Abstract
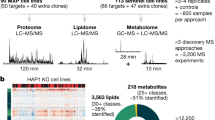
Isolated complex I (CI) deficiencies are a major cause of primary mitochondrial disease. A substantial proportion of CI deficiencies are believed to arise from defects in CI assembly factors (CIAFs) that are not part of the CI holoenzyme. The biochemistry of these CIAFs is poorly defined, making their role in CI assembly unclear, and confounding interpretation of potential disease-causing genetic variants. To address these challenges, we devised a deep mutational scanning approach to systematically assess the function of thousands of NDUFAF6 genetic variants. Guided by these data, biochemical analyses and cross-linking mass spectrometry, we discovered that the CIAF NDUFAF6 facilitates incorporation of NDUFS8 into CI and reveal that NDUFS8 overexpression rectifies NDUFAF6 deficiency. Our data further provide experimental support of pathogenicity for seven novel NDUFAF6 variants associated with human pathology and introduce functional evidence for over 5,000 additional variants. Overall, our work defines the molecular function of NDUFAF6 and provides a clinical resource for aiding diagnosis of NDUFAF6-related diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All unique/stable reagents generated in this study are available upon request with a materials transfer agreement. The raw next-generation sequencing data from the DMS experiment have been deposited in the Sequence Read Archive (BioProject accession PRJNA1007392). DMS variant and count data have been deposited on MaveDB (MaveDB accession urn:mavedb:00000663-a). The raw proteomics data from the XL-MS experiment have been deposited in the MassIVE repository (MassIVE ID MSV000094276). All deposited data are publicly available. Source data are provided with this paper.
Code availability
All original code has been deposited on GitHub (https://github.com/AYSung/af6-dms) and is publicly available.
References
Sazanov, L. A. A giant molecular proton pump: structure and mechanism of respiratory complex I. Nat. Rev. Mol. Cell Biol. 16, 375–388 (2015).
Kirby, D. M. et al. Respiratory chain complex I deficiency: an underdiagnosed energy generation disorder. Neurology 52, 1255–1264 (1999).
Schon, K. R. et al. Use of whole genome sequencing to determine genetic basis of suspected mitochondrial disorders: cohort study. Br. Med. J. 375, e066288 (2021).
Nouws, J., Nijtmans, L. G. J., Smeitink, J. A. & Vogel, R. O. Assembly factors as a new class of disease genes for mitochondrial complex I deficiency: cause, pathology and treatment options. Brain 135, 12–22 (2012).
Swalwell, H. et al. Respiratory chain complex I deficiency caused by mitochondrial DNA mutations. Eur. J. Hum. Genet. 19, 769–775 (2011).
Formosa, L. E., Dibley, M. G., Stroud, D. A. & Ryan, M. T. Building a complex complex: assembly of mitochondrial respiratory chain complex I. Semin. Cell Dev. Biol. 76, 154–162 (2018).
Rensvold, J. W. et al. Defining mitochondrial protein functions through deep multiomic profiling. Nature 606, 382–388 (2022).
Formosa, L. E. et al. Optic atrophy-associated TMEM126A is an assembly factor for the ND4-module of mitochondrial complex I. Proc. Natl. Acad. Sci. 118, e2019665118 (2021).
Jackson, T. D. et al. Sideroflexin 4 is a complex I assembly factor that interacts with the MCIA complex and is required for the assembly of the ND2 module. Proc. Natl Acad. Sci. USA 119, e2115566119 (2022).
Dibley, M. G. et al. The mitochondrial acyl-carrier protein interaction network highlights important roles for LYRM family members in complex I and mitoribosome assembly. Mol. Cell. Proteom. 19, mcp.RA119.001784 (2019).
Ribeiro, A. J. M. et al. Emerging concepts in pseudoenzyme classification, evolution, and signaling. Sci. Signal 12, eaat9797 (2019).
