Abstract
Mitochondrial biology and behavior are central to the physiology of liver. Multiple mitochondrial quality control mechanisms remodel mitochondrial homeostasis under physiological and pathological conditions. Mitochondrial dysfunction and damage induced by overnutrition lead to oxidative stress, inflammation, liver cell death, and collagen production, which advance hepatic steatosis to nonalcoholic steatohepatitis (NASH). Accumulating evidence suggests that specific interventions that target mitochondrial homeostasis, including energy metabolism, antioxidant effects, and mitochondrial quality control, have emerged as promising strategies for NASH treatment. However, clinical translation of these findings is challenging due to the complex and unclear mechanisms of mitochondrial homeostasis in the pathophysiology of NASH.
Similar content being viewed by others
Introduction
Due to caloric excess and sedentary lifestyles, obesity and metabolic syndrome have become global epidemics. Nonalcoholic fatty liver disease (NAFLD) is defined as ectopic lipid accumulation in the liver in the absence of excessive alcohol intake or other attributable causes. As a hepatic consequence of metabolic syndrome and obesity, NAFLD is estimated to affect up to 25% of the adult population worldwide, and it may progress to nonalcoholic steatohepatitis (NASH) in approximately 20% of patients, which may lead to cirrhosis or hepatocellular carcinoma (HCC) [1]. Notably, China experienced an unexpected rapid increase in NAFLD, with a prevalence of 29.2% and the largest number of patients, over a short period from 2008 to 2018 [2]. Given the prevalence and burden, rising awareness and urgent actions are required to control the NAFLD pandemic. Novel therapeutic targets and a greater understanding of the pathophysiology of NAFLD are also urgently needed for improved treatment [3].
NAFLD is a spectrum of chronic liver diseases varying from isolated excessive hepatic triglyceride accumulation and steatosis (nonalcoholic fatty liver, NAFL), to a more serious process with inflammation and hepatocyte damage (steatohepatitis) [1]. Patients with only NAFL carry a very low risk of adverse outcomes, but the presence of NASH increases the risks progress to cirrhosis, liver failure, and HCC [4]. NAFLD has very different rates of highly variable progression between individuals and different clinical manifestations, which reflects the complex and undefined pathogenesis. The ‘two-hit’ theory suggests that in the setting of steatosis alone (i.e., NAFL), a second ‘hit’ from other factors (e.g., oxidant stress) was required for the development of NASH, which was the original hypothesis model of steatohepatitis pathogenesis 20 years ago [5]. However, recent studies indicated that identical pathogenic drivers in all patients are unlikely. The ‘two-hit’ hypothesis is now considered outdated because it does not explain the several molecular and metabolic changes that occur in NAFLD [6]. The ‘multiple hit’ hypothesis considers simultaneous multiple insults acting on genetically predisposed subjects to induce NAFLD and provides a more accurate explanation of NAFLD pathogenesis. These ‘hits’ include insulin resistance, hormones secreted from adipose tissue, nutritional factors, gut microbiota, genetic and epigenetic factors [7].
Mitochondria are central powerhouses that perform many key functions in the cell, including oxidative phosphorylation, reactive oxygen species (ROS) generation, nutrient metabolism, and intracellular signaling cascades [8]. Mitochondrial homeostasis is maintaining the healthy mitochondrial functions that are responsible for intracellular signaling cascades. Mitochondrial dysfunction underlies the etiology of a broad spectrum of diseases, including neurodegenerative diseases, NAFLD/NASH, and other metabolic diseases [9]. The present review discusses the distinct and diverse mechanisms of mitochondrial dysfunction in the pathology and etiology of hepatic steatosis, such as the adaptation and ‘remodeling’ of mitochondrial energetics, morphology, mitochondrial DNA (mtDNA), oxidative stress, nuclear transcriptional regulation, and autophagy-mediated quality control. We also review current therapeutic approaches in NASH with an emphasis on mitochondria as potential therapeutic targets.
Mitochondrial biology in liver
Mitochondrial function plays an important role in normal physiology and cellular function in the liver [8, 10]. This organelle arose approximately two billion years ago from an ancestral bacterium and contains its own genome (mtDNA). Mammalian mtDNA encodes 13 proteins that are involved in the respiratory chains. All other proteins are encoded by nuclear genes and imported into mitochondria primarily via the translocase of the outer membrane (TOM) and translocase of the inner membrane (TIM) complexes [11, 12].
Mitochondrial metabolism
Mitochondria play important roles in energy production
One of the important functions of mitochondria in the liver is to produce energy via oxidative phosphorylation [OXPHOS] [13, 14]. Energy production, primarily in the form of ATP, is controlled by mitochondria that link oxidative respiration with the metabolism of nutrients, such as carbohydrates, fatty acids, and amino acids. Two major steps, oxidation of NADH or FADH2 and phosphorylation of ADP to form ATP by ATP synthase, are required for oxidative respiration to produce ATP in mitochondria. These two reactions are coupled, and OXPHOS is the most efficient pathway for energy production in the tricarboxylic acid cycle (TCA cycle). Reducing equivalents (NADH or FADH2) are produced in respiratory complex I or complex II, respectively, during the catabolism of carbon substrates into acetyl-CoA from pyruvate (glycolysis) or acyl-CoA (fatty acid β-oxidation), which are then oxidized to NAD or FAD. The protons produced during oxidation are pumped to the inter-mitochondrial membrane via respiratory complexes I, III, and IV. Electrons from NADH or FADH2 are transferred via a series of respiratory chain complexes to O2, which ultimately generates H2O in complex IV. The proton gradient across the inter-membrane and mitochondrial matrix is the driving force of ATP production from ADP by ATP synthase (F1F0-ATPase) [15]. ATP is transported to the cytoplasm via adenine nucleotide translocators (ANTs) by exchange with ADP and is used for various biological processes [16]. Notably, the mitochondrial membrane potential (Δψm) is essential for ATP production, and it sustains many other mitochondrial functions, including ion and metabolite exchanges and the importation of mitochondrial precursor proteins from the cytosol [17].
Mitochondria fully participate in metabolic flexibility
The liver plays an important role in energy homeostasis by regulating diverse carbon metabolism of nutrients in the mitochondria, which are involved in hepatic anabolic pathways (de novo lipogenesis, gluconeogenesis, and folate metabolism) and catabolic pathways (TCA cycle, urea cycle, fatty acid β-oxidation, ketogenesis, amino acid metabolism, and ROS production) [18]. The capacity to use distinct substrates under a wide variety of stimuli enables the continuous work of mitochondria in the liver. Under a fasting state, β-oxidation is initiated from fatty acids, and glycolysis is initiated from glucose, which constitutes the most prominent sources of acetyl-CoA for the TCA cycle [19]. The abundant acetyl-CoA catalyzed from β-oxidation induces ketone formation, which is exported from liver and used by peripheral tissue [20]. During a feeding state, mitochondrial citrate is transported to cytoplasm and leads to cytosolic acetyl-CoA formation catalyzed by ATP citrate lyase (ACLY), which is used for de novo lipogenesis (DNL) or epigenetic histone acetylation [21, 22].
Acetyl-CoA is the beginning of TCA cycle. Fatty acid β-oxidation (FAO) is an efficient metabolic pathway for acetyl-CoA production. Briefly, free fatty acids (FFAs) are converted into acyl-CoA by acyl-CoA synthetase (ACSLs) and metabolized to acetyl-CoA via multiple catalytic enzymes in hepatic mitochondria [23]. Branched-chain amino acids (BCAAs) are also involved in acetyl-CoA production. BCAAs are trans-aminated into branched-chain keto acids (BCKAs) in muscle and adipose tissue, which are shuttled to liver and catabolized to acetyl-CoA in mitochondria [24]. Glutamine-α-ketoglutarate (α-KG) metabolism catalyzed by glutamate dehydrogenase (GDH) and glutamine synthetase (GS) in mitochondria, drives the TCA cycle and ATP production [25].
Mitochondrial metabolism also closely controls histone epigenetic modification. Cytosolic acetyl-CoA derived from mitochondria is transferred into nucleus and participates in histone acetylation via histone acetyl transferases (HATs) in the progression of metabolic stress [22, 26]. One carbon produced from serine links folate, and the methionine cycle contributes to methyl donor production and DNA methylation. During the process of DNA methylation, serine is catalyzed by serine hydroxymethyltransferase 2 (SHMT2) and 5,10-methylene tetrahydrofolate dehydrogenase 2 (MTHFD2) in mitochondria to induce the production of folate [27]. Glutamine metabolism in mitochondria is also closely associated with serine metabolism and epigenetic modification via one carbon and α-ketoglutarate (α-KG) [28].
Mitochondrion is an important source of ROS
Mitochondria are the major sites of ROS formation in the cell and play key roles in maintaining normal energy cell redox homeostasis in multiple life processes [29]. ROS is generated in a physiological range. However, the partial reduction of O2 or mitochondrial proton leakage leads to the production of the primary ROS named superoxide anions (O2•−). H2O2 is generated via the spontaneous or superoxide dismutase (SOD)-catalyzed dismutation of O2•− in mitochondria. O2•− and H2O2 are converted into hydroxyl radicals (OH•) via the Fe2+-mediated Fenton reaction, and it is a highly reactive oxygen species [30]. OH• initiates the formation of lipid (L•) and lipid peroxyl (LOO•) radicals. H2O2 is then converted into H2O via mitochondrial peroxiredoxins, catalase (CAT), and glutathione peroxidases (GPX) [31].
Under physiological conditions, complex I is the main site of mitochondrial ROS production, and complex III is also a site for O2•− production. A recent study indicated that inducing reverse electron transport (RET) in vivo increased mitochondrial ROS and improved mitochondrial function [32]. Another major source of intracellular ROS production is catalyzed by NADPH oxidase (NOX) family proteins, which transfer electrons from NADPH to molecular oxygen [33]. The freely diffusible nitric oxide (NO) formed elsewhere also crosses mitochondrial membranes to react with superoxide and form peroxynitrite, which causes the formation of 3-nitrotyrosine residues on several proteins, including respiratory complex I and V subunits [34]. Normal levels of ROS under physiological conditions act as signaling molecules that play critical roles in most intra- and extracellular processes in the liver. However, intracellular ROS overload induces cellular dysfunction and pathological processes.
Mitochondrial biogenesis
Mitochondrial biogenesis is highly plastic in response to cellular energy demands that are triggered by developmental signals and environmental stimuli. Due to their bacterial origin, mitochondria have their own genome and auto-replicate [35]. Human mitochondrial proteins are encoded by nuclear and mitochondrial genomes, which means that mitochondrial biogenesis requires both the regulations of mitochondrial and nuclear genome. We provide a brief overview of mitochondrial biogenesis regulations, including mtDNA replication, transcription and translation, nuclear factor-mediated transcription regulation, and nuclear posttranscriptional regulation, such as microRNA interference, alternative splicing, and RNA stability (Fig. 1).
