Abstract
The first report of transmissible carbapenem resistance encoded by blaIMP-1 was discovered in Pseudomonas aeruginosa GN17203 in 1988, and blaIMP-1 has since been detected in other bacteria, including Enterobacterales. Currently, many variants of blaIMPs exist, and point mutations in the blaIMP promoter have been shown to alter promoter strength. For example, the promoter (Pc) of blaIMP-1, first reported in P. aeruginosa GN17203, was a weak promoter (PcW) with low-level expression intensity. This study investigates whether point mutations in the promoter region have helped to create strong promoters under antimicrobial selection pressure. Using bioinformatic approaches, we retrieved 115 blaIMPs from 14,529 genome data of Pseudomonadota and performed multiple alignment analyses. The results of promoter analysis of the 115 retrieved blaIMPs showed that most of them used the Pc located in class 1 integrons (n = 112, 97.4%). The promoter analysis by year revealed that the blaIMP population with the strong promoter, PcS, was transient. In contrast, the PcW-TG population, which had acquired a TGn-extended −10 motif in PcW and had an intermediate promoter strength, gradually spread throughout the world. An inverse correlation between Pc promoter strength and Intl1 integrase excision efficiency has been reported previously [1]. Because of this trade-off, it is unlikely that blaIMPs with strong promoters will increase rapidly, but the possibility that promoter strength will increase with the use of other integrons cannot be ruled out. Monitoring of the blaIMP genes, including promoter analysis, is necessary for global surveillance of carbapenem-resistant bacteria.
Similar content being viewed by others
Introduction
The first report of transmissible carbapenem resistance was made in 1988 in Pseudomonas aeruginosa GN17203 isolated from a hospital in Toyama Prefecture, Japan [2]. Three years later, Osano et al. reported a novel metallo-beta-lactamase (IMP-1), which is encoded by blaIMP-1, from Serratia marcescens Tn9106 isolated from a patient with a urinary tract infection at a hospital in Aichi Prefecture, Japan in 1991 [3]. Subsequent analysis revealed that P. aeruginosa GN17203 harbored the same blaIMP-1 [4]. Furthermore, Iyobe et al. found that the blaIMP-1 genes of P. aeruginosa GN17203 and S. marcescens Tn9106 were located on the gene cassette of a class 1 integron [5]. On the other hand, blaIMP-1 of S. marcescens isolated from another Aichi Prefecture hospital was transmitted by a novel integron [6], which was later shown to be a class 3 integron [7]. In 2016, blaIMP-27 was reported from Proteus mirabilis on the gene cassette of a class 2 integron [8]. Thus, blaIMPs have been reported to be transmitted by the class 1, 2 and 3 integrons. Here we report that class 1 integrons are primarily involved in the blaIMP propagation.
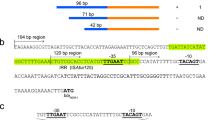
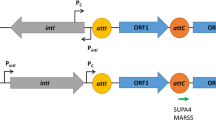
Class 1 integrons consist of an integron integrase gene (intI1) encoding tyrosine recombinase (IntI1), an integron recombination site (attI), and a cassette array consisting of 1–200 contiguous gene cassettes [9, 10]. Intl1 catalyzes the insertion and excision of the gene cassette. All gene cassettes have a cassette recombination site (attC) and generally consist of a single open reading frame (ORF). Multi-resistant integrons (MRIs), which contain multiple gene cassettes encoding drug resistance, have been implicated in multidrug resistance in bacteria [11]. Gene cassettes are generally promoter-less, and their genes are transcribed from the promoter Pc within the coding sequence of Intl1 (Fig. 1a), with transcription levels dependent on polymorphisms in the promoter sequence [12,13,14,15,16,17,18]. Specifically, several Pc variants are derived by point mutations in the −35 and −10 hexamer sequences. Among them, the four major Pc variants are classified as PcW (weak promoter), PcS (strong promoter), PcH1 (intermediate promoter), and PcH2 (intermediate promoter), according to promoter strength. PcH1 and PcH2 are hybrid-type promoters consisting of −35 and −10 hexamers of PcW and PcS in opposite combinations. In addition to mutations in the −10 and −35 hexamers, Nešvera et al. reported a C-to-G mutation 2 bp upstream of the −10 hexamer of PcW, which forms a TGn-extended −10 motif, in a streptomycin/spectinomycin resistance gene, aadA2a [19], and this mutation increased transcription efficiency [19,20,21,22]. Jové et al. investigated the promoter expression intensities of four major combinations of Pc and Pc with TGn-extended −10 motifs using a lacZ reporter system [1]; their findings are summarized in Fig. 1d. In addition, in some cases there is a second promoter, P2, downstream of Pc, with the spacing between the −35 and −10 hexamers optimized to 17 bp by insertion of three G residues [23], and the sequence of P2 is shown in the bottom row of Fig. 1c. Although the diversity of Pc variants and promoter intensities has been analyzed in detail as mentioned above, how such diversity is involved in the spread of IMP-producing bacteria has not been comprehensively analyzed.