Jeffery, C. J. The demise of catalysis, but new functions arise: pseudoenzymes as the phoenixes of the protein world. Biochem. Soc. Trans. 47, 371–379 (2019).
Frazier, A. E., Thorburn, D. R. & Compton, A. G. Mitochondrial energy generation disorders: genes, mechanisms, and clues to pathology. J. Biol. Chem. 294, 5386–5395 (2019).
Schon, K. R., Ratnaike, T., Ameele, J., van den, Horvath, R. & Chinnery, P. F. Mitochondrial diseases: a diagnostic revolution. Trends Genet. 36, 702–717 (2020).
Fowler, D. M. & Fields, S. Deep mutational scanning: a new style of protein science. Nat. Methods 11, 801–807 (2014).
Marchler-Bauer, A. et al. CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 45, D200–D203 (2017).
Lemire, B. D. Evolution, structure and membrane association of NDUFAF6, an assembly factor for NADH:ubiquinone oxidoreductase (complex I). Mitochondrion 35, 13–22 (2017).
Pagliarini, D. J. et al. A mitochondrial protein compendium elucidates complex I disease biology. Cell 134, 112–123 (2008).
McKenzie, M. et al. Mutations in the gene encoding C8orf38 block complex I assembly by inhibiting production of the mitochondria-encoded subunit ND1. J. Mol. Biol. 414, 413–426 (2011).
Stroud, D. A. et al. Accessory subunits are integral for assembly and function of human mitochondrial complex I. Nature 538, 123–126 (2016).
Robinson, B. H., Petrova-Benedict, R., Buncic, J. R. & Wallace, D. C. Nonviability of cells with oxidative defects in galactose medium: a screening test for affected patient fibroblasts. Biochem. Med. Metab. Biol. 48, 122–126 (1992).
Gohil, V. M. et al. Nutrient-sensitized screening for drugs that shift energy metabolism from mitochondrial respiration to glycolysis. Nat. Biotechnol. 28, 249–255 (2010).
Faure, A. J., Schmiedel, J. M., Baeza-Centurion, P. & Lehner, B. DiMSum: an error model and pipeline for analyzing deep mutational scanning data and diagnosing common experimental pathologies. Genome Biol. 21, 207 (2020).
Fukasawa, Y. et al. MitoFates: improved prediction of mitochondrial targeting sequences and their cleavage sites. Mol. Cell. Proteom. 14, 1113–1126 (2015).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Varadi, M. et al. AlphaFold protein structure database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2021).
Imai, K. & Mitaku, S. Mechanisms of secondary structure breakers in soluble proteins. Biophysics 1, 55–65 (2005).
Henikoff, S. & Henikoff, J. G. Amino acid substitution matrices from protein blocks. Proc. Natl Acad. Sci. 89, 10915–10919 (1992).
Lin, F. Y. et al. Mechanism of action and inhibition of dehydrosqualene synthase. Proc. Natl Acad. Sci. USA 107, 21337–21342 (2010).
Gautier, R., Douguet, D., Antonny, B. & Drin, G. HELIQUEST: a web server to screen sequences with specific α-helical properties. Bioinformatics 24, 2101–2102 (2008).
Oughtred, R. et al. The BioGRID database: a comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci. 30, 187–200 (2021).
Guerrero-Castillo, S. et al. The assembly pathway of mitochondrial respiratory chain complex I. Cell Metab. 25, 128–139 (2017).
Alston, C. L. et al. Pathogenic bi-allelic mutations in NDUFAF8 cause Leigh syndrome with an isolated complex I deficiency. Am. J. Hum. Genet. 106, 92–101 (2020).
Guarani, V. et al. TIMMDC1/C3orf1 functions as a membrane-embedded mitochondrial complex I assembly factor through association with the MCIA complex. Mol. Cell. Biol. 34, 847–861 (2014).
Kohda, M. et al. A comprehensive genomic analysis reveals the genetic landscape of mitochondrial respiratory chain complex deficiencies. PLoS Genet. 12, e1005679 (2016).