Mitochondria arose from an ancestral bacterium and contain their own genome (mtDNA), which encodes 13 proteins involved in the respiratory chains. However, greater than 98% of the total protein complement of the organelle is encoded by the nuclear genome and plays a crucial role in mitochondrial function. An overview of the regulation of mitochondrial and nuclear genomes in mitochondrial gene expression and the signaling pathways is summarized in this figure. MtDNA gene replication, transcription, and translation regulations are all involved in the assembly of OXPHOS complexes, which play an important role in mitochondrial function. The nuclear genomes encoded mitochondrial function-related proteins via transcription (chromatin remodeling, DNA methylation, transcription factors) and posttranscription (miRNA interference, alternative splicing, RNA stability) regulation and control the mitochondrial function, such as OXPHOS, FAO, TCA cycle, mitochondrial biogenesis, mitochondrial fission/fusion, and mitophagy (see text for additional information).
mtDNA replication, transcription, and translation
Mammalian mtDNA is a double-stranded circular molecule that contains 37 genes encoding 13 polypeptides of the OXPHOS system, 22 other tRNAs (transfer RNAs), and two rRNAs (ribosomal RNAs), which are necessary for the translation of respiratory subunit mRNAs within the mitochondrial matrix [11, 36]. Therefore, the expression of mtDNA is vital for the assembly and function of oxidative phosphorylation complexes. Defects in the mechanisms regulating mtDNA gene replication, transcription, and translation are associated with deficiencies in the assembly of these complexes and result in mitochondria-related diseases.
mtDNA replication regulations
Mammalian mitochondria contain multiple copies of the mtDNA genome, and a dedicated DNA replication machinery is required for its maintenance. The specific mechanism of mtDNA replication is not clear, but a strand displacement model was presented in 1972 [37]. According to this model, mtDNA replication is initiated at the H-strand origin of replication (OH) and continues to produce a new H-strand. Mitochondrial ssDNA-binding protein (mtSSB) binds and protects the exposed parental H-strand from mitochondrial RNA polymerase (POLRMT), which initiates random RNA synthesis [38]. When the replisome passes the L-strand origin of replication (OL), the H-strand folds into a stem-loop structure and blocks mtSSB binding. Therefore, a single-stranded loop region remains accessible, which allows POLRMT to initiate primer synthesis [39]. After approximately 25 nt, mitochondrial DNA polymerase γ (POLγ) replaces POLRMT at the 3′-end of the primer and initiates L-strand DNA synthesis. Synthesis of the two strands proceeds in a continuous manner until two full, double-stranded DNA molecules are formed. Another study questioned the strand-displacement model and proposed a RITOLS model in which the processed transcripts are successively hybridized to the paternal H-strand and function as the replication fork advances [40]. Under certain conditions, strand-coupled replication may function as a backup replication mode in mammalian mitochondria [41].
mtDNA transcription regulations
Functional mammalian mitochondrial biogenesis requires the activation of mitochondrial transcription. Mammalian mtDNA transcription originates in the major noncoding region with the L-strand (LSP) and H-strand (HSP) promoters [42]. In the transcription initiation complex, mitochondrial transcription factor A (TFAM), bound to DNA, recruits POLRMT to the promoter via its N-terminal extension, and mitochondrial transcription factor B2 (TFB2M) modifies the structure of POLRMT to induce opening of the promoter [43, 44]. Mitochondrial transcription elongation factor (TEFM) interacts with POLRMT via its C-terminal domain and strongly promotes POLRMT processivity because it stimulates the formation of longer transcripts for the elongation stage around the mtDNA downstream [45]. However, the mechanism of transcription termination is not clear. A previous study suggested that mitochondrial termination factor 1 (MTERF1) induced transcription termination via base flipping and DNA unwinding [46]. However, more recent evidence contradicts this hypothesis [47], and further study is needed.
mtDNA translation regulations
The regulation of mammalian mitochondrial translation is fully dependent on various nuclear-encoded regulatory proteins. The mtDNA-encoded genes in mammalian mitochondria are translated into proteins with the assistance of specific translation factors (encoded by nuclear DNA, nDNA), such as initiation factors 2 and 3 (mtIF2 and mtIF3), elongation factors Tu, Ts, and G1 (mtEFTu, mtEFTs, and mtEFG1), translational release factor-1 (mtRF1) and recycling factors (mtRRF1 and mtRRF2) [48, 49]. Translation is initiated with a methionine residue, but only a single tRNAMet is used for initiation and elongation in mammalian mitochondria. Therefore, a formylation of methionine (fMet) is formed to increase its affinity for mtIF2, which directs the association of fMet-tRNAMet with the mRNA. MtIF3 positions the AUG or AUA initiation codons of the mRNA at the peptidyl (P) site and initiates translation [50]. MtEFTu forms a complex with GTP and aminoacyl tRNA during elongation, which directs the tRNA to the acceptor (A) site and pairs with the mRNA at the codon–anticodon site. GTP hydrolysis and mtEFTu release catalyzes peptide bond formation. MtEFG1 releases the deacetylated tRNA from the P-site and translocates the peptidyl-tRNAs from the A and P sites to the P and exit (E) sites, which causes the mRNA to move along by one codon. The GTP:EFTu complex is re-established by EFTs [51]. The termination of mitochondrial translation is triggered by a stop codon at the A-site, where mtRF1 catalyzes the hydrolysis of peptidyl tRNA and releases the polypeptide [52]. Mitochondrial ribosomal recycling factors, mtRRF, catalyze the release of mRNAs, deacetylated tRNAs and ribosomal subunits [53].
Nuclear transcription regulations
Mammalian mitochondria contain more than 1000 proteins, but their genome encodes only 13 proteins. Most mitochondrial genes are situated in the nucleus, and the transcription complexes at the promoters of these genes control their expression. The following section highlights some of the major nuclear transcriptional complexes, such as nuclear respiratory factors, nuclear hormone receptors, and transcriptional coactivators, that regulate mitochondrial gene expression.
Nuclear respiratory factors
Nuclear respiratory factor 1 (NRF-1) was initially identified as an important regulator of cytochrome c gene expression via promoter sequence analysis. NRF-1 controls the expression of a significant number of the five respiratory complex proteins, and mitochondrial import proteins [54]. NRF-1 also modulates the gene expression of TFAM and transcription factor B proteins (TFBs) and controls the transcriptional and replicative activity of the mitochondrial genome. Another nuclear respiratory factor, nuclear respiratory factor 1 (NRF-2), is involved in the promoter region of cytochrome c oxidase complex subunit IV and regulates the expression of proteins in the electron transport chain [55]. Similar to NRF1, NRF2 also controls the expression of TFAM and TFBs and integrates nuclear control with mitochondrial DNA transcription and replication.
Nuclear hormone receptors
Peroxisome proliferator-activated receptors (PPARs), which belong to the nuclear hormone receptor superfamily, are activated by long-chain fatty acids and eicosanoids and control mitochondrial function and biogenesis [56]. PPARα regulates the constitutive transcription of genes encoding fatty acid-metabolizing enzymes and mitochondrial β-oxidation activity primarily in the liver [57]. Thyroid hormone receptors (THRs) also directly promote mitochondrial biogenesis by driving the transcription of nuclear-encoded genes and indirectly via the thyroid hormone-mediated upregulation of NRF-1 [58]. A truncated form of THRα is localized to the mitochondrion and directly activates the transcription of mtDNA genes [59]. Another set of nuclear hormone receptors, estrogen-related receptors (ERRα/γ/δ), also promote mitochondrial function, including mitochondrial biogenesis, oxidative phosphorylation, fatty acid oxidation, the TCA cycle, and mitochondrial fusion/fission [60].
Other transcription factors
cAMP-activated transcription factor (CREB) promotes the expression of several mitochondrial genes involved in the TCA cycle and β-oxidation [61]. Many mitochondrial genes contain transcription factor Yin and Yang 1 (YY1) binding sites within their promoter regions, and YY1 works in conjunction with peroxisome proliferator-activated receptor-γ coactivator-1α (PGC-1α) to regulate their expression [62].
Transcriptional coactivators
Transcriptional coactivators do not bind to DNA but coactivate many different DNA-binding transcription factors, such as the peroxisome proliferator-activated receptor gamma coactivator 1 (PGC-1) family (PGC-1α, PGC-1β, and PRC). These factors potentiate the activity of several transcription factors involved in basic mitochondrial functions and biogenesis. PGC-1α is the master regulator of mtDNA transcription and mitochondrial biogenesis [63, 64]. PGC-1α induces mitochondrial biogenesis by activating different transcription factors, including NRF-1, NRF-2, and ERR-α, which interact with TFAM and promote its expression as the final effector of mtDNA transcription and replication. Epigenetic modifications, such as chromatin remodeling and DNA methylation, also play critical roles in mitochondrial biogenesis and functions. For example, recent studies demonstrated a critical role for histone-modifying proteins in the epigenetic control of the expression of genes implicated in mitochondrial fatty acid β-oxidation [65].
Nuclear posttranscription regulations
The nuclear genome also controls mitochondrial biogenesis and functions in a posttranscriptional regulation manner, including microRNA interference, RNA processing (RNA alternative splicing), and RNA stability.
Mitochondrial microRNAs interference
Microribonucleic acids (miRNAs) are short, single-stranded, noncoding ribonucleic acid (RNA) molecules (19–23 nucleotides) that prevent messenger RNA (mRNA) translation or induce the degradation of mRNA transcripts [66]. Although miRNAs are primarily located in the cytosol or nucleus, a subset of ~150 different miRNAs, called mitochondrial microRNA (mt-miRNA), localize to mitochondrial fractions. Mt-microRNA is transcribed from nuclear or mitochondrial genome and localize with the subunits of the RNA-induced silencing complex (RISC), which is the protein complex through which miRNAs normally act to prevent translation of their mRNA targets. Mt-miRNAs play important roles in mitochondrial function regulation. For example, miR-122, one of the most abundant adult hepatic miRNAs, is required for mitochondrial translation of respiratory proteins, improvement of mitochondrial respiratory enzyme activity, and enhancement of mitochondrial proteostasis in the liver [67]. An increasing number of mt-miRNAs modulate mitochondrial homeostasis by directly targeting mitochondrial function-related genes, such as miR-21, miR-29a, miR-33, and miR-34a.
