Featured
-
-
Article
| Open Access Plasma proteome profiling reveals dynamic of cholesterol marker after dual blocker therapy
Plasma proteome profiling reveals dynamic of cholesterol marker after dual blocker therapy
Dual blockade therapy is currently being trialled for multiple tumour types, but efficacy is variable. Here, the authors use longitudinal proteomics profiling of 22 patients to develop a predictive model of therapy response.
- Jiacheng Lyu
- , Lin Bai
- & Chen Ding
-
Article
| Open Access BERNN: Enhancing classification of Liquid Chromatography Mass Spectrometry data with batch effect removal neural networks
BERNN: Enhancing classification of Liquid Chromatography Mass Spectrometry data with batch effect removal neural networks
Liquid Chromatography Mass Spectrometry (LC-MS) is a powerful method for profiling biological samples. Here, the authors have developed a suit of Batch Effect Removal Neural Networks (BERNN) to remove batch effects in large LC-MS experiments to maximize sample classification between conditions.
- Simon J. Pelletier
- , Mickaël Leclercq
- & Arnaud Droit
-
Article
| Open Access Accurately clustering biological sequences in linear time by relatedness sorting
Accurately clustering biological sequences in linear time by relatedness sorting
Accurately clustering biological sequences is an increasingly important task but is challenging for large datasets. This study introduces a new approach called ‘relatedness sorting’ to accurately cluster sequences with linear-time scalability.
- Erik Wright
-
Article
| Open Access Systematic HOIP interactome profiling reveals critical roles of linear ubiquitination in tissue homeostasis
Systematic HOIP interactome profiling reveals critical roles of linear ubiquitination in tissue homeostasis
Authors perform an in vivo mass spectrometry-based interactome analysis of HOIL-1-interacting protein, a key component of linear ubiquitination assembly complex.
- Yesheng Fu
- , Lei Li
- & Lingqiang Zhang
-
Article
| Open Access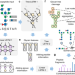 Prediction of glycopeptide fragment mass spectra by deep learning
Prediction of glycopeptide fragment mass spectra by deep learning
Deep learning has achieved a notable success in proteomics and is now emerging in glycoproteomics. Here, the authors develop a neural network-based method to predict mass spectra of intact glycopeptides and demonstrate its potential in data-dependent and data-independent acquisition glycoproteomics.
- Yi Yang
- & Qun Fang
-
Article
| Open Access AlphaPept: a modern and open framework for MS-based proteomics
AlphaPept: a modern and open framework for MS-based proteomics
Mass spectrometry-based proteomics faces the challenge of processing vast data amounts. Here, the authors introduce AlphaPept, an open-source, Python-based framework that offers high speed analysis and easy integration for large-scale proteome analysis.
- Maximilian T. Strauss
- , Isabell Bludau
- & Matthias Mann
-
Comment
| Open Access Fudging the volcano-plot without dredging the data
Fudging the volcano-plot without dredging the data
Selecting omic biomarkers using both their effect size and their differential status significance (i.e., selecting the “volcano-plot outer spray”) has long been equally biologically relevant and statistically troublesome. However, recent proposals are paving the way to resolving this dilemma.
- Thomas Burger
-
Article
| Open Access MARS an improved de novo peptide candidate selection method for non-canonical antigen target discovery in cancer
MARS an improved de novo peptide candidate selection method for non-canonical antigen target discovery in cancer
Detection of neoepitopes from tumours is time consuming and requires the integration of genomic and/or RNA sequencing expression data. Here, the authors propose a machine learning method to enable direct identification of additional, tumour-specific sequences using mass spectrometry through integration of de novo peptide sequencing scores, MHC class I binding prediction, and peptide retention time prediction.
- Hanqing Liao
- , Carolina Barra
- & Nicola Ternette
-
Article
| Open Access Deep learning-driven fragment ion series classification enables highly precise and sensitive de novo peptide sequencing
Deep learning-driven fragment ion series classification enables highly precise and sensitive de novo peptide sequencing
Accurate and high-throughput sequencing methods for proteins are lacking. Here the authors report Spectralis which improves de novo peptide sequencing using a convolutional layer that connects peaks in spectra spaced by amino acid masses, fragment ion series classification and a peptide-spectrum match confidence score.
