic Arabidopsis seedlings treated with MeJA. Data were presented as the mean ± SE of three separate measurements for each independent line. Different lower-case letters above the bars indicated significantly different at p < 0.05.
Discussion
The 997 bp promoter fragment upstream of the AtSMAP2 gene is sufficient for efficient expression of this gene
When an exogenous gene is introduced into a recipient plant, the expression must be driven by a promoter. Therefore, in the field of plant engineering, the study of promoters is of great significance to the expression of exogenous genes. In this study, it was found that the AtSMAP2 gene promoter contains core elements that regulate gene expression, such as the TATA box, which is necessary for RNA polymerase II transcription initiation, and the CAAT box, which enhances gene transcription. In addition, there are hormone response elements, light regulatory elements, stress response elements, etc., indicating that AtSMAP2 gene expression may be affected by various factors, such as light, hormones, and environmental stress.
In this study, two promoter fragments of different lengths of AtSMAP2 gene, 997 bp (− 1 to − 997 bp) and 1993 bp (− 1 to − 1993 bp), were designed, and the corresponding GUS expression vector was constructed. The expression activity of two lengths of the promoter was studied by Arabidopsis genetic transformation. Notably, this study showed that the GUS expression activity in the PAtSMAP2-997-GUS strain was always higher than that in the PAtSMAP2-1993-GUS strain. In fact, the 1993 bp promoter fragment contains all the cis-acting elements contained in the 997 bp fragment and contains more typical core elements, TATA-box and CATA-box, so it should have higher expression activity than the 997 bp promoter fragment. However, the results of our study were the opposite, in transgenic Arabidopsis, the 997 bp promoter fragment always had higher expression activity than the 1993 bp promoter from the germination to maturation stages and under exogenous hormone-addition conditions. It was therefore hypothesized that there are sequences in − 1 to − 997 bp that ensure efficient expression of the AtSMAP2 gene, while there are sequences in − 998 to − 1993 bp that repress the expression of this gene. For example, the MeJA response elements are located at − 1150 and − 1110 bp, and there is no MeJA response element in the 997 bp fragment, yet the promoter fragment of 997 bp still has higher expression activity than that of 1993 bp when exogenous MeJA is added. It was further demonstrated that there is a cis-acting element in − 998 to − 1993 bp that represses the expression of this gene, but the exact location needs to be verified by truncation experiments, which will be the focus of our subsequent work. In addition, which specific sequences in − 1 to − 997 bp are essential for the expression of this gene also still need to be verified by subsequent experiments.
The AtSMAP2 gene promoter showed different responses to IAA and 2,4-D
The addition of exogenous IAA or 2,4-D usually results in the inhibition of plant root growth12, so it has long been widely believed that IAA and 2,4-D inhibit root growth through a similar mechanism. Later studies reported that IAA and 2,4-D elicit differential responses in regulating root growth. The root growth rate is regulated by two interrelated processes, namely, cell expansion and cell production28. 2,4-D mainly affects cell division, while IAA affects cell elongation29. In addition, studies have demonstrated that 2,4-D-induced inhibition of root or plant growth is related to cellular actin status29,30. 2,4-D can alter actin structure in long-term and short-term assays, and the effects of 2,4-D on actin filament organization and root growth were found to be mediated by synergistic interactions between SMAP1 and SCFTIR1 ubiquitin proteasome components, as studied with the 2,4-D-specific mutant aar1-1 and the ubiquitin–proteasome mutants tir1-1 and axr1-1231.
In this study, cis-acting element prediction showed an auxin response element, AuxRR-core, at − 108 to − 114 bp upstream of the AtSMAP2 gene. To investigate whether the AtSMAP2 gene promoter responds to auxin and whether it responds differentially to 2,4-D and IAA, the PAtSMAP2-997-GUS and PAtSMAP2-1993-GUS lines were treated with exogenous 2,4-D and IAA, respectively. The results of this study indicate that the PAtSMAP2-997 promoter is insensitive to IAA, while it can be induced to express by 2,4-D. The PAtSMAP2-1993 promoter is sensitive to IAA, which results in the inhibition of its expression activity, and insensitive to 2,4-D. The two promoters respond differently to both IAA and 2,4-D, but the exact location of their actions and the mechanisms are necessary to be further verified.