Alternative splicing regulations
Most eukaryotic pre-mRNAs contain coding exonic sequences and noncoding intron sequences. An alternative splicing mechanism is needed to remove intron sequences and generate the correct concatenation of exonic sequences. One single gene may produce multiple mRNAs and structurally different proteins via alternative splicing and may affect more than 90% of human genes, including nuclear-encoded mitochondrial function-related genes [68]. Our later study revealed, for the first time, that DRAK2-SRSF6-mediated RNA alternative splicing of mitochondrial function-related genes may be one of the complementary mechanisms of mitochondrial homeostasis in hepatic steatosis to NASH [69]. We found a pathological alternative splicing form of Polg2, the coding gene of mtDNA polymerase γ2, in NASH model mouse livers and demonstrated that the pathological polg2 alternative splicing form influenced mitochondrial biogenesis and function. Notably, these results suggest an underlying mechanism by which the alternative splicing modulation of nuclear-encoded mitochondrial function-related genes plays an important role in mitochondrial biogenesis or other functions.
RNA stability
Post-transcriptional control of RNA stability is central to the regulation of gene expression and cellular function. This post-transcriptional process in mitochondria is also vital for proper expression of the 13 proteins encoded by the mitochondrial genome, mitochondrial tRNAs, and rRNAs. All factors involved in mtRNA stability are encoded by the nucleus and must be imported into the organelle. Defects in the machinery involved in human mitochondrial RNA stability are known causes of mitochondrial genome mutation and mitochondrial biogenesis, but the details still unknown [70]. Several studies in skeletal muscle investigated the stability of nuclear-encoded transcripts of mitochondrial biogenesis-related factors, such as NRF2, TFAM, and PGC-1α, that control mitochondrial biogenesis [71, 72]. However, the influence of mRNA stability on the expression of genes encoding mitochondrial proteins remains relatively unexplored and needs further study.
Mitochondrial fission and fusion dynamics
Mitochondria are highly dynamic organelles that morphologically continuously remodel and adapt to diverse cellular pathways, such as metabolism, intracellular calcium signaling, apoptosis, mitosis, and mitochondrial DNA replication [73]. Mitochondria undergo a continuous cycle that involves fusing together to form larger mitochondria and fission to break into smaller mitochondria. During fission, a single mitochondrion divides into two mitochondria via cleavage of the inner mitochondrial membrane (IMM) and outer mitochondrial membrane (OMM). During fusion, two mitochondria form one larger mitochondrion by the joining of the OMM and IMM, which occur in equilibrium to ensure mitochondrial network connectivity. Abnormal mitochondrial dynamics are associated with morphological, genetic, and biochemical mitochondrial recalibrations that trigger cellular stress responses and mitochondrial diseases [74].
Mitochondrial fission
Constriction and scission of the IMM and OMM are the key events of mitochondrial fission. Dynamin-related protein 1 (Drp1, also known as DNM1L) is a GTPase that is recruited to the OMM via the help of mitochondrial fission factor (MFF) and mitochondrial dynamics proteins 49 and 51 (MID49/51), and it is responsible for OMM constriction [75]. Once recruited, Drp1 oligomerizes to wrap around the outer membrane. Upon GTP hydrolysis, Drp1 dissociates MID49/51 to shrink the oligomeric ring and constricts the OMM to drive membrane scission. Drp1-dependent fission at mitochondria-ER contacts, which are the marker sites of mitochondrial division, is facilitated by actin assembly, and the inhibition of actin polymerization reduces fission frequency and Drp1 recruitment to mitochondria [76, 77]. Two actin nucleating proteins, formin protein inverted formin 2 (INF2) and actin-nucleating protein Spire (Spire1C), promote actin assembly and mitochondrial constriction and induce fission [78]. Recent advances revealed that nonmuscle myosin II (NMII) was located near mitochondrial constrictions and was involved, with actin and INF2, in fission, which is consistent with findings that NMII promotes Drp1 recruitment to mitochondria [79, 80]. Dynamin-2 (Dnm2), another dynamin GTPase, was found at fission sites after Drp1 recruitment and influenced mitochondrial fission [81]. Whereas outer membrane scission depends on Drp1, the mechanism of inner membrane scission is less clear. Recent studies showed that the IMM constriction and division at mitochondria-ER contacts were dependent on INF2-mediated actin polymerization and NMII, similar to outer membrane constriction, but the subsequent mechanism of IMM scission is not known [82].
Although the ER primarily coordinates mitochondrial fission, several other factors determine the sites of fission. RAB7-GTP is recruited to the OMM by mitochondrial fission protein 1 (Fis1) and promotes mitochondria-lysosome contact formation, which are also marker sites of fission, to restrict mitochondrial motility [83]. The trans-Golgi network (TGN) also modulates fission. After Drp1 recruitment, the small GTPase ADP-ribosylation factor 1 (Arf1) and its effector, phosphatidylinositol 4-kinase-III-b [PI(4)KIIIb], are recruited to fission sites on TGN vesicles and promote fission [84]. Notably, TGN vesicles converge with lysosomes and the ER at fission sites. Further study is necessary to understand how these organelles coordinate to promote fission.
Mitochondrial fusion
The dynamin family GTPases mitofusin 1 (Mfn1), mitofusin 2 (Mfn2), and optic atrophy protein 1 (Opa1) are required for mitochondrial fusion mechanisms and regulation. Fusion begins with Mfn1/2-mediated OMM tethering and merging followed by Opa1-mediated joining of the IMM [85]. Mfn1 and Mfn2 anchor to the mitochondrial outer membrane via the C-terminal membrane-binding domain, which extrudes the N-terminal GTPase domain to the cytoplasm [86]. However, their mechanisms of action in mitochondrial membrane fusion are not known. Opa1 has two isoforms, a long isoform (L-Opa1) containing a transmembrane domain and a short isoform (S-Opa1) that lacks the transmembrane domain due to proteolytic cleavage of L-Opa1 by the proteases ATP-dependent zinc metalloprotease (Yme1L) or OMA1 zinc metallopeptidase (Oma1) [87]. One study found that L-Opa1 was sufficient for fusion, without Yme1L and Oma1, and S-Opa1 overexpression in these cells resulted in mitochondrial fragmentation [88]. However, Opa1 processing tightly regulates fusion, with higher and lower ratios of S-Opa1 to L-Opa1 inhibiting fusion [89].
Additional insight into mitochondrial fusion comes from mitochondria-ER contact sites. Mitochondrial fusion also occurs at mitochondria-ER contact sites, similar to fission [90]. Fission and fusion proteins colocalize at mitochondria-ER contacts for membrane dynamics. Therefore, the ER regulates multiple aspects of mitochondrial dynamics at contact sites. How these separate machineries are coordinated to promote a single process needs further exploration.
Mitophagy
The autophagic system, termed mitophagy, targets impaired mitochondria and delivers them to lysosomes for degradation, which is a fundamental mechanism that is conserved from yeast to humans that regulates mitochondrial quality and quantity control. Mitophagy is promoted via specific mitochondrial outer membrane receptors or ubiquitin molecules conjugated to proteins on the mitochondrial surface, which leads to the formation of autophagosomes surrounding mitochondria. Mitophagy-mediated elimination of mitochondria plays an important role in modulating mitochondrial fitness and number in normal and disease physiology.
PINK1/Parkin pathway
The mitophagy field was constructed on investigations of the PINK1/Parkin pathway, which is the most characterized mitophagy pathway. Under normal conditions, PTEN-induced putative kinase protein 1 (PINK1) is imported into the mitochondria then exposed to and cleaved by the mitochondrial proteases MPP and PARL [91]. Once mitochondrial depolarization or misfolded mitochondrial proteins accumulate, PINK1 cannot be imported into mitochondria and stabilized on the surface [92]. PINK1 phosphorylates ubiquitin attached to the E3 ligase Parkin or other OMM proteins [93]. Phosphorylation encourages Parkin recruitment to these mitochondria and ubiquitinates multiple surface proteins, such as NDP52 and OPTN, which results in more phosphorylation and an amplification loop that sequentially escalates the signal for the degradation on the surface of mitochondria [94, 95].
Although the PINK1/Parkin pathway is indisputable for mitophagy in vitro assays, mice lacking PINK1 or Parkin do not exhibit a phenotype, and the loss of either protein in the heart or brain does not affect the levels of basal mitophagy [96]. PINK1/Parkin-mediated mitophagy is triggered by severe stress, and there are other pathways that maintain basal levels of mitophagy when stress is milder. Therefore, it makes sense to have a ubiquitin-independent group of mitophagy receptors that may be activated and balance the mitophagy level via other means.
Mitophagy receptor pathways
Mitophagy receptors contain an LIR motif that enables the recruitment of LC3 and the growing mitophagophore to the mitochondria [97]. The mitophagy receptor NIP3-like protein X (NIX) mediates mitophagy during red blood cell (RBC) differentiation or the hypoxia-driven glycolytic switch during metabolic transitions [98, 99]. BCL2/adenovirus E1B 19 kDa protein-interacting protein 3 (BNIP3) is also involved in PINK1 stabilization, DRP1 translocation, and BECN1 freeing during PINK1-Parkin mitophagy pathway activation [100, 101]. FUN14 domain-containing protein 1 (FUNDC1), the final mitophagy receptor induced by hypoxia, is also regulated via phosphorylation within the LIR motif, instead of being transcriptionally regulated. In addition to hypoxia-mediated mitophagy, FUNDC1 was also implicated in depolarization-induced mitophagy via interaction with IP3R2 and is regulated through direct phosphorylation by Unc-51-like kinase 1 (ULK1) [102, 103].
There are also several less studied receptors associated with mitophagy. FK506-binding protein 8 (FKBP8), another mitophagy receptor, mediates mitophagy and fission by binding LC3A, independent of Parkin [104]. Bcl-2-like protein 13 (BCL2L13), another OMM protein, mediates mitophagy by binding LC3 and promoting mitochondrial fission [105]. Prohibitin 2 (PHB2), located at the IMM, regulates proteasome-driven OMM rupture and is involved in PINK1-Parkin mitophagy. Upon OMM rupture, mitochondrial depolarization and PHB2 regulate mitophagy via PINK1 stability on mitochondria and LC3 binding [106, 107]. NLR family member X1 (NLRX1) is a NOD-like receptor located within the mitochondrial matrix and contains an LIR motif [108]. Cardiolipin is a unique phospholipid in the IMM that is also a mitophagy receptor. Upon mitochondrial damage, cardiolipin translocates to the OMM, where it interacts with LC3 and may be involved in PINK1-Parkin mitophagy [109].
Mitochondrial dysfunction in NAFLD and NASH
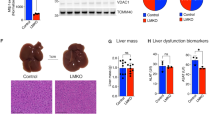
The major feature of NAFLD/NASH is excess lipid accumulation in the liver, with inflammation and liver damage. During NAFLD development, there is a constant dysfunction of mitochondria, including alterations in enzyme activities, protein expression, and signaling networks [110]. These processes are tightly coupled with mitochondrial quality control, hepatocyte cell death, and inflammatory responses (Fig. 2).