Characterization of the blaIMP promoters derived from class 1 (a–d) and class 2 (e) integrons. a Distribution of blaIMPs transcribed by Pc on the gene cassette of a class 1 integron. aac(6’), aminoglycoside 6’-N-acetyltransferase gene; fosI, fosfomycin resistance gene; qacG, quaternary ammonium compound resistance gene; and aac(6’)-Ib3, aminoglycoside N-acetyltransferase gene. b Nucleotide sequences of promoter Pc. One point mutation is observed in each sequence of the −35 hexamer, −10 hexamer, and 17 bp spacer. c Classification of promoter Pc. Pcs were retrieved from the complete level (Comp.) genomic data (n = 112) and from the scaffold level (Scaff.) genomic data (n = 89) and the right column indicates the number of promoters detected. d Color gradient representing the expression intensity of Pcs obtained by the β-galactosidase assay [1]. e Promoter located on a class 2 integron
Here, we retrieved 115 blaIMP genes from 14,529 complete-level genome data of Pseudomonadota deposited in the GenBank database and performed promoter analysis. We found that the blaIMP population transcribed by the strong promoter PcS is transient and regionally localized, whereas the blaIMP population with the intermediate promoter PcW-TG has increased since the late 2000s. Our findings clearly demonstrate that IMP-producing bacteria with intermediate promoter strength of the blaIMP are gradually spreading throughout the world.
Materials and methods
Collection of the bla IMP family from the GenBank database
The blaIMP family was retrieved from 14,529 complete-level genome data (including chromosomes and plasmids) from Pseudomonadota (synonym Proteobacteria) deposited in the GenBank database using Biopython [24] annotated to metallo-beta-lactamase IMP-X on the annotations already made (data retrieval was done on October 7, 2022). As a result, 115 blaIMP genes were retrieved and their upstream regions were used for promoter analysis. Additional blaIMP4 searches were performed using BLAST on 36,214 scaffold-level genome data from Pseudomonadota in the GenBank database. This search yielded 200 genes for blaIMP4 and its variants (showing >90% homology and 100% query coverage with the IMP-4 protein) and their upstream 290 bp region containing the intl1-blaIMP gene order was used for further promoter analysis. This additional search was completed on February 28, 2023.
Promoter analysis of bla IMP and phylogenetic analysis of the bla IMP coding region
The promoter region of blaIMPs was subjected to multiple alignment analysis using MAFFT (https://mafft.cbrc.jp/alignment/server/) to detect point mutations in the −35 and −10 hexamers, and in the 17 bp spacer sequence. For illustration of the phylogenetic tree, the coding region of blaIMP was analyzed by MAFFT and the resulting data were rendered by Phylo.io [25] using the MAFFT sequence alignment server [26]. For the Pc distribution, the world map was rendered by using the rworldmap [27] package in R.