- Daniela Klaproth-Andrade
- , Johannes Hingerl
- & Julien Gagneur
-
Article
| Open Access Mapping protein states and interactions across the tree of life with co-fractionation mass spectrometry
Mapping protein states and interactions across the tree of life with co-fractionation mass spectrometry
Co-fractionation mass spectrometry (CF-MS) is a powerful technique for mapping protein interactions under physiological conditions. Here, the authors uniformly re-process 411 CF-MS experiments and carry out meta-analyses of protein abundance, protein-protein interactions, and phosphorylation sites in the resulting resource.
- Michael A. Skinnider
- , Mopelola O. Akinlaja
- & Leonard J. Foster
-
Article
| Open Access DeepRTAlign: toward accurate retention time alignment for large cohort mass spectrometry data analysis
DeepRTAlign: toward accurate retention time alignment for large cohort mass spectrometry data analysis
Retention time (RT) alignment is a crucial step in large cohort proteomics and metabolomics studies. Here, the authors introduce DeepRTAlign, a deep learning tool for RT alignment that shows high identification sensitivity and quantitative accuracy.
- Yi Liu
- , Yun Yang
- & Cheng Chang
-
Article
| Open Access Proteomic characterization of epithelial ovarian cancer delineates molecular signatures and therapeutic targets in distinct histological subtypes
Proteomic characterization of epithelial ovarian cancer delineates molecular signatures and therapeutic targets in distinct histological subtypes
The molecular phenotypic features of epithelial ovarian cancer (EOC) remain elusive. Here, the authors perform mass spectrometry-based proteomic profiling for 269 EOC patients and reveal molecularly distinct features and potential therapeutic targets among the histological subtypes of EOC.
- Ting-Ting Gong
- , Shuang Guo
- & Qi-Jun Wu
-
Article
| Open Access Spectroscape enables real-time query and visualization of a spectral archive in proteomics
Spectroscape enables real-time query and visualization of a spectral archive in proteomics
Proteomics data repositories are deluged with data that is scarcely reused. Here, the authors developed Spectroscape, an interactive web-based tool for efficient similarity search of a query spectrum against a repository-scale spectral archive, and real-time visualization of its neighborhood.
- Long Wu
- , Ayman Hoque
- & Henry Lam
-
Article
| Open Access SEPepQuant enhances the detection of possible isoform regulations in shotgun proteomics
SEPepQuant enhances the detection of possible isoform regulations in shotgun proteomics
Protein isoform quantification in shotgun proteomics is challenging due to the mapping of many peptides to multiple protein isoforms. Here, the authors present a computational method SEPepQuant and demonstrate its utility in revealing protein isoform level regulation in shotgun proteomics.
- Yongchao Dou
- , Yuejia Liu
- & Bing Zhang
-
Article
| Open Access Global impact of somatic structural variation on the cancer proteome
Global impact of somatic structural variation on the cancer proteome
The relevance of non-coding somatic mutations in cancer remains elusive. Here, the combination of mass spectrometry-based proteomics and whole genome sequencing data across multiple cancer types helps to assess the effects of somatic structural variant breakpoint patterns on protein expression of nearby genes.
- Fengju Chen
- , Yiqun Zhang
- & Chad J. Creighton
-
Article
| Open Access Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis
Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis
Deep neural networks hold significant promise in capturing the complexity of biological systems. However, they suffer from a lack of interpretability. Here, authors present a generalizable method for developing, interpreting, and visualizing biologically informed neural networks for proteomics data.
- Erik Hartman
- , Aaron M. Scott
- & Johan Malmström
-
Article
| Open Access MSBooster: improving peptide identification rates using deep learning-based features
MSBooster: improving peptide identification rates using deep learning-based features
There is a need for accessible ways to improve peptide spectrum match rescoring with deep learning predictions in bottom-up proteomics. Here, the authors demonstrate robust gains in peptide/protein identifications across various experiments, from single cell proteomics to immunopeptidomics.
- Kevin L. Yang
- , Fengchao Yu
- & Alexey I. Nesvizhskii
-
Article
| Open Access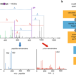 Detecting diagnostic features in MS/MS spectra of post-translationally modified peptides
Detecting diagnostic features in MS/MS spectra of post-translationally modified peptides
Protein modifications increase the complexity of data analysis in mass spectrometry-based proteomics, which may impair the comprehensive mapping of modification sites. Here, the authors develop an algorithm to extract diagnostic fragmentation patterns to improve modified peptide recovery and localization.