Western diet drives the mitochondrial TCA cycle and induces lipogenesis and lipid droplet formation. Hepatic lipid accumulation accelerates insulin resistance in the liver and adipose tissue, which results in a massive flux of FFAs into the liver from adipose tissue. FFAs overload in the liver and hepatic insulin resistance results in inefficient β-oxidation and uncouples mitochondrial TCA cycle activity from OXPHOS, which leads to excessive ROS generation. Hepatic ROS accumulation and lipotoxicity caused by lipid overload promote hepatocyte cell death via apoptosis or necrosis. Dysfunctional hepatocytes secrete microparticles containing chemotactic signals (inflammatory or necrosis factor) into the extracellular matrix, which induce hepatic resistant immune cells (macrophages) or recruit immune cells from bone marrow activation. The fibrogenic signal derived from dysfunctional hepatocytes and/or inflammatory cells activates HSCs and promotes the development of liver fibrosis. Overnutrition or intestinal microbiota-derived signal stimulation induces mitochondrial metabolic remodeling in macrophages and HSCs, which further accelerates the development of NASH directly or indirectly. Abbreviations: ROS reactive oxygen species, TCA cycle tricarboxylic acid cycle, FFA free fatty acid, OXPHOS oxidative phosphorylation system, mtDNA mitochondrial DNA, HSCs hepatic stellate cell, NASH nonalcoholic steatohepatitis.
Changes in mitochondrial lipid metabolism in NAFLD livers
Under normal conditions, the liver balances lipid degradation and lipogenesis. However, lipid homeostasis is altered in NAFLD. During the pathophysiology of NAFLD, lipid accumulation is excessive, and the oxidative catabolism of FFAs is insufficient. Under fasting conditions, the proportion of triglycerides (TGs) stored in the liver of NAFLD patients is 59.0% ± 9.9% from plasma FFAs, 26.1% ± 6.7% from de novo lipogenesis (DNL), and 14.9% ± 7.0% from dietary intake, which were traced using orally fed stable isotopes in humans, and the DNL rate is up to three-fold higher in NAFLD patients than healthy humans [111, 112]. The expression of lipogenic-related factors and enzymes, such as sterol regulatory element-binding protein (SREBP1c), carbohydrate-responsive element-binding protein (ChREBP), ACLY, fatty acid synthase (FAS), acetyl-CoA carboxylase 1 (ACC1), and acyl-CoA desaturase 1 (SCD1), is much higher in fatty livers than normal healthy livers [113]. Moreover, the expression of SREBP1c and lipogenesis can also be induced by tumor necrosis factor-alpha (TNF-α) [113]. Mitochondria from NAFLD livers show increased TCA cycle function, and the overloading of acetyl-CoA in the cytoplasm accelerates lipogenesis. Overactivated TCA cycle produces excessive ROS and mitochondrial damage, which also induce systemic insulin resistance and inflammation. High levels of insulin contribute to lipogenic enzyme expression and lipogenesis.
Continuously increased lipogenesis inhibits fatty acid oxidation for intermediate malonyl-CoA production by inhibiting carnitine palmitoyltransferase 1 (CPT1) activity [114]. However, the change in mitochondrial β-oxidation is closely coordinated with steatohepatitis and the function of mitochondria in patients with NAFLD or NASH [110, 115]. Fatty acid utilization is excessive in the early stage of NAFLD development [116]. However, the expression of genes related to β-oxidation is decreased in the progressive NASH livers compared to simple NAFL, and these genes are partially regulated by PPARα. Although hepatic PPARα levels do not differ between NAFLD patients with simple steatosis and healthy controls, PPARα was downregulated in patients with NASH compared to patients with steatosis and healthy controls [117, 118]. Therefore, overactivated β-oxidation in mitochondria leads to a remarkable increase in ROS in the simple steatosis stage, which promotes the occurrence of NASH by inducing oxidative stress and inflammation.
Ketogenesis in the liver is the third flux for the acetyl-CoA pool produced by FAO and glycolysis (the other two are the TCA cycle and lipogenesis). The serum β-hydroxybutyrate concentration negatively correlates with the fat content in the liver. Compared to high-fat diet (HFD)-fed mice, ketogenic insufficiency aggravated hepatic lipogenesis, inflammation, and injury in HFD-fed mice [115, 119]. Hepatocyte-derived acetoacetate (AcAc) acts as the energy substrate for macrophages, which ameliorates diet-induced hepatic fibrosis [120]. All of these results indicate that the extent of FAO and the function of mitochondria are altered during the progression of simple steatosis and NASH, which are tightly associated with mitochondrial metabolism and ROS generation.
Mitochondrial defects stimulate ROS production and oxidative stress in fatty liver
The intracellular balance of ROS and antioxidants is disrupted with the overproduction of free radicals in fatty liver [121]. The electronic respiratory chain of mitochondria in hepatocyte generates the major content of ROS in the process of NAFLD. The excessive ROS produced by dysfunctional mitochondria trigger pathological redox signaling and lead to serious hepatocyte DNA damage, apoptosis or necrosis, immune cell infiltration, and hepatic stellate cell (HSC) activation, which participates in the progression of NAFLD and NASH development [122].
The mitochondrial β-oxidation rate is increased significantly during rapid lipid overload in the liver to restrain hepatic fat accretion, which accelerates the accumulation of intercellular ROS. Inactivation of complexes I and III in the presence of fatty acids, especially polyunsaturated fatty acids (PUFAs), enhances O2•- production via the electronic respiratory chain [123, 124]. Fatty acid incorporation into the inner membrane of mitochondria also increases membrane fluidity and promotes electron leakage. Hepatic iron overload occurs in some NASH patients and facilitates the conversion of H2O2 to extremely toxic HO• via the Fenton reaction [125]. Lipotoxicity induces unfolded and misfolded protein accumulation and leads to the unfolded protein response (UPR), which generates stress in the ER and increases ROS production [126].
ROS production in the liver contributes to a large range of pathologies in the progression of NAFLD development. ROS overload decreases mitochondrial ETC activity and opens the mitochondrial permeability transition pore (mPTP), and the subsequent release of cytochrome c to the cytosol from mitochondria induces the apoptotic pathway in the liver [29, 127]. Excess ROS oxidize phospholipids, which increase the activation of proinflammatory and apoptotic signaling pathways in the liver [128]. NAFLD-induced lipid overload decreases sarco/ER Ca2+-ATPase (SERCA) activity. The excess calcium released into the cytoplasm is absorbed by the mitochondria and exacerbates cell death via mPTP [129].
During ROS-induced liver injury, the activation of redox-sensitive transcription factors, such as nuclear factor kappa B (NF-κB), are increased, and the inflammatory response is activated. The nucleotide-binding domain and leucine-rich repeat containing PYD-3 (NLRP3) inflammasome induce caspase 1-dependent release of the proinflammatory cytokines interleukin-1β (IL-1β) and interleukin-18 (IL-18), which induce liver cell death under lipid overload conditions. As one of many important NLRP3 inflammasome activators, ROS are important NLRP3 activators that promote NAFLD development [130]. ROS activate macrophages and HSCs, which induce fibrosis in hepatic tissue. ROS-induced macrophage activation triggers innate and adaptive immune responses with the release of proinflammatory cytokines and chemokines, which activate natural killer T cells and HSCs [131, 132].
Mitochondrial dysfunction and liver cell death
Animal models and clinical studies clearly demonstrated that hepatocyte death was an important driver of NASH progression. Liver injury, induced by multiple stimulations, is at the center of NASH development [133]. Various forms of cell death are observed in the liver, including apoptosis, necrosis, and necroptosis. Hepatocyte apoptosis induced by lipotoxicity (including fatty acids and cholesterol) and ROS stress are well-recognized cell death pathways in NASH. Apoptosis signal-regulating kinase 1 (ASK1) is ubiquitously expressed and activated by pathological stimulation in human and animal NASH livers [134]. Therefore, caspase inhibitors, such as pancaspase inhibitors and ASK1 inhibitors, were protective in animal models of NASH [135].
Homodimerization of the apoptosis-related proteins Bax and Bak within the OMM, which belong to the Bcl-2 family of proteins, is a direct activator of hepatocyte apoptosis [136]. Mitochondrial dysfunction is tightly associated with hepatocyte apoptosis. Excessive lipid immersion in hepatocytes promotes fatty acid β-oxidation, which promotes ROS and related byproducts, such as oxidized phospholipids (OxPLs) and oxidized cholesterol accumulation. Oxidative stress initiates hepatocyte apoptosis and liver injury. A recent study indicated that loss-of-function augmentation of liver regeneration (ALR), which is a flavin-containing oxidase localized in the mitochondrial intermembrane space, induced mitochondrial release of cytochrome c and accelerated the development of NASH [137]. Mitochondrial cytochrome c release leads to caspase-mediated hepatic apoptosis, and the energy sensor AMP-activated protein kinase (AMPK) phosphorylates proapoptotic caspase-6 protein to inhibit its activation via cytochrome c from mitochondria [138]. Apoptotic hepatocytes stimulate immune cell activation or infiltration and HSC activation toward the progression of NASH via the production of inflammasomes and cytokines [139]. These data suggest that mitochondria-initiated apoptosis is extremely important in hepatic injury and NASH development.
Mitochondrial metabolism regulates macrophage infiltration in fatty liver
Macrophage phenotypes exhibit marked metabolic plasticity in the liver, which depends on environmental stimulation. Kupffer cells and monocyte-derived macrophages represent distinct origins of hepatic macrophage populations that perform a range of metabolic functions. Various dangerous molecules and fatty acids promote a proinflammatory phenotype in macrophages, which contributes to the development of persistent, low-grade inflammation in the early stage of NAFLD. Proinflammatory cytokines, chemokines, mtDNA, apoptosis, or necrosis signals, which are transmitted from dysfunctional hepatocytes, the gut-liver axis, and adipocytes, stimulate the activation or infiltration of macrophages into the liver [140, 141].
Macrophage metabolic disorder induced by multiple energy stresses also plays a role in NASH development. The activities of isocitrate dehydrogenase (IDH) and succinate dehydrogenase (SDH) are much lower than quiescent macrophages, which induce the accumulation of citrate and succinate in macrophages. Citrate is exported into the cytoplasm and cleaved by ACLY into acetyl-CoA and oxaloacetate, which acts as an epigenetic modifier or NADPH provider and induces inflammatory factor production [142]. Succinate induces mitochondrial ROS production or HIF-1α and acts as a signal to activate proinflammatory-related gene expression and cytokine secretion [143]. Itaconate is generated by the mitochondria-associated enzyme immune responsive gene 1 (IRG1) from the TCA cycle metabolite cis-aconitate in the mitochondrial matrix, which potently modulates macrophage metabolism by inhibiting succinate dehydrogenase-mediated oxidation of succinate in the TCA cycle [144]. Glutamine is effective in inducing the polarization of M2 macrophages via the glutamine-UDP-N-acetylglucosamine pathway and α-ketoglutarate produced via glutaminolysis, and succinate synthesized via glutamine-dependent anerplerosis or the γ-aminobutyric acid shunt promotes the polarization of M1 macrophages. Macrophage dysfunction induces mtDNA release, which stimulates innate immune receptors and activates the inflammatory response in the liver [145, 146].