Results
Characterization of the bla NDM-1 promoters
To investigate the blaIMP promoters, we retrieved promoter and coding regions of blaIMPs from 14,529 full-length Pseudomonadota genome data deposited in the GenBank database. A total of 115 blaIMPs were retrieved; they were located in plasmids (91 genes) and chromosomes (24 genes) (Fig. 2a). blaIMP was first detected in P. aeruginosa GN17203, but it is now evident that it has spread to various strains, including Enterobacterales (Fig. 2a). In this study, IMP-4 was the most abundant IMP-type, followed by IMP-1 (Fig. 2b). Promoter analysis of the 115 retrieved blaIMPs showed that they were broadly classified into two groups: those derived from class 1 integrons (n = 112, 97.4%) and those from class 2 integrons (n = 3, 2.6%).
Analysis of promoter Pc within the class 1 integrons
In the 115 blaIMP analyses, 112 Pcs derived from the class 1 integrons were detected. 95 of the 112 blaIMPs belonged to the intl1-blaIMP gene order, while the other 17 genes had other resistance genes arranged between the intl1 and blaIMP in a cassette-like arrangement (Fig. 1a). Multiple sequence alignment analysis using MAFFT (See the Materials and Methods) demonstrated that one point mutation was detected in each of the −35 and −10 hexamers of Pc, which were in coding sequence of the Intl1 (Fig. 1a). For example, among the 112 Pc promoters retrieved, 86 of the −35 hexamers utilized TGGACA and 26 utilized TTGACA. Similarly, there was a frequency bias in the −10 hexamer, with 84 sequences utilizing TAAGCT and 28 sequences utilizing TAAACT (Fig. 1b). Thus, the combination of −35 and −10 hexamers generates PcW, PcH1, PcH2, and PcS as described in the Introduction (Fig. 1c). For the 17 bp spacer sequence between −35 and −10 hexamers, a C-to-G point mutation from two bases upstream of the −10 hexamer was observed and formed the typical TGn-extended −10 motif [22]. For convenience, the PcX promoter with a TGn-extended −10 motif is denoted here as PcX-TG. Including the TGn-mutation, eight promoters could logically be expected to be generated. In this study, we have detected seven Pc derivatives, not including PcH2 (Fig. 1c). In contrast, the second promoter downstream from the Pc region, i.e., P2 [23], was not detected. Thus, the multiple alignment analysis revealed a bias in the Pc region of blaIMP. Furthermore, the reason for limiting point mutations to three sites is that the Pc is located on the coding sequence of Intl1, and mutations that destroy the Intl1 coding region are not allowed.
Yearly transition of the bla IMP Pc promoters
In 1988, IMP-1-producing P. aeruginosa GN17203 was discovered for the first time, and it was shown that blaIMP-1 is transcribed by the weak promoter, PcW [5]. This leads us to ask, Has the blaIMPs family shifted to blaIMP with a strong promoter under the selection pressure of antimicrobial agents? To answer this question, we chronologically sorted Pc promoters (n = 109) from IMP-producing bacteria with known isolation years as shown in Fig. 2, and categorized their promoter types (Fig. 3). As a result, we found that PcW-TG with intermediate promoter strength began to appear around 2007, and promoters with PcW-TG accounted for 47% (n = 51) of the IMP-producing bacteria (n = 109). The blaIMPs utilizing promoters PcH1 and PcH2-TG were detected sporadically and were a minor population. Interestingly, blaIMPs with the strong promoter, PcS, were only detected between 2012 and 2017, and were restricted to Taiwan [28, 29], China [30, 31], and Australia [32] (Table 1). Thus, the emergence of IMP-producing bacteria with blaIMPs with the strong promoter was transient, but blaIMPs transcribed PcW-TG with the intermediate promoter were found to be dominant.