The AtSMAP2 gene promoter may also control this gene in response to ABA and MeJA
Judging from the predicted results of cis-acting elements and the differences in expression between the two promoters, it appears that AtSMAP2 may respond to abscisic acid, gibberellin, auxin, salicylic acid and MeJA. By overexpressing the AtSMAP1 and AtSMAP2 genes in aar1 mutants, it was found that AtSMAP2 functions similarly to AtSMAP1 and can respond to 2,4-D responses in Arabidopsis roots27, but how AtSMAP2 responds to other phytohormones remains to be studied.
It has been reported that several inducible genes contain conserved ABRE motifs (ACGT) in their promoter regions32. In this study, two ABRE motifs were present upstream of the AtSMAP2 gene and the PAtSMAP2-997-GUS and PAtSMAP2-1993-GUS lines were treated with exogenous ABA, which showed that both the PAtSMAP2-1993 and PAtSMAP2-997 promoters responded to ABA stress. ABA represses the expression activity of the PAtSMAP2-1993 and PAtSMAP2-997 promoters. MeJA induces the synthesis of defensive compounds and initiates the expression of genes associated with pathogenesis. Therefore, MeJA can be used to combat pathogens, salt stress, drought stress, low temperature, heavy metal stress and toxicity of other elements. In this study, two sites upstream of the AtSMAP2 gene promoter at -1150 and -1110 were predicted to have MeJA-responsive cis-acting elements. Treatment of PAtSMAP2-997-GUS and PAtSMAP2-1993-GUS strains by exogenous MeJA showed that the PAtSMAP2-997 promoter was insensitive to exogenous MeJA, whereas the PAtSMAP2-1993 promoter showed sensitivity to exogenous MeJA, which was caused by the probable reason that the PAtSMAP2- 997 promoter sequence does not contain MeJA-responsive cis-acting elements. Gene expression patterns may be a direct indicator of the role of promoters in stress and development. Both ABA and MeJA are phytohormones associated with plant stress response and the AtSMAP2 gene promoter exhibits a corresponding response to both of them, so it is hypothesized that the properties of this promoter may confer a role to the AtSMAP2 gene related to stress resistance.
Materials and methods
Experimental materials
The Arabidopsis thaliana ecotype Col-0, Agrobacterium strain GV3101, Escherichia coli strains TOP10 and DB3.1, and the plant expression vector pMDC162 plasmid used in this study were all provided by our laboratory. TOPO vectors (pENTR™ Directional TOPO® Cloning Kits) were purchased from Life Technologies, ClonExpress® II One Step Cloning Kit C112 was purchased from Vazyme, and GUS staining kits were purchased from Solarbio.
AtSMAP2 gene promoter sequence acquisition and bioinformatics analysis
According to the sequence of the AtSMAP2 gene (AT3G24280) in The Arabidopsis Information Resource (TAIR) (https://www.arabidopsis.org/), the fragment was obtained as a 1998 bp full-length promoter sequence upstream of the transcription initiation codon ATG. This sequence was analyzed using the online tools Plant Cis-Acting Regulatory DNA Elements (PLACE) (https://www.dna.afrc.go.jp/PLACE/?action=newplace)33 and Plant Cis-Acting Regulatory Element (PlantCARE) (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/)34 for the identification of cis-acting elements in the putative promoter.
