Abstract
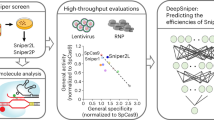
Finely tuned enzymatic pathways control cellular processes, and their dysregulation can lead to disease. Developing predictive and interpretable models for these pathways is challenging because of the complexity of the pathways and of the cellular and genomic contexts. Here we introduce Elektrum, a deep learning framework that addresses these challenges with data-driven and biophysically interpretable models for determining the kinetics of biochemical systems. First, it uses in vitro kinetic assays to rapidly hypothesize an ensemble of high-quality kinetically interpretable neural networks (KINNs) that predict reaction rates. It then employs a transfer learning step, where the KINNs are inserted as intermediary layers into deeper convolutional neural networks, fine-tuning the predictions for reaction-dependent in vivo outcomes. We apply Elektrum to predict CRISPR–Cas9 off-target editing probabilities and demonstrate that Elektrum achieves improved performance, regularizes neural network architectures and maintains physical interpretability.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All training and testing data presented in this paper are publicly available. The in vitro kinetic data were downloaded from ref. 20 (https://github.com/finkelsteinlab/nucleaseq/tree/master). The in vivo CIRSPR–Cas9 off-targeting dataset was downloaded from CRISPR-Net29 (https://codeocean.com/capsule/9553651/tree/v2). Source data is available with this paper.
Code availability
The Elektrum code is available on GitHub at https://github.com/zj-zhang/Elektrum and Zenodo at https://doi.org/10.5281/zenodo.8044859 ref. 51.
References
Gebauer, F., Schwarzl, T., Valcárcel, J. & Hentze, M. W. RNA-binding proteins in human genetic disease. Nat. Rev. Genet. 22, 185–198 (2021).
Masoud, G. N. & Li, W. Hif-1α pathway: role, regulation and intervention for cancer therapy. Acta Pharm. Sin. B 5, 378–389 (2015).
Santamaria, S. & Groot, R. ADAMTS proteases in cardiovascular physiology and disease. Open Biol. 10, 200333 (2020).
Flinn, A.M. & Gennery, A.R. Adenosine deaminase deficiency: a review. Orphanet J. Rare Dis. 13, 65 (2018).
Kim, R. Q. et al. Kinetic analysis of multistep USP7 mechanism shows critical role for target protein in activity. Nat. Commun. 10, 231 (2019).
Persikov, A. V. et al. A systematic survey of the Cys2His2 zinc finger DNA-binding landscape. Nucleic Acids Res. 43, 1965–1984 (2015).
Liepelt, S. & Lipowsky, R. Kinesin’s network of chemomechanical motor cycles. Phys. Rev. Lett. 98, 258102 (2007).
Schreiber, G. Kinetic studies of protein–protein interactions. Curr. Opin. Struct. Biol. 12, 41–47 (2002).
Zhou, J. & Troyanskaya, O. G. Predicting effects of noncoding variants with deep learning-based sequence model. Nat. Methods 12, 931–934 (2015).
Fudenberg, G., Kelley, D. R. & Pollard, K. S. Predicting 3D genome folding from DNA sequence with Akita. Nat. Methods 17, 1111–1117 (2020).
Li, V. R., Zhang, Z. & Troyanskaya, O. G. CROTON: an automated and variant-aware deep learning framework for predicting CRISPR/Cas9 editing outcomes. Bioinformatics 37, 342 (2021).
Avsec, Ž. et al. Effective gene expression prediction from sequence by integrating long-range interactions. Nat. Methods 18, 1196–1203 (2021).
Wong, A. K., Sealfon, R. S., Theesfeld, C. L. & Troyanskaya, O. G. Decoding disease: from genomes to networks to phenotypes. Nat. Rev. Genet. 22, 774–790 (2021).
Ching, T. et al. Opportunities and obstacles for deep learning in biology and medicine. J. R. Soc. Interface 15, 20170387 (2018).
Tareen, A. & Kinney, J. B. Biophysical models of cis-regulation as interpretable neural networks. Preprint at bioRxiv https://doi.org/10.1101/835942 (2019).