Promoter analysis within the class 2 integrons
In 115 blaIMP analyses, we detected three promoters (herein referred to as Pintl2 for convenience) on the gene cassette of the class 2 integrons as depicted in Fig. 1e. Previously, blaIMP-27 utilizing Pintl2 was detected in IMP-27-producing Proteus mirabilis isolated from two patients in the Upper Plains region of the United States in 2009 and 2015, respectively [8]. In the present analysis, blaIMP-27 utilizing Pintl2 was detected in Morganella morganii and P. mirabilis isolated in Winnipeg, Canada [33]. In addition, blaIMP-64 utilizing Pintl2 was detected in Providencia rettgeri in Quebec, Canada (Table 2). IMP-64 is a variant of IMP-27 with one amino acid substitution. Collectively, blaIMPs with Pintl2 are involved in the spread of IMP-27 and its variant, IMP-64, and these are geographically restricted.
The promoter type of bla IMP-1 and bla IMP-4 is shifting to PcW-TG
Our results suggest that Pc is being shifted to PcW-TG in the blaIMP family (Fig. 3). To further validate the utilization frequency of Pc, we retrieved the blaIMP-4 and its variants (>90% homology with the blaIMP-1 product), which are mostly represented in Fig. 2, from 36,214 scaffold-level genome data from Pseudomonadota deposited in the GenBank database. As a result, 89 blaIMPs, which belonged to the intl1-blaIMP gene order, encoding IMP-4 and its one amino acid variant, IMP-38, and IMP-1 and its one amino acid variant, IMP-34, were retrieved. These data were collected after 2007, and by using these data, we can speculate how the Pc of blaIMPs has recently shifted and dominated. Consequently, the Pcs of blaIMP-1 and blaIMP-4 collected after 2007 are also dominated by PcW-TGs (n = 52, 58.4%) (Fig. 4). Furthermore, we also confirmed that blaIMPs with a strong promoter, PcS and PcS-TG, were restricted.
Geographic spread of IMP-1- and IMP-4-producing bacteria and biased promoter utilization
Promoter analysis of blaIMP-1 and blaIMP-4 demonstrated that blaIMP with PcW-TG is currently spreading (Fig. 4). On the other hand, the correlation between promoter types and countries has not been fully elucidated. Therefore, we retrieved the Pcs of blaIMP-1 (n = 74) and blaIMP-4 (n = 73) from the full-length and scaffold-level genomic data and examined their spread and promoter types by country (Fig. 5). Our data demonstrated that blaIMP-1 with PcW-TG has been detected mainly in Asian countries, although blaIMP-1 genes were also detected in the UK, Brazil, and multiple African countries (Fig. 5b). Thus, the emergence of IMP-1-type-producing bacteria with different Pc promoters, which are geographically distant from Japan, appears to be sporadically distributed, and we further investigated this using a phylogenetic tree analysis (Fig. 6). On the other hand, detection of blaIMP-4 genes was limited to China and Australia, and again blaIMP-4 with PcW-TG were dominant (Fig. 5c).