Mitochondrial-related HSC activation
Activated HSCs produce extracellular matrix (ECM) proteins and sustain the wound-healing process to gradually develop liver fibrosis when hepatocytes lose their function [147]. Mitochondrial signals from neighboring cells, such as hepatocytes and immune cells, lead to HSCs activation. Hepatocytes release damage-associated molecular patterns derived from mitochondria (mito-DAMPs) upon injury or death [148]. Hepatocyte mito-DAMPs are elevated in human NAFLD/NASH and directly trigger profibrogenic HSCs activation [149]. Hepatocytes release microparticles to activate HSCs upon increased oxidative stress and apoptotic death induced by chemical hypoxia [150].
Decades of studies indicated that mitochondrial dysfunction-induced oxidative stress is a major inducer of HSCs activation in the development of NASH [151]. The NAPDH oxidase enzyme proteins NOX1, NOX2, NOX4, and NOX5 are upregulated prominently in activated HSCs [152,153,–154]. Intracellular ROS provoke NLRP3 inflammasome activation, which contributes to HSC activation [155]. Recent research identified that the redox enzyme Shc (p66Shc) was significantly increased in human fibrosis and mediated mitochondrial ROS generation and triggered NLRP3 inflammasome and HSCs activation [156]. Morphological stimulation also manipulates HSCs activation. The downregulation of ALR in HSCs, which is associated with F-actin assembly, is also attributed to the promotion of HSCs migration and mitochondrial fusion during hepatic fibrosis [157].
An important source of ECM in the liver is hepatocyte epithelial mesenchymal transition (EMT). Oxidative stress-induced hepatic damage and mtDNA deletion are closely associated with abnormal mitochondrial fission, EMT, and liver fibrosis via PGC-1α transcriptional regulation [158]. Therefore, antioxidant reagents may attenuate transforming growth factor β1 (TGF-β1)- and platelet-derived growth factor (PDGF)-stimulated HSCs activation and improve liver diseases [159].
Therapies targeting mitochondria in NAFLD and NASH
The goal of anti-NASH drugs is to reduce the accumulation of liver fat, inflammation, and fibrosis. Despite the high prevalence and potential consequences, there are no approved treatments for NASH. Calorie restriction and weight loss are the only effective means to clinically reduce fat in the liver. Mitochondria are the center of cellular metabolism. A total of 31%–40% of patients with steatohepatitis have lower hepatic mitochondrial respiratory levels than obese patients. This difference indicates that improvements in mitochondrial dysfunction may contribute to the treatment of NASH.
PPARs agonists for NASH treatment
Hepatic lipid overload is a hallmark of NASH. Therefore, stimulating mitochondrial fatty acid oxidation reduces lipotoxicity and improves hepatic dysfunction. Gene expression controlled by PPAR-α and -β is related to mitochondrial and peroxisomal β-oxidative catabolism and ketogenesis. PPAR-γ is a critical regulator of adipocyte differentiation and lipogenesis and acts as a functional insulin sensitizer. Therefore, a series of PPARs regulators were identified and used in NASH treatment [160].
PPARα agonists (primarily fibrates) reduce alanine aminotransferase (ALT) and aspartate aminotransferase (AST) activity and inflammatory factor content in the serum. However, a long-term clinical study demonstrated that fenofibrate or clofibrate treatment did not improve the liver histology of NASH [161]. Oral administration of the PPARα/δ agonist elafibranor reduced hepatic steatosis inflammation and fibrosis in a mouse model, but the antifibrotic effects were limited in phase II and phase III studies [162]. The PPAR-γ agonist pioglitazone reduced hepatic steatosis and inflammation but not the concomitant fibrosis [163]. The pan-PPARs agonist lanifibranor, which targets the three PPAR isotypes, reduced hepatic steatosis and inflammation in different mouse models of NASH. A recent phase 2b trial showed that lanifibranor treatment achieved significant therapeutic effects on NASH resolution and did not worsen fibrosis [164].
Fibroblast growth factor 21 (FGF21) is a hepatokine that is expressed and secreted in response to PPARα activation in the fasting state. Recent FGF21 analogs demonstrated efficacy in animal models and humans with NASH by inducing β-oxidation and improving mitochondrial function [165]. Farnesoid X receptor (FXR) is a metabolic regulator that plays a key role in bile acid metabolism. FXR agonists demonstrated promising clinical results in the treatment of NASH [166]. FXR activation increased the expression of PPARα and related genes in humans [167]. Whether FXR agonist treatment improves mitochondrial function in NASH patients requires further study.
Thyroxine receptorβ (THRβ) agonists for NASH treatment
The THRβ isoform is responsible for the metabolic rate induced by thyroxine in the liver. THR activation stimulates mitochondrial fatty acid β-oxidation and oxidative phosphorylation in the liver [168]. Studies from patients also showed an inverse relationship between NASH liver function and serum thyroxine levels [169]. Therefore, THRβ agonists are a treatment strategy for NASH. Resmetirom (MGL-3196) is a recently developed liver-directed THRβ-selective agonist by Madrigal Pharmaceuticals that exhibited great potential for the treatment of NASH. Resmetirom administration reduced hepatic fat and significantly decreased the expression of inflammation and fibrosis marker genes, which was a positive phase II clinical trial result in patients with NASH. Mechanistically, THRβ agonists decrease steatosis in the liver by increasing fatty acid β-oxidation via the induction of autophagy, mitochondrial biogenesis, and enzyme expression for lipid oxidation [170].
Beyond transcriptional regulation, other potential NASH therapeutic strategies in clinical trials are tightly associated with mitochondrial function improvement. GLP-1 receptor agonists were promising anti-NASH candidates in clinical phase III studies. A recent study showed that the GLP-1R/GcgR dual agonist cotadudidide improved NASH by modulating mitochondrial function [171]. The cellular energy sensor AMPK improves mitochondrial function via multiple mechanisms [172, 173]. The AMPK activator PXL770 met its primary efficacy endpoint in a phase II trial for NASH treatment [174].
Antioxidants for NAFLD and NASH treatment
Intra/extra-mitochondrial oxidative stress is a major player that triggers the progression of NASH. Recent literature strongly suggests that vitamin E or other antioxidant supplementation is beneficial in improving the clinical biochemical and histological parameters in NASH.
Vitamin E is a chain-breaking antioxidant in free radical reactions and may be an option for the treatment of NAFLD and NASH. Preclinical animal studies showed that vitamin E administration reduced mitochondrial lipid peroxidation and corrected oxidative stress, improved TGFβ1-induced fibrosis, and reduced the content of TNF-α. Notable reductions in oxidative stress and cytokine markers in NASH patients treated with vitamin E were observed, which support the role of vitamin E as an antioxidant [175, 176]. Vitamin E treatment improved liver function and reduced liver fat in NASH patients in clinical trial studies. Vitamin E combined with vitamin C was a potent antioxidant for NASH treatment or was protective against NAFLD-related liver damage [177]. However, a recent study also indicated that vitamin E alone improved steatosis but did not significantly change inflammation, ballooning, or fibrosis in NASH patients with type 2 diabetes, which indicates the multilevel and multifactorial pathogenesis of NASH development [178]. Several other antioxidants, such as N-acetylcysteine (NAC), betaine, and probucol, were also evaluated for their potential therapeutic effects in NASH patients [179,180,–181]. Accumulating data demonstrated that antioxidants may be effective candidates for NASH treatment.
Dissipation of abnormal liver lipid accumulation by mitochondrial uncoupler
Mitochondrial uncoupling is a process that shuttles protons across the IMM via a pathway that is independent of ATP synthesis. Therefore, the energy derived from fat oxidation in OXPHOS is directly converted to heat rather than ATP synthesis [182]. Small-molecule mitochondrial uncouplers mimic the mild physiological effects of mitochondrial uncoupling in the liver. Rodent and nonhuman primate studies indicated that mitochondrial uncouplers safely reversed NAFLD/NASH [183]. The therapeutic effects of mitochondrial uncouplers are often linked to increased fatty acid β-oxidation, activation of the AMPK response to inefficient ATP production, and a reduction in ROS production.
The most notable mitochondrial uncoupler, 2,4-dinitrophenol (DNP), is widely used as a weight-loss agent in obese humans [184]. However, a series of toxic side effects, including hyperthermia, cataracts, agranulocytosis, and death, were reported continuously, which caused the US Food and Drug Administration (FDA) to ban its use in the 1930s [185]. Several attempts were made to develop pharmacological agents based on DNP to discover safe chemical uncouplers. Perry et al. developed a prodrug named DNPME that targeted DNP in the liver with a 50–200-fold greater therapeutic window than DNP [186]. They also identified an extended-release formulation of DNP (CRMP). Oral administration of CRMP prominently increased hepatic lipid utilization and reduced hepatic steatosis [187]. Wei et al. developed a liquid crystal gel formulation with extended-release properties (DNP-LC gel) that safely reduced dyslipidemia and hepatic steatosis in a rat model [188].
Researchers also identified a series of chemical uncouplers that exhibited therapeutic promise for NAFLD treatment. We synthesized a novel compound (6j) with mitochondrial uncoupling activity and pyruvate dehydrogenase activation effects. The administration of 6j improved hyperglycemia and hepatic steatosis [189]. Jian et al. identified that sorafenib, an extracellular signal-regulated kinase (ERK) inhibitor for HCC treatment, acted as a mitochondrial uncoupler to improve NASH in rodent and nonhuman primate models [190]. Salamoun and Alexopoulos et al. identified that BAM15 and its two BAM15 derivatives were efficacious for anti-NASH in a STAM murine model [191, 192]. All of these studies indicate that modestly increased uncoupling mimics some of the benefits of calorie restriction, and small molecule mitochondrial uncoupler treatment increases cellular energy expenditure, which is a promising therapeutic strategy to improve fatty liver.
Mitochondrial microRNAs interference
Many studies revealed the significance of mt-miRNA in modulating mitochondrial homeostasis and improving the pathophysiology of NAFLD by directly targeting key genes. Therefore, abnormal mt-miRNA expression may result in mitochondrial dysfunction, which alters lipid metabolism, oxidative stresses, and inflammation in the liver and plays a crucial role in the pathophysiology of NAFLD.