- Daniel J. Geiszler
- , Daniel A. Polasky
- & Alexey I. Nesvizhskii
-
Article
| Open Access Analysis of DIA proteomics data using MSFragger-DIA and FragPipe computational platform
Analysis of DIA proteomics data using MSFragger-DIA and FragPipe computational platform
DIA-MS has emerged as a widely used technological platform for quantitative protein profiling. Here, the authors develop MSFragger-DIA, a robust and fast tool to directly identify peptides from DIA spectra. It demonstrates excellent performance across applications from large-scale tumor studies to single-cell proteomics.
- Fengchao Yu
- , Guo Ci Teo
- & Alexey I. Nesvizhskii
-
Article
| Open Access Glycopeptide database search and de novo sequencing with PEAKS GlycanFinder enable highly sensitive glycoproteomics
Glycopeptide database search and de novo sequencing with PEAKS GlycanFinder enable highly sensitive glycoproteomics
Accurate identification of intact glycopeptides from mass spectrometry data is essential for the characterization of glycosylation events in biological samples. Here, the authors propose GlycanFinder, a database search and de novo sequencing tool for the analysis of intact glycopeptides.
- Weiping Sun
- , Qianqiu Zhang
- & Baozhen Shan
-
Article
| Open Access A pharmacoproteomic landscape of organotypic intervention responses in Gram-negative sepsis
A pharmacoproteomic landscape of organotypic intervention responses in Gram-negative sepsis
Sepsis can cause organ damage through disparate immunological and metabolic processes. Here the authors demonstrate a proteomics-based scoring strategy for quantifying quantitative and organotypic changes in relationship to dosing, timing, and potential synergistic intervention combinations during sepsis.
- Tirthankar Mohanty
- , Christofer A. Q. Karlsson
- & Johan Malmström
-
Article
| Open Access Unraveling the glycosylated immunopeptidome with HLA-Glyco
Unraveling the glycosylated immunopeptidome with HLA-Glyco
Protein glycosylation plays a vital role in antigen recognition and immune modulation. Here, the authors present a computational workflow for identifying glycosylated peptides from mass spectrometry-based immunopeptidome data and investigate the properties of glycosylated MHC associated peptides.
- Georges Bedran
- , Daniel A. Polasky
- & Alexey I. Nesvizhskii
-
Article
| Open Access Cell-selective proteomics segregates pancreatic cancer subtypes by extracellular proteins in tumors and circulation
Cell-selective proteomics segregates pancreatic cancer subtypes by extracellular proteins in tumors and circulation
“In-depth cell-selective proteomics and secretomics has remained challenging. Here, the authors devise an optimised azidonorleucine labelling, mass spectrometry method and detect over 10,000 proteins in a pancreatic ductal adenocarcinoma model.
- Jonathan J. Swietlik
- , Stefanie Bärthel
- & Felix Meissner
-
Article
| Open Access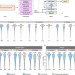 DeepFLR facilitates false localization rate control in phosphoproteomics
DeepFLR facilitates false localization rate control in phosphoproteomics
Protein phosphorylation is a critical modification in many cellular processes. Here, the authors present DeepFLR, a deep learning-based framework to accurately predict phosphopeptide tandem mass spectra and effectively control false localization rates in phosphoproteomics.
- Yu Zong
- , Yuxin Wang
- & Liang Qiao
-
Article
| Open Access PepQuery2 democratizes public MS proteomics data for rapid peptide searching
PepQuery2 democratizes public MS proteomics data for rapid peptide searching
Billions of MS/MS spectra are available in public proteomics data repositories, but their usage has been limited to informatics experts. Here, the authors provide a solution to democratize these data for rapid peptide searching and demonstrate utilities in a wide range of biological applications
- Bo Wen
- & Bing Zhang
-
Article
| Open Access Hierarchical graph learning for protein–protein interaction
Hierarchical graph learning for protein–protein interaction
Despite recent progress, machine learning methods remain inadequate in modeling the natural protein-protein interaction (PPI) hierarchy for PPI prediction. Here, the authors present a double-viewed hierarchical graph learning model, HIGH-PPI, to predict PPIs and extrapolate the molecular details involved.