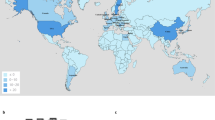
Geographic distribution of blaIMP-1 and blaIMP-4 (a) and promoter diversity (b). Pc promoters of blaIMP-1 (n = 74) and blaIMP-4 (n = 73) from the full-length and scaffold-level genomic data were collected, and their spread and promoter types were investigated by country. Red and blue circles indicate the number of blaIMP-1 and blaIMP-4 detected, respectively. The world map was rendered by rworldmap as described in the Materials and Methods
The amino acid sequences translated from blaIMP-1 verified above are identical and form the IMP-1 family, but the coding sequence may contain several silent mutations. Therefore, coding sequences of blaIMP-1 and blaIMP-4 used in this analysis (Fig. 5) were subjected to multiple alignment analysis and the resulting data were used for phylogenetic tree analysis as described in the Materials and Methods. Several silent mutations were observed in the blaIMP-1 coding region, resulting in the phylogenetic tree shown in Fig. 6. Of the blaIMP-1 coding regions (n = 74) used in the analysis, 74.7% (n = 59) shared the same coding sequence, grouped in cluster A, while the remaining 15 genes were grouped into four clusters B ~ E (Fig. 6). As described elsewhere for blaIMP-1, it was concluded that IMP-1-producing bacteria from Asia, including Japan and China, grouped in cluster A, and IMP-1-producing bacteria from Europe are geographically independent [34]. In particular, seven strains isolated in the UK, grouped in cluster D, including blaIMP-1 of Enterobacter complex and blaIMP-1 of Klebsiella aerogenes [35] had the same silent mutation and the same promoter, PcH2-TG, suggesting that they were not derived from Asia but sporadically distributed in the UK (Table 3). Similarly, three strains from the IMP-1-producing P. aeruginosa found in Africa, grouped in cluster E, had silent mutations in their coding sequences. In contrast, the coding sequences of the four blaIMP-1 with PcS isolated in Taiwan and the one blaIMP-1 with PcS-TG isolated in Brazil (see Fig. 5b) were the same as that of the blaIMP-1 isolated in Japan as grouped in cluster A. Interestingly, no silent mutations were observed in the coding sequence of blaIMP-4 that spread to China and Australia. Between 1994 and 1998, imipenem-resistant Acinetobacter strains were detected in Hong Kong, and this is the first report of IMP-producing bacteria with blaIMP-1 [36]. Taken together, these results suggest that the blaIMP-1 detected in the UK and continental Africa may not have been of Asian origin and was most likely derived from a geographically isolated source and that IMP-4-producing bacteria spread from Hong Kong and mainland China to Australia.
Discussion
Promoter analysis was performed on 115 blaIMP genes retrieved from 14,529 complete genomic data of Pseudomonadota. The results showed that the blaIMPs were divided into those derived from class 1 integrons (n = 112, 97.4%) and those from class 2 integrons (n = 3, 2.6%), respectively, and no promoters from class 3 integrons were detected. The Pc promoter is characterized by the presence of −35 and −10 hexamers within the coding sequence of Intl1; therefore, promoter mutations disrupting the translation of the coding sequence are not acceptable. The purpose of this study was to analyze alterations in and the spread of the blaIMP promoter under the Intl1 functional constraints.
It is known that there is an inverse correlation between the strength of Pc and the integrase efficiency for the integration and excision of gene cassettes [1], and in the present study, we demonstrated that the blaIMP with PcW-TG, the promoter of intermediate strength, diffuses in this trade-off relationship. Multiple alignment analysis revealed no silent mutations in the blaIMP-4 coding sequence, but polymorphisms were observed in the Pc promoter. Thus, point mutations in the Pc region occur independently of mutations in the blaIMP coding sequence. Interestingly, IMP-4-producing E. coli 1585m1 has been isolated from the silver gull [32] (Table 1). This strain harbors two plasmids and each plasmid contains blaIMP-4 with the strong promoter PcS. Thus, silver gulls living near urban areas may be potential reservoirs of carbapenem resistance.
Due to the fact that most studies are related to IMP typing or enzyme functions of metallo-beta-lactamase, and these data do not cover upstream promoter regions, there is a paucity of information on blaIMP promoter regions. In addition, our analysis method is based on the retrieval of blaIMP according to known annotations using Biopython. For example, the first report of a blaIMP was from P. aeruginosa strain GN17203 isolated in 1988 [2], but we were unable to search for the blaIMP because it was not annotated at that time. We expect that more accurate genomic information will be obtained by alteration of the retrieval conditions of the blaIMP, which is an issue for the future.
In this study, we modeled the spread of blaIMP with strong promoters as a means for resistant bacteria to survive in the environment, but in reality, we found that blaIMPs with promoters of intermediate strength were widely distributed. In the future, a more detailed analysis of blaIMP diffusion will be possible by including the plasmid replicon type.