Cloning of the AtSMAP2 promoter
Genomic DNA was extracted from fresh leaves of wild-type Arabidopsis Col-0 by the CTAB method35. Based on the prediction of the cis-acting element of the 1998 bp sequence upstream of the AtSMAP2 gene transcription start codon ATG, the 997 bp and 1993 bp sequences were intercepted as promoter sequences, respectively. The primers were designed according to the following sequences: PAtSMAP2-1993 –F (CACCAAGCTGTTTTGAAATTTATATCCAG); PAtSMAP2-997 –F (CACCTTGACTTTGAAGTAAAAGTTTGGAC); and PAtSMAP2-1993/PAtSMAP2-997 –R (AACAAAACCTAAGGATAAAGAGAAAG). The PCR reactions were performed using Arabidopsis genomic DNA as a template, and the reaction volume was 50 µL. The PCR conditions for amplifying AtSMAP2 promoters were as follows: 95 °C for 3 min; 35 cycles of 95 °C for 30 s, 60 °C for 30 s, 72 °C for 2 min; and then final extension at 72 °C for 5 min. After the PCR amplification product was recovered using a purification kit, ligated to the TOPO vector, and transformed into competent cells of E. coli TOP10 by the heat shock method, single colonies were picked at random, and the positive colonies were sent to the company for sequencing verification after colony PCR verification.
Construction of expression vectors and genetic transformation of Arabidopsis
The correctly sequenced TOPO vector and the plant expression vector pMDC162 plasmid were digested separately and recombined using the ClonExpress® II One Step Cloning Kit C112. The recombinant product was transformed into E. coli DB3.1 competent cells, and single colonies were randomly picked. After colony PCR verification, the positive colonies were sent to the company for sequencing verification.
The correct vector confirmed by sequencing is the constructed expression vector of the AtSMAP2 promoter driving the GUS gene, named PAtSMAP2-997-GUS and PAtSMAP2-1993-GUS. The above recombinant vector was transformed into Agrobacterium strain GV3101 competent cells, and Agrobacterium colonies were identified by PCR to obtain GV3101 strains carrying PAtSMAP2-997-GUS and PAtSMAP2-1993-GUS. Flower buds of Arabidopsis thaliana Col-0 ecotype at the flowering stage were then infested with the above bacterial solution using the floral dipping method36 and the T1 generation seeds were obtained.
Culture and identification of transgenic Arabidopsis
The seeds of Arabidopsis were first sterilized with 75% alcohol for 5 min, washed with sterile water, soaked in 30% NaClO for 10 min, and then washed with sterile water 5 to 6 times, and vernalized at 4 °C for 3 ~ 4 days. The T1 transformants were selected on the half-strength Murashige and Skoog (MS) plates containing hygromycin at a final concentration of 50 mg/L. And then these plants were cultured in a growth chamber with a light intensity of 120 μmol·m−2 s−1, a humidity of approximately 60%, 24 °C light for 16 h, and 20 °C darkness for 8 h. The regenerated T1 seedlings with hygromycin resistance were confirmed by PCR amplification using a promoter-GUS specific primer (PAtSMAP2-1993-GUS/PAtSMAP2-997-GUS-R: 5′CTGCTTTTTCTTGCCGTTTTCGTCG3′; PAtSMAP2-1993-GUS-F: 5′GTACTCTTGATCTTTATTACCTTGG3′, the length of the amplification product was 1524 bp; PAtSMAP2-997-GUS-F: 5′TGTTGATGACACATAAAGAAAGAAA3′, the length of the amplification product was 638 bp) and then transplanted into the soil and grown to maturity in a greenhouse. The T2-generation transgenic lines that displayed a segregation ratio of 3:1 by hygromycin resistance screening were used for propagation. Finally, the homozygous transgenic lines with a single insertion of the AtSMAP2 promoter-GUS were screened for functional analyses through segregation ratio analysis.
Treatments with different concentrations of 2,4-D, IAA, ABA and MeJA
Wild-type (Col-0) and transgenic seedlings were grown on MS medium solidified with 0.8% agar in a growth chamber for 7 days. Twenty seedlings per transgenic line were used for each treatment. The hormone treatments were carried out for 12 h and involved culturing 7-day-old seedlings in liquid MS medium containing 50nM, 100nM, 150nM 2,4-D and IAA, 25µM, 50µM, 100µM ABA, 1µM, 25µM, and 50µM MeJA, respectively, while control seedlings were cultured in liquid MS medium without any hormones.