Tareen, A. et al. MAVE-NN: learning genotype–phenotype maps from multiplex assays of variant effect. Genome Biol. 23, 1–27 (2022).
Fowler, D. M. & Fields, S. Deep mutational scanning: a new style of protein science. Nat. Methods 11, 801–807 (2014).
Faure, A. J. et al. Mapping the energetic and allosteric landscapes of protein binding domains. Nature 604, 175–183 (2022).
Kretz, C. A. et al. Massively parallel enzyme kinetics reveals the substrate recognition landscape of the metalloprotease ADAMTS13. Proc. Natl Acad. Sci. USA 112, 9328–33 (2015).
Jones, S. K. et al. Massively parallel kinetic profiling of natural and engineered CRISPR nucleases. Nat. Biotechnol. 39, 84–93 (2021).
Zhang, Z., Park, C. Y., Theesfeld, C. L. & Troyanskaya, O. G. An automated framework for efficiently designing deep convolutional neural networks in genomics. Nat. Mach. Intell. 3, 392–400 (2021).
Tsai, S. Q. et al. Circle-seq: a highly sensitive in vitro screen for genome-wide CRISPR–Cas9 nuclease off-targets. Nat. Methods 14, 607–614 (2017).
Haapaniemi, E., Botla, S., Persson, J., Schmierer, B. & Taipale, J. CRISPR–Cas9 genome editing induces a p53-mediated DNA damage response. Nat. Med. 24, 927–930 (2018).
Cancellieri, S. et al. Human genetic diversity alters off-target outcomes of therapeutic gene editing. Nat. Genet. 55, 34–43 (2023).
Eslami-Mossallam, B. et al. A kinetic model predicts SpCas9 activity, improves off-target classification, and reveals the physical basis of targeting fidelity. Nat. Commun. 13, 1–10 (2022).
Klein, M., Eslami-Mossallam, B., Arroyo, D. G. & Depken, M. Hybridization kinetics explains CRISPR-Cas off-targeting rules. Cell Rep. 22, 1413–1423 (2018).
Fu, R. et al. Systematic decomposition of sequence determinants governing CRISPR/Cas9 specificity. Nat. Commun. https://doi.org/10.1038/s41467-022-28028-x (2022).
Concordet, J.-P. & Haeussler, M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 46, 242–245 (2018).
Lin, J., Zhang, Z., Zhang, S., Chen, J. & Wong, K.-C. CRISPR-Net: a recurrent convolutional network quantifies CRISPR off-target activities with mismatches and indels. Adv. Sci. 7, 1903562 (2020).
Listgarten, J. et al. Prediction of off-target activities for the end-to-end design of CRISPR guide RNAs. Nat. Biomed. Eng. 2, 38–47 (2018).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–cas9. Nat. Biotechnol. 34, 184–191 (2016).
Haeussler, M. et al. Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol. 17, 1–12 (2016).
Cameron, P. et al. Mapping the genomic landscape of CRISPR-Cas9 cleavage. Nat. Methods 14, 600–606 (2017).
Kleinstiver, B. P. et al. Engineered CRISPR–Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Zhuo, C. et al. Spatiotemporal control of CRISPR/Cas9 gene editing. Signal Transduct. Target. Ther. 6, 1–18 (2021).
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J. & Wojna, Z. Rethinking the inception architecture for computer vision. In Proc. IEEE Conference on Computer Vision and Pattern Recognition 2818–2826 (IEEE, 2016).
Kelley, D. R. et al. Sequential regulatory activity prediction across chromosomes with convolutional neural networks. Genome Res. 28, 739–750 (2018).
Lundberg, S. M. & Lee, S.-I. A unified approach to interpreting model predictions. Adv. Neural Inf. Process. Syst. 30, 4765–4774 (2017).
Moreno-Mateos, M. A. et al. CRISPRscan: designing highly efficient sgRNAs for CRISPR–Cas9 targeting in vivo. Nat. Methods 12, 982–988 (2015).
Moreb, E. & Lynch, M. Genome dependent Cas9/gRNA search time underlies sequence dependent gRNA activity. Nat. Commun. 12, 5034 (2021).