References
Jové T, Re SD, Denis F, Mazel D, Ploy M-C. Inverse correlation between promoter strength and excision activity in class 1 integrons. PLoS Genet. 2010;6:e1000793.
Watanabe M, Iyobe S, Inoue M, Mitsuhashi S. Transferable imipenem resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1991;35:147–51.
Osano E, et al. Molecular characterization of an enterobacterial metallo β-lactamase found in a clinical isolate of Serratia marcescens that shows imipenem resistance. Antimicrob Agents Chemother. 1994;38:71–8.
Minami S, et al. Imipenem and cephem resistant Pseudomonas aeruginosa carrying plasmids coding for class B β-lactamase. J Antimicrob Chemother. 1996;37:433–44.
Iyobe S, Yamada H, Minami S. Insertion of a carbapenemase gene cassette into an integron of a Pseudomonas aeruginosa plasmid. J. Antimicrob Chemother. 1996;38:1114–5.
Arakawa Y, et al. A novel integron-like element carrying the metallo-β-lactamase gene blaIMP. Antimicrob Agents Chemother. 1995;39:1612–5.
Collis CM, Kim M-J, Partridge SR, Stokes HW, Hall RM. Characterization of the class 3 integron and the site-specific recombination system it determines. J Bacteriol. 2002;184:3017–26.
Dixon N, et al. IMP-27, a unique metallo-β-lactamase identified in geographically distinct isolates of Proteus mirabilis. Antimicrob Agents Chemother. 2016;60:6418–21.
Stokes HW, Hall RM. A novel family of potentially mobile DNA elements encoding site‐specific gene‐integration functions: integrons. Mol Microbiol. 1989;3:1669–83.
Ghaly TM, et al. The natural history of integrons. Microorganisms. 2021;9:2212.
Partridge SR, Tsafnat G, Coiera E, Iredell JR. Gene cassettes and cassette arrays in mobile resistance integrons. FEMS Microbiol Rev. 2009;33:757–84.
Lévesque C, Brassard S, Lapointe J, Roy PH. Diversity and relative strength of tandem promoters for the antibiotic-resistance genes of several integron. Gene. 1994;142:49–54.
Bunny KL, Hall RM, Stokes HW. New mobile gene cassettes containing an aminoglycoside resistance gene, aacA7, and a chloramphenicol resistance gene, catB3, in an integron in pBWH301. Antimicrob Agents Chemother. 1995;39:686–93.
Papagiannitsis CC, Tzouvelekis LS, Miriagou V. Relative strengths of the class 1 integron promoter hybrid 2 and the combinations of strong and hybrid 1 with an active P2 promoter. Antimicrob. Agents Chemother. 2009;53:277–80.
Yamamoto M, et al. Molecular analysis of a blaIMP-1-harboring class 3 integron in multidrug-resistant Pseudomonas fulva. Antimicrob Agents Chemother. 2018;62:e00701–18.
Wang T, Zhu Y, Zhu W, Cao M, Wei Q. Molecular characterization of class 1 integrons in carbapenem-resistant Enterobacterales isolates. Microb Pathog. 2023;177:106051.
Wei Q, et al. Diversity of gene cassette promoter variants of class 1 integrons in uropathogenic Escherichia coli. Curr Microbiol. 2013;67:543–9.
Vinué L, Jové T, Torres C, Ploy M-C. Diversity of class 1 integron gene cassette Pc promoter variants in clinical Escherichia coli strains and description of a new P2 promoter variant. Int J Antimicrob Agents. 2011;38:526–9.
Nešvera J, Hochmannová J, Pátek M. An integron of class 1 is present on the plasmid pCG4 from Gram‐positive bacterium Corynebacterium glutamicum. FEM Microbiol Lett. 1998;169:391–5.
Kumar A, et al. The minus 35-recognition region of Escherichia coli sigma 70 is inessential for initiation of transcription at an “extended minus 10” promoter. J Mol Biol. 1993;232:406–18.