Lake, N. J., Compton, A. G., Rahman, S. & Thorburn, D. R. Leigh syndrome: one disorder, more than 75 monogenic causes. Ann. Neurol. 79, 190–203 (2016).
McCormick, E. M. et al. Expert panel curation of 113 primary mitochondrial disease genes for the Leigh syndrome spectrum. Ann. Neurol. 94, 696–712 (2023).
Landrum, M. J. et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 46, D1062–D1067 (2017).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–423 (2015).
Brnich, S. E. et al. Recommendations for application of the functional evidence PS3/BS3 criterion using the ACMG/AMP sequence variant interpretation framework. Genome Med .12, 3 (2019).
Couvillion, M. T., Soto, I. C., Shipkovenska, G. & Churchman, L. S. Synchronized mitochondrial and cytosolic translation programs. Nature 533, 499–503 (2016).
Lee, et al.Transcriptome analysis of complex I-deficient patients reveals distinct expression programs for subunits and assembly factors of the oxidative phosphorylation system. BMC Genomics 16, 691 (2015).
Sung, A. Y., Floyd, B. J. & Pagliarini, D. J. Systems biochemistry approaches to defining mitochondrial protein function. Cell Metab. 31, 669–678 (2020).
Forny, P. et al. Diagnosing mitochondrial disorders remains challenging in the omics era. Neurol. Genet. 7, e597 (2021).
Pronicka, E. et al. New perspective in diagnostics of mitochondrial disorders: two years’ experience with whole-exome sequencing at a national paediatric centre. J. Transl. Med. 14, 174 (2016).
Taylor, R. W. et al. Use of whole-exome sequencing to determine the genetic basis of multiple mitochondrial respiratory chain complex deficiencies. J. Am. Med. Assoc. 312, 68–77 (2014).
Theunissen, T. E. J. et al. Whole exome sequencing is the preferred strategy to identify the genetic defect in patients with a probable or possible mitochondrial cause. Front. Genet. 9, 400 (2018).
Wortmann, S. B., Koolen, D. A., Smeitink, J. A., Heuvel, L. & Rodenburg, R. J. Whole exome sequencing of suspected mitochondrial patients in clinical practice. J. Inherit. Metab. Dis. 38, 437–443 (2015).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
VanderPlas, J. et al. Altair: interactive statistical visualizations for python. J. Open Source Softw. 3, 1057 (2018).
Frezza, C., Cipolat, S. & Scorrano, L. Organelle isolation: functional mitochondria from mouse liver, muscle and cultured fibroblasts. Nat. Protoc. 2, 287–295 (2007).
Wittig, I., Braun, H. P. & Schagger, H. Blue native PAGE. Nat. Protoc. 1, 418–428 (2006).
Schertl, P. & Braun, H.-P. Plant mitochondria, methods and protocols. Methods Mol. Biol. 1305, 131–138 (2015).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Gietz, R. D. & Schiestl, R. H. High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 31–34 (2007).
James, P., Halladay, J. & Craig, E. A. Genomic libraries and a host strain designed for highly efficient two-hybrid selection in yeast. Genetics 144, 1425–1436 (1996).
Yau, E. H. & Rana, T. M. Next generation sequencing, methods and protocols. Methods Mol. Biol. 1712, 203–216 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Cock, P. J. A. et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423 (2009).
Mölder, F. et al. Sustainable data analysis with Snakemake. F1000Research 10, 33 (2021).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Alston, C. L. et al. A recurrent mitochondrial p.Trp22Arg NDUFB3 variant causes a distinctive facial appearance, short stature and a mild biochemical and clinical phenotype. J. Med. Genet. 53, 634–641 (2016).
Legati, A. et al. New genes and pathomechanisms in mitochondrial disorders unraveled by NGS technologies. Biochim. Biophys. Acta Bioenerg. 1857, 1326–1335 (2016).