- Ziqi Gao
- , Chenran Jiang
- & Jia Li
-
Article
| Open Access Integrative proteomic characterization of adenocarcinoma of esophagogastric junction
Integrative proteomic characterization of adenocarcinoma of esophagogastric junction
The molecular subtypes of adenocarcinoma of the esophagogastric junction (AEG) remain to be identified. Here, the authors perform proteogenomic characterisation of AEG tumours with paired normal adjacent tissues and suggest three proteomic subtypes and potential druggable targets.
- Shengli Li
- , Li Yuan
- & Xiang-Dong Cheng
-
Article
| Open Access Parallelized multidimensional analytic framework applied to mammary epithelial cells uncovers regulatory principles in EMT
Parallelized multidimensional analytic framework applied to mammary epithelial cells uncovers regulatory principles in EMT
Epithelial-to-mesenchymal transition (EMT) is a complex process regulated at multiple molecular levels. Here, the authors implement an analytic framework - PAMAF - to integrate data from twelve distinct omics modalities, which they use to understand the molecular changes and regulation during EMT in vitro.
- Indranil Paul
- , Dante Bolzan
- & Andrew Emili
-
Article
| Open Access Sample multiplexing-based targeted pathway proteomics with real-time analytics reveals the impact of genetic variation on protein expression
Sample multiplexing-based targeted pathway proteomics with real-time analytics reveals the impact of genetic variation on protein expression
Targeted proteomics enables robust hypothesis-driven research. Here, Yu et al. present a multiplexed approach for targeted pathway proteomics and apply it to quantify protein families across 480 fully genotyped Diversity Outbred mice, revealing impacts of genetic variation on protein expression and lipid metabolism.
- Qing Yu
- , Xinyue Liu
- & Steven P. Gygi
-
Article
| Open Access Benchmarking commonly used software suites and analysis workflows for DIA proteomics and phosphoproteomics
Benchmarking commonly used software suites and analysis workflows for DIA proteomics and phosphoproteomics
Many software suites and spectral libraries have been developed for DIA proteomics data analysis. Here, the authors create benchmark data sets to evaluate four commonly used software tools combined with seven spectral libraries in both global proteomics and phosphoproteomics analysis.
- Ronghui Lou
- , Ye Cao
- & Wenqing Shui
-
Article
| Open Access Benchmarking tools for detecting longitudinal differential expression in proteomics data allows establishing a robust reproducibility optimization regression approach
Benchmarking tools for detecting longitudinal differential expression in proteomics data allows establishing a robust reproducibility optimization regression approach
Longitudinal proteomics holds great promise for biomarker discovery, but the data interpretation has remained a challenge. Here, the authors evaluate several tools to detect longitudinal differential expression in proteomics data and introduce RolDE, a robust reproducibility optimization approach.
- Tommi Välikangas
- , Tomi Suomi
- & Laura L. Elo
-
Article
| Open Access pGlycoQuant with a deep residual network for quantitative glycoproteomics at intact glycopeptide level
pGlycoQuant with a deep residual network for quantitative glycoproteomics at intact glycopeptide level
Software tools for larger-scale intact glycopeptide quantification lag far behind, which hinders exploring the differential sitespecific glycosylation. Here, the authors report pGlycoQuant, a generic tool with a deep learning model for quantitative glycoproteomics at intact glycopeptide level.
- Siyuan Kong
- , Pengyun Gong
- & Weiqian Cao
-
Article
| Open Access RG/RGG repeats in the C. elegans homologs of Nucleolin and GAR1 contribute to sub-nucleolar phase separation
RG/RGG repeats in the C. elegans homologs of Nucleolin and GAR1 contribute to sub-nucleolar phase separation
Spaulding et al. survey RG/RGG repeats in C. elegans and identify the homologs of Nucleolin (NUCL-1) and GAR1 (GARR-1). RG/RGG repeats are dispensable for nucleolar accumulation but critical for sub-nucleolar phase separation.