MiR-29a protects mitochondrial structural integrity, restricts mito-DAMPs, and exerts an anti-inflammatory effect on the pathogenesis of NAFLD by targeting voltage-dependent anion channels and Bcl-2-associated X genes, whose oligomerization is involved in mPTP opening and mito-DAMPs release [193, 194]. One computational analysis revealed that miR-29a targeted DRP1 [195], which suggests its potential role in regulating mitochondrial dynamics and mitophagy. MiR-29a inhibits glycogen synthase kinase-3β (GSK3β) to suppress sirtuin 1 (SIRT1)-mediated mitochondrial biogenesis, which leads to alleviation of mitochondrial proteostatic stress and UPRmt in the context of NASH [196]. Hepatic miR‑34a expression is increased from steatosis to less and more severe NASH [197]. Inhibition of miR-34a mitigated steatosis in an experimental NAFLD model [198]. MiR-34a exhibited suppressive activity on PPARα via direct association with its mRNA 3′UTR [198], which decreased mitochondrial biogenesis and increased oxidative stress and the inflammatory response. The miR-34a/SIRT1/AMPK pathway caused mitochondrial dynamics dysfunction in a mouse NASH model [199]. MiR-34a impaired the mitochondrial quality control mechanism via SIRT3/FoxO3a/PINK1 signaling in an experimental mouse model of liver inflammation [200]. GLP-1 treatment increased the expression of the mitochondrial protective gene PGC-1α by downregulating miR-23a to inhibit hepatocyte apoptosis [201].
Other miRNAs also impact pathways that regulate mitochondrial functions in the liver, such as miR-122, miR-370, and miR-21 [202,203,–204]. Many miRNAs play important roles in mitochondrial function, but their roles in liver diseases are not clear, such as miRNA-144 [205], miRNA-145 [206], and miRNA-126 [207]. However, other miRNAs, such as miRNA-103/107 and miRNA-221/222, play key roles during NAFLD pathophysiology via a mechanism other than mitochondrial function regulation [208, 209].
The available pieces of evidence suggest a new framework for considering and understanding mt-miRNA as a novel biomarker in mitochondrial dysfunction by linking it with many complex diseases, including NAFLD and NASH. The analysis of miRNA profiles in serum, plasma, and blood cells linked with the development and progression of mitochondrial dysfunctions may lead to novel therapeutic strategies for NAFLD/NASH. MiRNAs may serve as potential noninvasive prognostic and diagnostic markers for NAFLD/NASH. Significant research endeavors must be exercised on miRNAs to determine their future use and clinical application. The applicability of miRNAs in mitochondrial research remains elusive.
Nuclear-encoded mitochondrial gene alternative splicing
Alterations in mRNA splicing are important causes of disease, and alterations in this pathway are found in many human diseases. Our recent study revealed a novel mechanism underlying NAFLD/NASH progression via the dysregulation of mitochondrial function-related gene alternative splicing via DRAK2-SRSF6 signaling and suggests that targeting this mechanism is a promising approach for NAFLD/NASH treatment. We found pathological alternative splicing forms of many mitochondrial function-related genes, such as Polg2, Rnasel, Nme4, Nudt13, and Guf1, which suggests a close relationship between the pathological alternative splicing form and mitochondrial function during NAFLD development. We also used the DRAK2 inhibitor 22b to explore the therapeutic potential of targeting the DRAK2-SRSF6 axis during the progression of NAFLD/NASH. Notably, HFD-induced alternative splicing alterations in mitochondrial function-related genes, including Polg2, Nudt13, Guf1, Rnasel, and Nme4, and SRSF6 hyperphosphorylation were largely blocked by 22b intervention, which further supports the feasibility of targeting nuclear-encoded mitochondrial gene alternative splicing to treat NAFLD/NASH [69].
PGC-1a is a transcriptional coactivator that is expressed as multiple alternatively spliced variants transcribed from different promoters that coordinate metabolic adaptation and protect against inflammation. Transcription initiated from the PGC-1a proximal promoter generates canonical PGC-1a1 (also known as PGC-1a-a) and NT-PGC-1a-a is a truncated version containing a 31-nucleotide insertion between exons 6 and 7 that generates a premature stop codon [210]. Overexpression of NT-PGC-1a-a and PGC-1a1 drives similar gene programs, including mitochondrial biogenesis and fatty acid oxidation [210, 211]. PGC-1a variants have distinct yet complementary roles in hepatic mitochondrial function via alternative RNA splicing.
The mitochondrial fission GTPase Drp1 also expresses multiple splicing variants. Mitochondrial fission is mediated by the GTPase activity of Drp1. Cooperative GTPase activity is contingent upon intra- and intermolecular interactions between the four major domains of Drp1 [212]. Different Drp1 isoforms may affect this process because variations in the GTPase and variable domain of Drp1 have an allosteric effect on the preferred curvature and cooperative GTPase activity of Drp1 polymers [213]. The posttranslational modifications of Drp1 vary for the alternate Drp1 isoforms and result in distinct functions [214, 215]. Therefore, different Drp1 isoforms have distinct and overlapping roles in mitochondrial fission and cell death, which suggests a potential role of Drp1 alternative splicing in NAFLD and NASH.
There are many other mitochondrial function-related genes with different isoforms arising from alternative splicing, such as 12 complex I-associated genes (NDUFA3, NDUFA13, NDUFA8, NDUFS2, NDUFS4, NDUFA4, NDUFA12, NDUFB6, NDUFV1, NDUFA5, NDUFB11, and NDUFA7), OPA1, and MRPL33 [216], but the functions of many different isoforms are not clear and need further study.
Risdiplam (Evrysdi™), an orally administered, survival motor neuron 2 (SMN2)-directed RNA splicing modifier developed by Roche, PTC Therapeutics Inc., and the SMA Foundation, received its first approval in the USA for the treatment of spinal muscular atrophy [217, 218], which indicates a novel therapeutic strategy targeting alternative splicing modification. Various studies investigated the role of alternative splicing in disease development and the use of alternative splicing events as diagnostic markers and therapeutic targets for various diseases. Given the role of alternative splicing in mitochondrial function, targeting nuclear-encoded mitochondrial gene alternative splicing may be a promising approach for NAFLD/NASH treatment. Further investigation is warranted to determine how these genes play different roles in mitochondrial function with different alternative splicing forms and to elucidate their relevance to human NAFLD/NASH etiology.
Conclusion and perspective
NAFLD is the most common chronic liver disease worldwide and impacts 25% of the world’s population. With NAFLD spreading, the prevalence of NASH, liver cirrhosis, and HCC inevitably increases. Given the overall burden of the disease, further studies are urgently needed to identify novel therapeutic targets for NAFLD prevention and treatment. Although several new drugs and molecular targets are promising, many clinical trials concluded that the optimal pharmacological approach will require modification of the complex pathogenesis of NAFLD and NASH. Comprehensive evidence supports a pivotal role of mitochondrial dysfunction in hepatic steatosis to NASH pathogenesis, which indicates one promising therapeutic strategy targeting mitochondrial homeostasis. The present review highlighted the mitochondrial biology and homeostasis involved in hepatic steatosis to NASH and reviewed the current therapeutic approaches in NAFLD with an emphasis on mitochondria as potential therapeutic targets. Despite progress in our understanding of mitochondrial dysfunction in NAFLD/NASH and the therapeutic potential of targeting mitochondrial homeostasis mechanisms, several questions remain.
First, many of these findings were derived from experimentation on animal models of simple steatosis and NASH, and translation of these results into human subjects with a spectrum of disease stages remains a major priority. Second, although comprehensive evidence supports a pivotal role of mitochondrial dysfunction in hepatic steatosis to NASH pathogenesis, mitochondrial biology and pathology during NAFLD/NASH are not fully understood. For example, the key point at which mitochondrial adaptation remains in hepatic steatosis but is lost in NASH and the balance of mitochondrial dynamics, mitophagy and mitochondrial biogenesis, are not well understood. Third, hepatic mitochondria maintain a tight balance between fat oxidation and the generation of ROS, and therapies aimed at increasing mitochondrial activity may reduce steatosis but at the cost of increasing inflammation and fibrosis, especially in humans [219]. Fourth, despite exciting preclinical data, translation of mitochondria-targeted agents into clinical use remains a major challenge because of their potential adverse effects, unclear mechanism of action, and the unclear mitochondrial biology and pathology underlying these conditions. Therefore, further studies should focus on identifying the role and regulation of mitochondrial homeostasis mechanisms during the development of hepatic steatosis to NASH pathogenesis to facilitate the discovery of pharmacological modulators to prevent and treat this disease.
References
Loomba R, Friedman SL, Shulman GI. Mechanisms and disease consequences of nonalcoholic fatty liver disease. Cell. 2021;184:2537–64.
Zhou F, Zhou J, Wang W, Zhang XJ, Ji YX, Zhang P, et al. Unexpected rapid increase in the burden of NAFLD in China from 2008 to 2018: A systematic review and meta-analysis. Hepatology. 2019;70:1119–33.
Diehl AM, Day C. Cause, pathogenesis, and treatment of nonalcoholic steatohepatitis. N Engl J Med. 2017;377:2063–72.
Anstee QM, Reeves HL, Kotsiliti E, Govaere O, Heikenwalder M. From NASH to HCC: Current concepts and future challenges. Nat Rev Gastroenterol Hepatol. 2019;16:411–28.
Day CP, James OF. Steatohepatitis: A tale of two “hits”? Gastroenterology. 1998;114:842–5.
Alonso C, Fernandez-Ramos D, Varela-Rey M, Martinez-Arranz I, Navasa N, Van Liempd SM, et al. Metabolomic identification of subtypes of nonalcoholic steatohepatitis. Gastroenterology. 2017;152:1449–61. e1447.
Buzzetti E, Pinzani M, Tsochatzis EA. The multiple-hit pathogenesis of non-alcoholic fatty liver disease (NAFLD). Metabolism. 2016;65:1038–48.
Friedman JR, Nunnari J. Mitochondrial form and function. Nature. 2014;505:335–43.
Chavez JD, Tang X, Campbell MD, Reyes G, Kramer PA, Stuppard R, et al. Mitochondrial protein interaction landscape of SS-31. Proc Natl Acad Sci USA. 2020;117:15363–73.
Mansouri A, Gattolliat CH, Asselah T. Mitochondrial dysfunction and signaling in chronic liver diseases. Gastroenterology. 2018;155:629–47.
Rath S, Sharma R, Gupta R, Ast T, Chan C, Durham TJ, et al. MitoCarta3.0: An updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucleic Acids Res. 2021;49:D1541–D1547.
Schmidt O, Pfanner N, Meisinger C. Mitochondrial protein import: From proteomics to functional mechanisms. Nat Rev Mol Cell Biol. 2010;11:655–67.
Rui L. Energy metabolism in the liver. Compr Physiol. 2014;4:177–97.
Morio B, Panthu B, Bassot A, Rieusset J. Role of mitochondria in liver metabolic health and diseases. Cell Calcium. 2021;94:102336.
Wilson DF. Oxidative phosphorylation: Regulation and role in cellular and tissue metabolism. J Physiol. 2017;595:7023–38.