GUS histochemical staining and quantitative real-time PCR (qRT‒PCR) analysis
According to the GUS staining kit, the transgenic Arabidopsis seedlings were completely immersed in GUS staining solution, incubated at 37 °C overnight, and then seedlings were moved to 75% ethanol for decolorization, observed for GUS staining, and photographed for preservation.
The total RNA of Arabidopsis thaliana was extracted with a Spectrum Plant Total RNA Kit (Sigma‒Aldrich, Germany), and reverse transcription synthesis was performed using StarScript II First-strand cDNA Synthesis Mix with gDNA Remover (GenStar, China). The enzyme used for qRT‒PCR was 2 × RealStar Green Fast Mixture with ROX (GenStar, China), and the equipment used was a StepOnePlus™ real-time PCR device (Thermo Fisher Scientific, Applied Biosystems, China). The primers for GUS were F: 5′- CGAACTGAACTGGCAGACTATCC-3′ and R: 5′- CGGCGTGGTGTAGAGCATTAC-3'; and the expression of Actin2 was used as an internal control, the primers for Actin2 were F: 5'-ACGAACTGAACTGGCAGACTATCC-3' and R: 5'-TCGGCGTGGTGTAGAGCATTAC-3'. Relative gene expression was calculated using the 2−ΔΔCt method, with Ct values averaged over three technical replicates, and the relative expression was the average of three biological replicates.
Data availability
The datasets analysed during the current study are available in the The Arabidopsis Information Resource (TAIR) (BioSample number: AT3G24280), https://www.arabidopsis.org/servlets/TairObject?id=37851&type=locus.
References
Odell, J. T., Nagy, F. & Chua, N. H. Identification of DNA sequences required for activity of the cauliflower mosaic virus 35S promoter. Nature 313, 810–812. https://doi.org/10.1038/313810a0 (1985).
Christensen, A. H., Sharrock, R. A. & Quail, P. H. Maize polyubiquitin genes: structure, thermal perturbation of expression and transcript splicing, and promoter activity following transfer to protoplasts by electroporation. Plant Mol. Biol. 18, 675–689. https://doi.org/10.1007/bf00020010 (1992).
Cornejo, M. J., Luth, D., Blankenship, K. M., Anderson, O. D. & Blechl, A. E. Activity of a maize ubiquitin promoter in transgenic rice. Plant Mol. Biol. 23, 567–581. https://doi.org/10.1007/bf00019304 (1993).
Gilmartin, P. M. & Chua, N. H. Spacing between GT-1 binding sites within a light-responsive element is critical for transcriptional activity. Plant Cell 2, 447–455. https://doi.org/10.1105/tpc.2.5.447 (1990).
Potenza, C., Aleman, L. & Sengupta-Gopalan, C. Targeting transgene expression in research, agricultural, and environmental applications: Promoters used in plant transformation. In Vitro Cell. Dev. Biol. Plant 40, 1–22. https://doi.org/10.1079/IVP2003477 (2004).
Xu, W. L. et al. Cotton PRP5 gene encoding a proline-rich protein is involved in fiber development. Plant Mol. Biol. 82, 353–365. https://doi.org/10.1007/s11103-013-0066-8 (2013).
Hirai, T., Kim, Y. W., Kato, K., Hiwasa-Tanase, K. & Ezura, H. Uniform accumulation of recombinant miraculin protein in transgenic tomato fruit using a fruit-ripening-specific E8 promoter. Transg. Res. 20, 1285–1292. https://doi.org/10.1007/s11248-011-9495-9 (2011).
Jiang, P. et al. Characterization of a strong and constitutive promoter from the Arabidopsis serine carboxypeptidase-like gene AtSCPL30 as a potential tool for crop transgenic breeding. BMC Biotechnol. 18, 59. https://doi.org/10.1186/s12896-018-0470-x (2018).