- Emily L. Spaulding
- , Alexis M. Feidler
- & Dustin L. Updike
-
Article
| Open Access FLASHIda enables intelligent data acquisition for top–down proteomics to boost proteoform identification counts
FLASHIda enables intelligent data acquisition for top–down proteomics to boost proteoform identification counts
Data acquisition suitable for top-down proteomics (TDP) has the potential to significantly improve proteoform analysis. Here, the authors present FLASHIda, an intelligent online data acquisition algorithm for TDP that nearly doubles the number of proteoform-level identifications in complex samples.
- Kyowon Jeong
- , Maša Babović
- & Oliver Kohlbacher
-
Article
| Open Access A streamlined platform for analyzing tera-scale DDA and DIA mass spectrometry data enables highly sensitive immunopeptidomics
A streamlined platform for analyzing tera-scale DDA and DIA mass spectrometry data enables highly sensitive immunopeptidomics
Immunopeptidomics benefits from highly sensitive mass spectrometry (MS). Here, the authors present a computational platform for integrating data-dependent and -independent acquisition MS approaches, and demonstrate its utility for deeper immunopeptidome profiling.
- Lei Xin
- , Rui Qiao
- & Ming Li
-
Article
| Open Access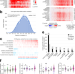 Proteogenomic characterization of 2002 human cancers reveals pan-cancer molecular subtypes and associated pathways
Proteogenomic characterization of 2002 human cancers reveals pan-cancer molecular subtypes and associated pathways
Pan-cancer proteomics analysis enables the analysis of protein expression across multiple cancer types. Here, the authors compare proteomics from 14 cancer types and show 11 distinct subtypes across multiple cancer types. Proteome data could link higher pathway activity levels with somatic alteration of specific genes in the pathway.
- Yiqun Zhang
- , Fengju Chen
- & Chad J. Creighton
-
Article
| Open Access Glyco-Decipher enables glycan database-independent peptide matching and in-depth characterization of site-specific N-glycosylation
Glyco-Decipher enables glycan database-independent peptide matching and in-depth characterization of site-specific N-glycosylation
Poor peptide fragmentation and unusual glycan structures limit mass spectrometry-based analysis of intact N-glycopeptides. Here, the authors develop Glyco-Decipher, a glycan-independent peptide search tool, to tackle these issues and improve the coverage of site-specific glycan analysis.
- Zheng Fang
- , Hongqiang Qin
- & Mingliang Ye
-
Article
| Open Access cyCombine allows for robust integration of single-cell cytometry datasets within and across technologies
cyCombine allows for robust integration of single-cell cytometry datasets within and across technologies
Combining single-cell cytometry datasets increases the analytical flexibility and the statistical power of data analyses. Here, the authors present a method to robustly integrate cytometry data from different batches, experiments, or even different experimental techniques.
- Christina Bligaard Pedersen
- , Søren Helweg Dam
- & Lars Rønn Olsen
-
Article
| Open Access Improved prediction of protein-protein interactions using AlphaFold2
Improved prediction of protein-protein interactions using AlphaFold2
Predicting the structure of protein complexes is extremely difficult. Here, authors apply AlphaFold2 with optimized multiple sequence alignments to model complexes of interacting proteins, enabling prediction of both if and how proteins interact with state-of-art accuracy.
- Patrick Bryant
- , Gabriele Pozzati
- & Arne Elofsson
-
Article
| Open Access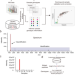 SMAP is a pipeline for sample matching in proteogenomics
SMAP is a pipeline for sample matching in proteogenomics
Sample mix-up is a potential problem in large-scale omic studies due to the complexity of sample processing. Here, the authors present a pipeline for sample matching in proteogenomics to verify sample identity and ensure data integrity.
- Ling Li
- , Mingming Niu
- & Xusheng Wang
-
Article
| Open Access An atlas of protein turnover rates in mouse tissues
An atlas of protein turnover rates in mouse tissues
Protein turnover underpins biology but is challenging to measure in vivo across the entire proteome. Here, the authors provide a comprehensive resource of protein turnover in mouse tissues and develop a visualization platform to analyze these data.