Enriquez JA. Supramolecular organization of respiratory complexes. Annu Rev Physiol. 2016;78:533–61.
Brown GC. The relative proton stoichiometries of the mitochondrial proton pumps are independent of the proton motive force. J Biol Chem. 1989;264:14704–9.
Ramakrishna R, Edwards JS, McCulloch A, Palsson BO. Flux-balance analysis of mitochondrial energy metabolism: consequences of systemic stoichiometric constraints. Am J Physiol Regul Integr Comp Physiol. 2001;280:R695–704.
Badman MK, Pissios P, Kennedy AR, Koukos G, Flier JS, Maratos-Flier E. Hepatic fibroblast growth factor 21 is regulated by PPARalpha and is a key mediator of hepatic lipid metabolism in ketotic states. Cell Metab. 2007;5:426–37.
McGarry JD, Foster DW. Regulation of hepatic fatty acid oxidation and ketone body production. Annu Rev Biochem. 1980;49:395–420.
Zhao S, Jang C, Liu J, Uehara K, Gilbert M, Izzo L, et al. Dietary fructose feeds hepatic lipogenesis via microbiota-derived acetate. Nature. 2020;579:586–91.
Wellen KE, Hatzivassiliou G, Sachdeva UM, Bui TV, Cross JR, Thompson CB. ATP-citrate lyase links cellular metabolism to histone acetylation. Science. 2009;324:1076–80.
Huh JY, Reilly SM, Abu-Odeh M, Murphy AN, Mahata SK, Zhang J, et al. TANK-binding kinase 1 regulates the localization of acyl-CoA synthetase ACSL1 to control hepatic fatty acid oxidation. Cell Metab. 2020;32:1012–27. e1017.
White PJ, McGarrah RW, Grimsrud PA, Tso SC, Yang WH, Haldeman JM, et al. The BCKDH kinase and phosphatase integrate BCAA and lipid metabolism via regulation of ATP-citrate lyase. Cell Metab. 2018;27:1281–93. e1287.
Xiao D, Zeng L, Yao K, Kong X, Wu G, Yin Y. The glutamine-alpha-ketoglutarate (AKG) metabolism and its nutritional implications. Amino Acids. 2016;48:2067–80.
Berndsen CE, Denu JM. Catalysis and substrate selection by histone/protein lysine acetyltransferases. Curr Opin Struct Biol. 2008;18:682–9.
Lucas S, Chen G, Aras S, Wang J. Serine catabolism is essential to maintain mitochondrial respiration in mammalian cells. Life Sci Alliance. 2018;1:e201800036.
Ducker GS, Rabinowitz JD. One-carbon metabolism in health and disease. Cell Metab. 2017;25:27–42.
Zorov DB, Juhaszova M, Sollott SJ. Mitochondrial reactive oxygen species (ROS) and ROS-induced ROS release. Physiol Rev. 2014;94:909–50.
Thomas C, Mackey MM, Diaz AA, Cox DP. Hydroxyl radical is produced via the Fenton reaction in submitochondrial particles under oxidative stress: Implications for diseases associated with iron accumulation. Redox Rep. 2009;14:102–8.
Murphy MP. How mitochondria produce reactive oxygen species. Biochem J. 2009;417:1–13.
Scialo F, Sriram A, Fernandez-Ayala D, Gubina N, Lohmus M, Nelson G, et al. Mitochondrial ROS produced via reverse electron transport extend animal lifespan. Cell Metab. 2016;23:725–34.
Bedard K, Krause KH. The NOX family of ROS-generating NADPH oxidases: Physiology and pathophysiology. Physiol Rev. 2007;87:245–313.
Ischiropoulos H, Zhu L, Chen J, Tsai M, Martin JC, Smith CD, et al. Peroxynitrite-mediated tyrosine nitration catalyzed by superoxide dismutase. Arch Biochem Biophys. 1992;298:431–7.
Pfanner N, Warscheid B, Wiedemann N. Mitochondrial proteins: From biogenesis to functional networks. Nat Rev Mol Cell Biol. 2019;20:267–84.
Fontana GA, Gahlon HL. Mechanisms of replication and repair in mitochondrial DNA deletion formation. Nucleic Acids Res. 2020;48:11244–58.
Robberson DL, Kasamatsu H, Vinograd J. Replication of mitochondrial DNA. Circular replicative intermediates in mouse L cells. Proc Natl Acad Sci USA. 1972;69:737–41.
Miralles Fuste J, Shi Y, Wanrooij S, Zhu X, Jemt E, Persson O, et al. In vivo occupancy of mitochondrial single-stranded DNA binding protein supports the strand displacement mode of DNA replication. PLoS Genet. 2014;10:e1004832.
Fuste JM, Wanrooij S, Jemt E, Granycome CE, Cluett TJ, Shi Y, et al. Mitochondrial RNA polymerase is needed for activation of the origin of light-strand DNA replication. Mol Cell. 2010;37:67–78.
Yasukawa T, Reyes A, Cluett TJ, Yang MY, Bowmaker M, Jacobs HT, et al. Replication of vertebrate mitochondrial DNA entails transient ribonucleotide incorporation throughout the lagging strand. EMBO J. 2006;25:5358–71.
Holt IJ, Lorimer HE, Jacobs HT. Coupled leading- and lagging-strand synthesis of mammalian mitochondrial DNA. Cell. 2000;100:515–24.
Montoya J, Christianson T, Levens D, Rabinowitz M, Attardi G. Identification of initiation sites for heavy-strand and light-strand transcription in human mitochondrial DNA. Proc Natl Acad Sci USA. 1982;79:7195–9.
Hillen HS, Morozov YI, Sarfallah A, Temiakov D, Cramer P. Structural basis of mitochondrial transcription initiation. Cell. 2017;171:1072–81. e1010.
Ramachandran A, Basu U, Sultana S, Nandakumar D, Patel SS. Human mitochondrial transcription factors TFAM and TFB2M work synergistically in promoter melting during transcription initiation. Nucleic Acids Res. 2017;45:861–74.
Minczuk M, He J, Duch AM, Ettema TJ, Chlebowski A, Dzionek K, et al. TEFM (c17orf42) is necessary for transcription of human mtDNA. Nucleic Acids Res. 2011;39:4284–99.
Yakubovskaya E, Mejia E, Byrnes J, Hambardjieva E, Garcia-Diaz M. Helix unwinding and base flipping enable human MTERF1 to terminate mitochondrial transcription. Cell. 2010;141:982–93.
Terzioglu M, Ruzzenente B, Harmel J, Mourier A, Jemt E, Lopez MD, et al. MTERF1 binds mtDNA to prevent transcriptional interference at the light-strand promoter but is dispensable for rRNA gene transcription regulation. Cell Metab. 2013;17:618–26.
Cai YC, Bullard JM, Thompson NL, Spremulli LL. Interaction of mitochondrial elongation factor Tu with aminoacyl-tRNA and elongation factor Ts. J Biol Chem. 2000;275:20308–14.
Lind C, Sund J, Aqvist J. Codon-reading specificities of mitochondrial release factors and translation termination at non-standard stop codons. Nat Commun. 2013;4:2940.
Spencer AC, Spremulli LL. Interaction of mitochondrial initiation factor 2 with mitochondrial fMet-tRNA. Nucleic Acids Res. 2004;32:5464–70.
D’Souza AR, Minczuk M. Mitochondrial transcription and translation: Overview. Essays Biochem. 2018;62:309–20.
Soleimanpour-Lichaei HR, Kuhl I, Gaisne M, Passos JF, Wydro M, Rorbach J, et al. mtRF1a is a human mitochondrial translation release factor decoding the major termination codons UAA and UAG. Mol Cell. 2007;27:745–57.
Rorbach J, Richter R, Wessels HJ, Wydro M, Pekalski M, Farhoud M, et al. The human mitochondrial ribosome recycling factor is essential for cell viability. Nucleic Acids Res. 2008;36:5787–99.
Scarpulla RC. Transcriptional paradigms in mammalian mitochondrial biogenesis and function. Physiol Rev. 2008;88:611–38.
Carter RS, Bhat NK, Basu A, Avadhani NG. The basal promoter elements of murine cytochrome c oxidase subunit IV gene consist of tandemly duplicated ets motifs that bind to GABP-related transcription factors. J Biol Chem. 1992;267:23418–26.
Desvergne B, Wahli W. Peroxisome proliferator-activated receptors: Nuclear control of metabolism. Endocr Rev. 1999;20:649–88.
Aoyama T, Peters JM, Iritani N, Nakajima T, Furihata K, Hashimoto T, et al. Altered constitutive expression of fatty acid-metabolizing enzymes in mice lacking the peroxisome proliferator-activated receptor alpha (PPARalpha). J Biol Chem. 1998;273:5678–84.
Venditti P, Bari A, Di Stefano L, Cardone A, Della Ragione F, D’Esposito M, et al. Involvement of PGC-1, NRF-1, and NRF-2 in metabolic response by rat liver to hormonal and environmental signals. Mol Cell Endocrinol. 2009;305:22–29.
Casas F, Rochard P, Rodier A, Cassar-Malek I, Marchal-Victorion S, Wiesner RJ, et al. A variant form of the nuclear triiodothyronine receptor c-ErbAalpha1 plays a direct role in regulation of mitochondrial RNA synthesis. Mol Cell Biol. 1999;19:7913–24.
Eichner LJ, Giguere V. Estrogen related receptors (ERRs): A new dawn in transcriptional control of mitochondrial gene networks. Mitochondrion. 2011;11:544–52.
Gopalakrishnan L, Scarpulla RC. Differential regulation of respiratory chain subunits by a CREB-dependent signal transduction pathway. Role of cyclic AMP in cytochrome c and COXIV gene expression. J Biol Chem. 1994;269:105–13.
Cunningham JT, Rodgers JT, Arlow DH, Vazquez F, Mootha VK, Puigserver P. mTOR controls mitochondrial oxidative function through a YY1-PGC-1alpha transcriptional complex. Nature. 2007;450:736–40.
Puigserver P, Wu Z, Park CW, Graves R, Wright M, Spiegelman BM. A cold-inducible coactivator of nuclear receptors linked to adaptive thermogenesis. Cell. 1998;92:829–39.
Deng XH, Liu JJ, Sun XJ, Dong JC, Huang JH. Benzoylaconine induces mitochondrial biogenesis in mice via activating AMPK signaling cascade. Acta Pharmacol Sin. 2019;40:658–65.
Seok S, Kim YC, Byun S, Choi S, Xiao Z, Iwamori N, et al. Fasting-induced JMJD3 histone demethylase epigenetically activates mitochondrial fatty acid beta-oxidation. J Clin Invest. 2018;128:3144–59.
Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993;75:843–54.
Bandiera S, Pfeffer S, Baumert TF, Zeisel MB. miR-122–a key factor and therapeutic target in liver disease. J Hepatol. 2015;62:448–57.