Efremova, L. N., Strelnikova, S. R., Gazizova, G. R., Minkina, E. A. & Komakhin, R. A. A synthetic strong and constitutive promoter derived from the Stellaria media pro-SmAMP1 and pro-SmAMP2 promoters for effective transgene expression in plants. Genes https://doi.org/10.3390/genes11121407 (2020).
Basak, P. et al. Functional characterization of two myo-inositol-1-phosphate synthase (MIPS) gene promoters from the halophytic wild rice (Porteresia coarctata). Planta 248, 1121–1141. https://doi.org/10.1007/s00425-018-2957-z (2018).
Li, Y. et al. Cloning and function analysis of an alfalfa (Medicago sativa L.) zinc finger protein promoter MsZPP. Mol. Biol. Rep. 39, 8559–8569. https://doi.org/10.1007/s11033-012-1712-y (2012).
Maher, E. P. & Martindale, S. J. Mutants of Arabidopsis thaliana with altered responses to auxins and gravity. Biochem. Genet. 18, 1041–1053. https://doi.org/10.1007/bf00484337 (1980).
Stasinopoulos, T. C. & Hangarter, R. P. Preventing photochemistry in culture media by long-pass light filters alters growth of cultured tissues. Plant Physiol. 93, 1365–1369. https://doi.org/10.1104/pp.93.4.1365 (1990).
Campanoni, P. & Nick, P. Auxin-dependent cell division and cell elongation. 1-Naphthaleneacetic acid and 2,4-dichlorophenoxyacetic acid activate different pathways. Plant Physiol. 137, 939–948. https://doi.org/10.1104/pp.104.053843 (2005).
Delbarre, A., Muller, P., Imhoff, V. & Guern, J. Comparison of mechanisms controlling uptake and accumulation of 2,4-dichlorophenoxy acetic acid, naphthalene-1-acetic acid, and indole-3-acetic acid in suspension-cultured tobacco cells. Planta 198, 532–541. https://doi.org/10.1007/bf00262639 (1996).
Jackson, R. G. et al. Over-expression of an Arabidopsis gene encoding a glucosyltransferase of indole-3-acetic acid: Phenotypic characterisation of transgenic lines. Plant J. Cell Mol. Biol. 32, 573–583. https://doi.org/10.1046/j.1365-313x.2002.01445.x (2002).
Staswick, P. E. et al. Characterization of an Arabidopsis enzyme family that conjugates amino acids to indole-3-acetic acid. Plant Cell 17, 616–627. https://doi.org/10.1105/tpc.104.026690 (2005).
Mockaitis, K. & Estelle, M. Auxin receptors and plant development: a new signaling paradigm. Annu. Rev. Cell Dev. Biol. 24, 55–80. https://doi.org/10.1146/annurev.cellbio.23.090506.123214 (2008).
Dharmasiri, N. et al. Plant development is regulated by a family of auxin receptor F box proteins. Dev. Cell 9, 109–119. https://doi.org/10.1016/j.devcel.2005.05.014 (2005).
Lau, S., Jürgens, G. & De Smet, I. The evolving complexity of the auxin pathway. Plant Cell 20, 1738–1746. https://doi.org/10.1105/tpc.108.060418 (2008).
Guilfoyle, T. J. & Hagen, G. Auxin response factors. Curr. Opin. Plant Biol. 10, 453–460. https://doi.org/10.1016/j.pbi.2007.08.014 (2007).
Oono, Y. et al. p-Chlorophenoxyisobutyric acid impairs auxin response in Arabidopsis root. Plant Physiol. 133, 1135–1147. https://doi.org/10.1104/pp.103.027847 (2003).
Biswas, K. K. et al. Genetic characterization of mutants resistant to the antiauxin p-chlorophenoxyisobutyric acid reveals that AAR3, a gene encoding a DCN1-like protein, regulates responses to the synthetic auxin 2,4-dichlorophenoxyacetic acid in Arabidopsis roots. Plant Physiol. 145, 773–785. https://doi.org/10.1104/pp.107.104844 (2007).