Alston, C. L. et al. Bi-allelic mutations in NDUFA6 establish its role in early-onset isolated mitochondrial complex I deficiency. Am. J. Hum. Genet. 103, 592–601 (2018).
Ogawa, E. et al. Clinical validity of biochemical and molecular analysis in diagnosing Leigh syndrome: a study of 106 Japanese patients. J. Inherit. Metab. Dis. 40, 685–693 (2017).
Frazier, A. E., Vincent, A. E., Turnbull, D. M., Thorburn, D. R. & Taylor, R. W. Assessment of mitochondrial respiratory chain enzymes in cells and tissues. Methods Cell. Biol. 155, 121–156 (2020).
Ioannidis, N. M. et al. REVEL: an ensemble method for predicting the pathogenicity of rare missense variants. Am. J. Hum. Genet. 99, 877–885 (2016).
Yariv, B. et al. Using evolutionary data to make sense of macromolecules with a ‘face‐lifted’ ConSurf. Protein Sci. 32, e4582 (2023).
Jurrus, E. et al. Improvements to the APBS biomolecular solvation software suite. Protein Sci. 27, 112–128 (2018).
Wheeler, T. J., Clements, J. & Finn, R. D. Skylign: a tool for creating informative, interactive logos representing sequence alignments and profile hidden Markov models. BMC Bioinform. 15, 7 (2014).
Acknowledgements
We thank the members of the Pagliarini and Keck laboratories as well as B. J. Floyd and R. D. Mitra for their helpful discussion and critical evaluation of the manuscript. This work was supported by NIH award nos R35GM131795 (to D.J.P.), T32GM140935 (to A.Y.S. and L.H.S.), T32AG000213 (to A.Y.S.) and T32GM008505 (to L.H.S.), as well as funds from the BJC Investigator Program (to D.J.P.), NIHR award no. PDF-2018-11-ST2-021 (to C.L.A.) and funds from the Wellcome Centre for Mitochondrial Research grant no. 203105/Z/16/Z, the Mitochondrial Disease Patient Cohort grant no. G0800674 and the Medical Research Council International Centre for Genomic Medicine in Neuromuscular Disease grant no. MR/S005021/1 (to R.W.T.). This work was partly supported by the Practical Research Project for Rare/Intractable Diseases from the Japan Agency for Medical Research and Development (grant no. JP23ek0109625; to K.M., Y.O. and M.S.) and by the Italian Ministry of Health (grant no. ERP-2019-23671045; to D.G.) and German Federal Ministry of Education and Research (grant no. 01GM1920A; to H. P.) through the European Joint Programme on Rare Diseases project GENOMIT. We thank the Genome Technology Access Center at the McDonnell Genome Institute at Washington University School of Medicine for help with genomic analysis. The Center is partially supported by NCI Cancer Center Support Grant no. P30 CA91842 to the Siteman Cancer Center from the National Center for Research Resources (NCRR), a component of the National Institutes of Health (NIH), and NIH Roadmap for Medical Research. This publication is solely the responsibility of the authors and does not necessarily represent the official view of NCRR or NIH, nor the NHS, the NIHR or the Department of Health and Social Care.
Author information
Authors and Affiliations
Contributions
A.Y.S. and D.J.P. conceived of the project and its design. A.Y.S. performed the deep mutational scanning, yeast two-hybrid assays, ultracentrifugation experiments and stable cell line generation. A.Y.S. and R.M.G. performed the blue native western blots and in-gel activity assays. R.M.G. performed the cross-linking mass spectrometry. L.H.S. performed the galactose growth assays. C.L.A., K.M., Y.O., M.S., H.P., D.G., A.T., R.C., A.R. and R.W.T. contributed the clinicopathological data relating to patients with mitochondrial disease and NDUFAF6 variants. D.J.P. and J.L.K. supervised the project. A.Y.S., R.M.G., L.H.S. and D.J.P. wrote this paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Douglas Fowler, David Thorburn and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Alfredo Giménez-Cassina, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Full DMS dataset.