- Zach Rolfs
- , Brian L. Frey
- & Nathan V. Welham
-
Article
| Open Access DeepPhospho accelerates DIA phosphoproteome profiling through in silico library generation
DeepPhospho accelerates DIA phosphoproteome profiling through in silico library generation
The coverage and throughput of data-independent acquisition (DIA)-based phosphoproteomics is limited by its dependence on experimental spectral libraries. Here the authors develop a DIA workflow based on in silico spectral libraries generated by a novel deep neural network to expand phosphoproteome coverage.
- Ronghui Lou
- , Weizhen Liu
- & Wenqing Shui
-
Article
| Open Access An integrative proteomics method identifies a regulator of translation during stem cell maintenance and differentiation
An integrative proteomics method identifies a regulator of translation during stem cell maintenance and differentiation
To characterize molecular changes during cell type transitions, the authors develop a method to simultaneously measure protein expression and thermal stability changes. They apply this approach to study differences between human pluripotent stem cells, their progenies, parental and allogeneic cells.
- Pierre Sabatier
- , Christian M. Beusch
- & Roman A. Zubarev
-
Article
| Open Access GproDIA enables data-independent acquisition glycoproteomics with comprehensive statistical control
GproDIA enables data-independent acquisition glycoproteomics with comprehensive statistical control
Data independent acquisition (DIA) proteomics provides deep coverage and high quantitative accuracy, but is not yet well established in glycoproteomics. Here, the authors develop a DIA-based glycoproteomics workflow with stringent statistical controls to enable accurate glycopeptide identification.
- Yi Yang
- , Guoquan Yan
- & Liang Qiao
-
Perspective
| Open Access A proteomics sample metadata representation for multiomics integration and big data analysis
A proteomics sample metadata representation for multiomics integration and big data analysis
The number of publicly available proteomics datasets is growing rapidly, but a standardized approach for describing the associated metadata is lacking. Here, the authors propose a format and a software pipeline to present and validate metadata, and integrate them into ProteomeXchange repositories.
- Chengxin Dai
- , Anja Füllgrabe
- & Yasset Perez-Riverol
-
Article
| Open Access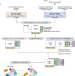 Spatiotemporal proteomic profiling of the pro-inflammatory response to lipopolysaccharide in the THP-1 human leukaemia cell line
Spatiotemporal proteomic profiling of the pro-inflammatory response to lipopolysaccharide in the THP-1 human leukaemia cell line
“Protein relocalisation plays a major role in the innate immune response but remains incompletely characterised. Here, the authors combine temporal proteomics with LOPIT, a spatial proteomic workflow, in a fully Bayesian framework to elucidate spatiotemporal proteomic changes during the LPS-induced immune response in THP-1 cells.
- Claire M. Mulvey
- , Lisa M. Breckels
- & Kathryn S. Lilley
-
Article
| Open Access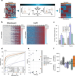 IceR improves proteome coverage and data completeness in global and single-cell proteomics
IceR improves proteome coverage and data completeness in global and single-cell proteomics
Label-free quantitative proteomics by data dependent acquisition offers high protein identification rates but is often limited by missing values. Here, the authors develop a quantification workflow that substantially reduces missing values while maintaining high identification rates and quantification accuracy.
- Mathias Kalxdorf
- , Torsten Müller
- & Jeroen Krijgsveld
-
Article
| Open Access Systematic detection of functional proteoform groups from bottom-up proteomic datasets
Systematic detection of functional proteoform groups from bottom-up proteomic datasets
Many proteins exist in various proteoforms but detecting these variants by bottom-up proteomics remains difficult. Here, the authors present a computational approach based on peptide correlation analysis to identify and characterize proteoforms from bottom-up proteomics data.
- Isabell Bludau
- , Max Frank
- & Ruedi Aebersold
-
Article
| Open Access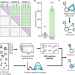 Reliable identification of protein-protein interactions by crosslinking mass spectrometry
Reliable identification of protein-protein interactions by crosslinking mass spectrometry
Cross-linking mass spectrometry (MS) can identify protein-protein interaction (PPI) networks but assessing the reliability of these data remains challenging. To address this issue, the authors develop and validate a method to determine the false-discovery rate of PPIs identified by cross-linking MS.
- Swantje Lenz
- , Ludwig R. Sinn
- & Juri Rappsilber

 Optimizing differential expression analysis for proteomics data via high-performing rules and ensemble inference
Optimizing differential expression analysis for proteomics data via high-performing rules and ensemble inference