Rahman, A. et al. A small acidic protein 1 (SMAP1) mediates responses of the Arabidopsis root to the synthetic auxin 2,4-dichlorophenoxyacetic acid. Plant J. Cell Mol. Biol. 47, 788–801. https://doi.org/10.1111/j.1365-313X.2006.02832.x (2006).
Nakasone, A. et al. SMALL ACIDIC PROTEIN1 acts with RUB modification components, the COP9 signalosome, and AXR1 to regulate growth and development of Arabidopsis. Plant Physiol. 160, 93–105. https://doi.org/10.1104/pp.111.188409 (2012).
Gray, W. M. & Estelle, I. Function of the ubiquitin-proteasome pathway in auxin response. Trends Biochem. Sci. 25, 133–138. https://doi.org/10.1016/s0968-0004(00)01544-9 (2000).
Nakasone, A. et al. A gene encoding SMALL ACIDIC PROTEIN 2 potentially mediates the response to synthetic auxin, 2,4-dichlorophenoxyacetic acid, Arabidopsis thaliana. J. Plant Physiol. 166, 1307–1313. https://doi.org/10.1016/j.jplph.2009.02.005 (2009).
Beemster, G. T. & Baskin, T. I. Analysis of cell division and elongation underlying the developmental acceleration of root growth in Arabidopsis thaliana. Plant Physiol. 116, 1515–1526. https://doi.org/10.1104/pp.116.4.1515 (1998).
Rahman, A. et al. Auxin, actin and growth of the Arabidopsis thaliana primary root. Plant J. Cell Mol. Biol. 50, 514–528. https://doi.org/10.1111/j.1365-313X.2007.03068.x (2007).
Rodríguez-Serrano, M. et al. 2,4-Dichlorophenoxyacetic acid promotes S-nitrosylation and oxidation of actin affecting cytoskeleton and peroxisomal dynamics. J. Exp. Bot. 65, 4783–4793. https://doi.org/10.1093/jxb/eru237 (2014).
Takahashi, M. et al. Small acidic protein 1 and SCF(TIR)(1) ubiquitin proteasome pathway act in concert to induce 2,4-dichlorophenoxyacetic acid-mediated alteration of actin in Arabidopsis roots. Plant J. Cell Mol. Biol. 89, 940–956. https://doi.org/10.1111/tpj.13433 (2017).
Grill, E. & Himmelbach, A. ABA signal transduction. Curr. Opin. Plant Biol. 1, 412–418. https://doi.org/10.1016/s1369-5266(98)80265-3 (1998).
Higo, K., Ugawa, Y., Iwamoto, M. & Korenaga, T. Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res. 27, 297–300. https://doi.org/10.1093/nar/27.1.297 (1999).
Lescot, M. et al. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 30, 325–327. https://doi.org/10.1093/nar/30.1.325 (2002).
Chen, K. S., Li, F., Xu, C. J., Zhang, S. L. & Fu, C. X. An efficient macro-method of genomic DNA isolation from Actinidia chinensis leaves. Yichuan Hereditas 26, 529–531 (2004).
Clough, S. J. & Bent, A. F. Floral dip: A simplified method for agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. Cell Mol. Biol. 16, 735–743. https://doi.org/10.1046/j.1365-313x.1998.00343.x (1998).
Acknowledgements
This research was supported by the Department of Jilin Province Science and Technology (20230402020GH).
Author information
Authors and Affiliations
Contributions
AB conducted experiments and wrote the manuscript. TJ and TH conducted experiments and analyzed the data. KC, WY, YL, HZ, JZ and SH conducted experiments. MW conceived and designed research. All the authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bao, A., Jiao, T., Hu, T. et al. Cloning of the Arabidopsis SMAP2 promoter and analysis of its expression activity. Sci Rep 14, 11451 (2024). https://doi.org/10.1038/s41598-024-61525-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-61525-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.