Heatmap representation of the DMS data. The residue position is labeled along the horizontal axis (missing residues in the dataset denoted by a black triangle), the amino acid substitution is labeled on the vertical axis. The color of the rectangles represents the fitness score of the variant. Gray dots mark the wild-type amino acid at each position. Additional annotations to aid interpretation are shown above and below the fitness score heatmaps. Conservation at each position (calculated using ConSurf) is shown using the ConSurf color scale. Predicted alpha-helical secondary structure based on the AlphaFold AF6 model is shown as a thick, grey line. Percent solvent-accessible surface area (% SASA) at each residue position, calculated from the AlphaFold AF6 model is shown as a bar graph.
Extended Data Fig. 2 Experimental design and quality control.
a. Experimental design of DMS experiment. AF6 KO1 and AF6 KO2 cells are split into five replicates and separately transduced with the AF6 variant library. Input samples are collected 6–8 days after transduction (with 4-5 days of selection with puromycin). Cells are then cultured in galactose media to select for cells expressing functional variants of AF6. Output samples are collected after three and six passages in galactose media. b. Histogram of input read count distribution of the variants by replicate. The dashed line represents a read depth of 100x. c. Binned scatterplot of fitness scores from individual replicates. The dot size represents the number of variants in each fitness score bin. Pearson correlation coefficients (r) for each pairwise comparison is provided in the upper left corner of each plot. d. Bayesian information criterion (BIC) scores for Gaussian mixture models (GMM) with 1–6 components. A three-component model (shown in red) was chosen for this analysis. e. Histogram of DMS fitness scores overlaid with a three-component GMM. The three components are represented as colored dashed lines while the overall model is represented by a solid black line. The component weight, mean, and standard deviation are shown in the table below. f. Histogram of DMS fitness scores of nonsense variants overlaid with the three components from the GMM. The red component likely represents variants with a strong functional impact. The red shaded region represents three standard deviations from the mean of the red component, [−1.729, −0.842], and encompasses the fitness scores for 274 out of 286 nonsense variants.
Extended Data Fig. 3 Comparisons of DMS data to predicted NDUFAF6 structure.
a. Schematic for the calculation of mutational sensitivity. At each residue position, the number of missense substitutions with a fitness ≤ −0.842 (threshold for variants with strong functional impact, see Extended Data Fig. 2f) are counted and scaled linearly to a value between 0 and 9. b. Mutational sensitivity and percent solvent-accessible surface area (% SASA) of helix 11 (residues 253–275) are shown as bars. A cartoon representation of this alpha helix from the AF6 AlphaFold model is shown below and colored by mutational sensitivity. c. Density plot of DMS fitness scores for proline substitutions grouped by predicted secondary structure. d. Binned scatterplot showing the distribution of percent solvent accessible surface area and mutational sensitivity of residues in AF6. The dot size represents the number of residues in each bin. e. Cartoon representation of the AF6 AlphaFold model colored by mutational sensitivity (top) and % SASA (bottom). A front view and a back view are shown.
Extended Data Fig. 4 Biochemical principles recapitulated by DMS.
a-d. DMS fitness scores of substitutions in alanine residues (a), aromatic residues (b), hydrophobic residues (c), and charged residues (d). BLOSUM62 similarity scores for each substitution is shown along the right. Predicted secondary structure is shown below, conservation (calculated using ConSurf), and percent solvent accessible surface area (% SASA) are shown below. e. Surface representation of AF6 AlphaFold model colored by conservation, mutational sensitivity, and electrostatics. f. Binned scatterplot showing the distribution of conservation (calculated by ConSurf) and mutational sensitivity of residues in AF6. The dot size represents the number of residues in each bin.
Extended Data Fig. 5 Additional XL-MS hits.