Moreb, E. A., Hutmacher, M. & Lynch, M. D. CRISPR–Cas “non-target" sites inhibit on-target cutting rates. CRISPR J. 3, 550–561 (2020).
Shen, Y., Pressman, A., Janzen, E. & Chen, I. A. Kinetic sequencing (k-seq) as a massively parallel assay for ribozyme kinetics: utility and critical parameters. Nucleic Acids Res. 49, 67 (2021).
King, E. L. & Altman, C. A schematic method of deriving the rate laws for enzyme-catalyzed reactions. J. Phys. Chem. 60, 1375–1378 (1956).
Cornish Bowden, A. An automatic method for deriving steady-state rate equations. Biochem. J. 165, 55–59 (1977).
Lam, C. F. & Priest, D. G. Enzyme kinetics: systematic generation of valid King–Altman patterns. Biophys. J. 12, 248–256 (1972).
Pelikan, M. Probabilistic model-building genetic algorithms. In Proc. 13th Annual Conference Companion on Genetic and Evolutionary Computation 913–940 (2011).
Wang, Wei, et al. "Backpropagation-friendly eigendecomposition." Advances in Neural Information Processing Systems 32, (2019).
Bae, S., Park, J. & Kim, J.-S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475 (2014).
Lewandowski, D., Kurowicka, D. & Joe, H. Generating random correlation matrices based on vines and extended onion method. J. Multivar. Anal. 100, 1989–2001 (2009).
Salvatier, J., Wiecki, T. V. & Fonnesbeck, C. Probabilistic programming in Python using PyMC3. PeerJ Comput. Sci. 2, 55 (2016).
Zhang, F. Z. & Lamson, A. R. zj-zhang/Elektrum: frozen publication version. Zenodo https://doi.org/10.5281/zenodo.8044859 (2023).
Liu, Q., He, D. & Xie, L. Prediction of off-target specificity and cell-specific fitness of CRISPR–Cas system using attention boosted deep learning and network-based gene feature. PLoS Comput. Biol. 15, 1007480 (2019).
Peng, Hui., Zheng, Yi., Zhao, Zhixun., Liu, Tao. & Li, Jinyan. Recognition of CRISPR/Cas9 off-target sites through ensemble learning of uneven mismatch distributions. Bioinformatics 34, 757–765 (2018).
Lin, J. & Wong, K.-C. Off-target predictions in CRISPR–Cas9 gene editing using deep learning. Bioinformatics 34, 656–663 (2018).
Alkan, F., Wenzel, A., Anthon, C., Havgaard, J. H. & Gorodkin, J. CRISPR–Cas9 off-targeting assessment with nucleic acid duplex energy parameters. Genome Biol. 19, 1–13 (2018).
Acknowledgements
We thank C. Theesfeld and E. Thiede for useful conversations. O.T. is supported by National Institutes of Health (NIH) grant R01GM071966, and Simons Foundation grant no. 395506.
Author information
Authors and Affiliations
Contributions
Z.Z. and A.R.L. conceived the study, wrote the code and performed the analysis. M.S. and O.T. supervised the study. Z.Z., A.R.L., M.S. and O.T. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Computational Science thanks Xin Gao, Christina Theodoris and Ka-Chun Wong for their contribution to the peer review of this work. Primary Handling Editor: Kaitlin McCardle, in collaboration with the Nature Computational Science team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Tables 1-3.
Source data
Source Data Fig. 2
Statistical source data for Fig. 2.
Source Data Fig. 3
Statistical source data for Fig. 3.
Source Data Fig. 4
Statistical source data for Fig. 4.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Z., Lamson, A.R., Shelley, M. et al. Interpretable neural architecture search and transfer learning for understanding CRISPR–Cas9 off-target enzymatic reactions. Nat Comput Sci 3, 1056–1066 (2023). https://doi.org/10.1038/s43588-023-00569-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43588-023-00569-1
This article is cited by
-
Interpretable model of CRISPR–Cas9 enzymatic reactions
Nature Computational Science (2023)