Barne KA, Bown JA, Busby SJ, Minchin SD. Region 2.5 of the Escherichia coli RNA polymerase σ70 subunit is responsible for the recognition of the “extended-10” motif at promoters. EMBO J. 1997;16:4034–40.
Burr T, Mitchell J, Kolb A, Minchin S, Busby S. DNA sequence elements located immediately upstream of the –10 hexamer in Escherichia coli promoters: a systematic study. Nucleic Acids Res. 2000;28:1864–70.
Wei Q, et al. Transcription of integron‐harboured gene cassette impacts integration efficiency in class 1 integron. Mol Microbiol. 2011;80:1326–36.
Chapman B, Chang J. Biopython: Python tools for computational biology. ACM SIGBIO Newsl. 2000;20:15–19.
Robinson O, Dylus D, Dessimoz C. Phylo.io: Interactive viewing and comparison of large phylogenetic trees on the web. Mol Bio Evol. 2016;33:2163–6.
Kuraku S, Zmasek CM, Nishimura O, Katoh K. aLeaves facilitates on-demand exploration of metazoan gene family trees on MAFFT sequence alignment server with enhanced interactivity. Nucleic Acids Res. 2013;41:W22–W28.
South A. rworldmap: a new R package for mapping global data. R J. 2011;3:35.
Chen F-J, et al. Molecular epidemiology of emerging carbapenem resistance in Acinetobacter nosocomialis and Acinetobacter pittii in Taiwan, 2010 to 2014. Antimicrob Agents Chemother. 2019;63:e02007–18.
Li L-H, et al. Clinical and molecular characterization of Acinetobacter seifertii in Taiwan. J. Antimicrob Chemother. 2020;76:312–21.
Chen F, et al. Uncovering the hidden threat: The widespread presence of chromosome-borne accessory genetic elements and novel antibiotic resistance genetic environments in Aeromonas. Virulence. 2023;14:2271688.
Yuan S, Wu G, Zheng B. Complete genome sequence of an IMP-8, CTX-M-14, CTX-M-3 and QnrS1 co-producing Enterobacter asburiae isolate from a patient with wound infection. J Glob Antimicrob Resist. 2019;18:52–54.
Wyrsch ER, et al. Urban Wildlife Crisis: Australian silver gull is a bystander host to widespread clinical antibiotic resistance. mSystems. 2022;7:e00158–22.
Boyd DA, et al. Emergence of Morganellaceae harboring blaIMP-27 metalloenzyme in Canada. mSphere. 2021;6:e00048–21.
Walsh TR, Toleman MA, Poirel L, Nordmann P. Metallo-β-Lactamases: the quiet before the storm? Clin Microbiol Rev. 2005;18:306–25.
Turton JF, et al. IncN3 and IncHI2 plasmids with an In1763 integron carrying blaIMP-1 in carbapenem-resistant Enterobacterales clinical isolates from the UK. J Méd Microbiol. 2020;69:739–47.
Chu Y-W, et al. IMP-4, a novel metallo-β-lactamase from nosocomial Acinetobacter spp. Collected in Hong Kong between 1994 and 1998. Antimicrob Agents Chemother. 2001;45:710–4.
Acknowledgements
This work was supported in part by grants from the Ministry of Education, Culture, Sports, Science and Technology, KAKENHI grants from the Japan Society for the Promotion of Science (nos. 19K07542 to A.A. and 20K07485, 23K06531 to A.K.) and AMED grants (nos. JP20nk0101552 to A.A. and JP22nk0101587 to A.A.). The funders had no role in the study design, data collection or analysis, the decision to publish, or the preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kikuchi, Y., Yoshida, M., Kuwae, A. et al. Correlation between the spread of IMP-producing bacteria and the promoter strength of blaIMP genes. J Antibiot 77, 315–323 (2024). https://doi.org/10.1038/s41429-024-00715-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-024-00715-5