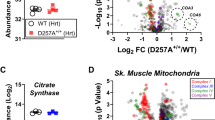
Volcano plot showing the mean log2 fold-change and the −log10 of the two-tailed Student’s t-test p-value (no adjustment for multiple comparisons) of proteins in mitochondria overexpressing FLAG-tagged AF6 compared to wild-type mitochondria, n = 4 biologically independent samples per condition. CI subunits and assembly factors are marked as red dots and labeled. The most significant hits are labeled in the magnified portion of the volcano plot.
Supplementary information
Supplementary Tables
Supplementary Table 1. DMS fitness and fitness estimate errors for the 5,714 AF6 variants tested in this study. The probability of abnormal or normal function (Pabnormal and Pnormal, respectively), as well as the resulting functional classification for each variant is also noted. Supplementary Table 2. Additional annotations for each residue position of AF6 to aid interpretation of DMS data. For each residue in the DMS variant library, the conservation (determined by ConSurf), the solvent-accessible surface area (relative percentage and absolute area), mutational sensitivity and the alpha-helical secondary structure are provided. Supplementary Table 3. All proteins identified in the XL-MS experiment. The mean log2(fold change) for each protein in the AF6-FLAG overexpression condition relative to the wild-type control and the −log10 of the two-tailed Student’s t-test P value (no adjustment for multiple comparisons) are provided, n = 4 biologically independent samples per condition. Supplementary Table 4. Control ClinVar variants. The genomic location, gnomAD allele frequency data, ClinVar accession and classification, ACMG criteria applied and resultant ACMG classification, DMS fitness and functional classification and REVEL score (in silico prediction) for the 14 non-VUS variants on ClinVar used as control variants in this study are described. In addition, updated ACMG classification upon incorporation of DMS functional data is also provided for each control variant. Supplementary Table 5. Calculation of OddsPath values for pathogenic and benign assessment of variant effect as described by the ClinGen Sequence Variant Interpretation Working Group. Supplementary Table 6. Patient information. The patient’s clinical presentation, biochemical data, genomic location of the variants, ClinVar accession and classification, DMS fitness and functional classification, REVEL score (in silico prediction) and ACMG criteria applied and resultant ACMG classification are provided for the 18 patients reported in this manuscript. Supplementary Table 7. Patient respiratory chain (RC) enzyme activity data. Absolute RC activity is normalized to control citrate synthase (CS) values. Relative RC activity is relative to the control samples. Reference ranges for each cohort (separated by double lines) are provided as the mean ± standard deviation. Patient genotypes and the tissue used for the assays are also indicated. Variant calls are based on RefSeq NM_152416.4. Note that the second column in the last cohort measures CI + CIII activity instead of CII activity. Supplementary Table 8. Cell lines used in this study. Supplementary Table 9. Plasmids used in this study. Supplementary Table 10. Oligonucleotides used in this study. Supplementary Table 11. Antibodies used in this study.
Source data
Source Data Fig. 1
Unprocessed immunoblot and in-gel activity assay.
Source Data Fig. 1
Seahorse OCR measurements and growth constant estimates.
Source Data Fig. 2
Residue numbers for AF6 features and mutational sensitivity and conservation for AF6 surface patches.
Source Data Fig. 3
Unprocessed yeast two-hybrid spot plate images and unprocessed blue native western blots.
Source Data Fig. 4
Unprocessed immunoblots.
Source Data Fig. 4
Cell counts and growth constant estimates.
Source Data Fig. 5
Unprocessed immunoblots.
Source Data Fig. 5
Gaussian mixture model for clinical analysis.
Source Data Extended Data Fig. 2
Variant input read counts by replicate, variant fitness estimates by replicate, Bayesian information criterion for Gaussian mixture model and Gaussian mixture model probability density function.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sung, A.Y., Guerra, R.M., Steenberge, L.H. et al. Systematic analysis of NDUFAF6 in complex I assembly and mitochondrial disease. Nat Metab (2024). https://doi.org/10.1038/s42255-024-01039-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42255-024-01039-2