Licatalosi DD, Darnell RB. RNA processing and its regulation: Global insights into biological networks. Nat Rev Genet. 2010;11:75–87.
Li Y, Xu J, Lu Y, Bian H, Yang L, Wu H. et al. DRAK2 aggravates nonalcoholic fatty liver disease progression through SRSF6-associated RNA alternative splicing. Cell Metab. 2021;33:2004–20 e2009.
Gagliardi D, Stepien PP, Temperley RJ, Lightowlers RN, Chrzanowska-Lightowlers ZM. Messenger RNA stability in mitochondria: different means to an end. Trends Genet. 2004;20:260–7.
Lai RY, Ljubicic V, D’Souza D, Hood DA. Effect of chronic contractile activity on mRNA stability in skeletal muscle. Am J Physiol Cell Physiol. 2010;299:C155–163.
D’Souza D, Lai RY, Shuen M, Hood DA. mRNA stability as a function of striated muscle oxidative capacity. Am J Physiol Regul Integr Comp Physiol. 2012;303:R408–417.
Fenton AR, Jongens TA, Holzbaur ELF. Mitochondrial dynamics: Shaping and remodeling an organelle network. Curr Opin Cell Biol. 2021;68:28–36.
Youle RJ, van der Bliek AM. Mitochondrial fission, fusion, and stress. Science. 2012;337:1062–5.
Palmer CS, Elgass KD, Parton RG, Osellame LD, Stojanovski D, Ryan MT. Adaptor proteins MiD49 and MiD51 can act independently of Mff and Fis1 in Drp1 recruitment and are specific for mitochondrial fission. J Biol Chem. 2013;288:27584–93.
Ji WK, Hatch AL, Merrill RA, Strack S, Higgs HN. Actin filaments target the oligomeric maturation of the dynamin GTPase Drp1 to mitochondrial fission sites. Elife. 2015;4:e11553.
Friedman JR, Lackner LL, West M, DiBenedetto JR, Nunnari J, Voeltz GK. ER tubules mark sites of mitochondrial division. Science. 2011;334:358–62.
Korobova F, Ramabhadran V, Higgs HN. An actin-dependent step in mitochondrial fission mediated by the ER-associated formin INF2. Science. 2013;339:464–7.
Yang C, Svitkina TM. Ultrastructure and dynamics of the actin-myosin II cytoskeleton during mitochondrial fission. Nat Cell Biol. 2019;21:603–13.
Korobova F, Gauvin TJ, Higgs HN. A role for myosin II in mammalian mitochondrial fission. Curr Biol. 2014;24:409–14.
Lee JE, Westrate LM, Wu H, Page C, Voeltz GK. Multiple dynamin family members collaborate to drive mitochondrial division. Nature. 2016;540:139–43.
Chakrabarti R, Ji WK, Stan RV, de Juan Sanz J, Ryan TA, Higgs HN. INF2-mediated actin polymerization at the ER stimulates mitochondrial calcium uptake, inner membrane constriction, and division. J Cell Biol. 2018;217:251–68.
Wong YC, Ysselstein D, Krainc D. Mitochondria-lysosome contacts regulate mitochondrial fission via RAB7 GTP hydrolysis. Nature. 2018;554:382–6.
Nagashima S, Tabara LC, Tilokani L, Paupe V, Anand H, Pogson JH, et al. Golgi-derived PI(4)P-containing vesicles drive late steps of mitochondrial division. Science. 2020;367:1366–71.
Cipolat S, Martins de Brito O, Dal Zilio B, Scorrano L. OPA1 requires mitofusin 1 to promote mitochondrial fusion. Proc Natl Acad Sci USA. 2004;101:15927–32.
Fritz S, Rapaport D, Klanner E, Neupert W, Westermann B. Connection of the mitochondrial outer and inner membranes by Fzo1 is critical for organellar fusion. J Cell Biol. 2001;152:683–92.
Mishra P, Carelli V, Manfredi G, Chan DC. Proteolytic cleavage of Opa1 stimulates mitochondrial inner membrane fusion and couples fusion to oxidative phosphorylation. Cell Metab. 2014;19:630–41.
Anand R, Wai T, Baker MJ, Kladt N, Schauss AC, Rugarli E, et al. The i-AAA protease YME1L and OMA1 cleave OPA1 to balance mitochondrial fusion and fission. J Cell Biol. 2014;204:919–29.
Ge Y, Shi X, Boopathy S, McDonald J, Smith AW, Chao LH. Two forms of Opa1 cooperate to complete fusion of the mitochondrial inner-membrane. Elife. 2020;9:e50973.
Abrisch RG, Gumbin SC, Wisniewski BT, Lackner LL, Voeltz GK. Fission and fusion machineries converge at ER contact sites to regulate mitochondrial morphology. J Cell Biol. 2020;219:e201911122.
Deas E, Plun-Favreau H, Gandhi S, Desmond H, Kjaer S, Loh SH, et al. PINK1 cleavage at position A103 by the mitochondrial protease PARL. Hum Mol Genet. 2011;20:867–79.
Narendra DP, Jin SM, Tanaka A, Suen DF, Gautier CA, Shen J, et al. PINK1 is selectively stabilized on impaired mitochondria to activate Parkin. PLoS Biol. 2010;8:e1000298.
Koyano F, Okatsu K, Kosako H, Tamura Y, Go E, Kimura M, et al. Ubiquitin is phosphorylated by PINK1 to activate parkin. Nature. 2014;510:162–6.
Lazarou M, Sliter DA, Kane LA, Sarraf SA, Wang C, Burman JL, et al. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature. 2015;524:309–14.
Heo JM, Ordureau A, Paulo JA, Rinehart J, Harper JW. The PINK1-PARKIN mitochondrial ubiquitylation pathway drives a program of OPTN/NDP52 recruitment and TBK1 activation to promote mitophagy. Mol Cell. 2015;60:7–20.
McWilliams TG, Prescott AR, Montava-Garriga L, Ball G, Singh F, Barini E, et al. Basal mitophagy occurs independently of PINK1 in mouse tissues of high metabolic demand. Cell Metab. 2018;27:439–49. e435
Wang Y, Liu N, Lu B. Mechanisms and roles of mitophagy in neurodegenerative diseases. CNS Neurosci Ther. 2019;25:859–75.
Schweers RL, Zhang J, Randall MS, Loyd MR, Li W, Dorsey FC, et al. NIX is required for programmed mitochondrial clearance during reticulocyte maturation. Proc Natl Acad Sci USA. 2007;104:19500–5.
Sandoval H, Thiagarajan P, Dasgupta SK, Schumacher A, Prchal JT, Chen M, et al. Essential role for Nix in autophagic maturation of erythroid cells. Nature. 2008;454:232–5.
Zhang T, Xue L, Li L, Tang C, Wan Z, Wang R, et al. BNIP3 protein suppresses PINK1 kinase proteolytic cleavage to promote mitophagy. J Biol Chem. 2016;291:21616–29.
Lee Y, Lee HY, Hanna RA, Gustafsson AB. Mitochondrial autophagy by Bnip3 involves Drp1-mediated mitochondrial fission and recruitment of Parkin in cardiac myocytes. Am J Physiol Heart Circ Physiol. 2011;301:H1924–1931.
Wu W, Tian W, Hu Z, Chen G, Huang L, Li W, et al. ULK1 translocates to mitochondria and phosphorylates FUNDC1 to regulate mitophagy. EMBO Rep. 2014;15:566–75.
Wu S, Lu Q, Wang Q, Ding Y, Ma Z, Mao X, et al. Binding of FUN14 domain containing 1 with inositol 1,4,5-trisphosphate receptor in mitochondria-associated endoplasmic reticulum membranes maintains mitochondrial dynamics and function in hearts in vivo. Circulation. 2017;136:2248–66.
Bhujabal Z, Birgisdottir AB, Sjottem E, Brenne HB, Overvatn A, Habisov S, et al. FKBP8 recruits LC3A to mediate Parkin-independent mitophagy. EMBO Rep. 2017;18:947–61.
Murakawa T, Yamaguchi O, Hashimoto A, Hikoso S, Takeda T, Oka T, et al. Bcl-2-like protein 13 is a mammalian Atg32 homologue that mediates mitophagy and mitochondrial fragmentation. Nat Commun. 2015;6:7527.
Wei Y, Chiang WC, Sumpter R Jr, Mishra P, Levine B. Prohibitin 2 is an inner mitochondrial membrane mitophagy receptor. Cell. 2017;168:224–38. e210.
Yan C, Gong L, Chen L, Xu M, Abou-Hamdan H, Tang M, et al. PHB2 (prohibitin 2) promotes PINK1-PRKN/Parkin-dependent mitophagy by the PARL-PGAM5-PINK1 axis. Autophagy. 2020;16:419–34.
Zhang Y, Yao Y, Qiu X, Wang G, Hu Z, Chen S, et al. Listeria hijacks host mitophagy through a novel mitophagy receptor to evade killing. Nat Immunol. 2019;20:433–46.
Chu CT, Ji J, Dagda RK, Jiang JF, Tyurina YY, Kapralov AA, et al. Cardiolipin externalization to the outer mitochondrial membrane acts as an elimination signal for mitophagy in neuronal cells. Nat Cell Biol. 2013;15:1197–205.
Koliaki C, Szendroedi J, Kaul K, Jelenik T, Nowotny P, Jankowiak F, et al. Adaptation of hepatic mitochondrial function in humans with non-alcoholic fatty liver is lost in steatohepatitis. Cell Metab. 2015;21:739–46.
Donnelly KL, Smith CI, Schwarzenberg SJ, Jessurun J, Boldt MD, Parks EJ. Sources of fatty acids stored in liver and secreted via lipoproteins in patients with nonalcoholic fatty liver disease. J Clin Invest. 2005;115:1343–51.
Lambert JE, Ramos-Roman MA, Browning JD, Parks EJ. Increased de novo lipogenesis is a distinct characteristic of individuals with nonalcoholic fatty liver disease. Gastroenterology. 2014;146:726–35.
Todoric J, Di Caro G, Reibe S, Henstridge DC, Green CR, Vrbanac A, et al. Fructose stimulated de novo lipogenesis is promoted by inflammation. Nat Metab. 2020;2:1034–45.
McGarry JD, Mannaerts GP, Foster DW. A possible role for malonyl-CoA in the regulation of hepatic fatty acid oxidation and ketogenesis. J Clin Invest. 1977;60:265–70.
Fletcher JA, Deja S, Satapati S, Fu X, Burgess SC, Browning JD. Impaired ketogenesis and increased acetyl-CoA oxidation promote hyperglycemia in human fatty liver. JCI Insight. 2019;4:e127737.
Sunny NE, Parks EJ, Browning JD, Burgess SC. Excessive hepatic mitochondrial TCA cycle and gluconeogenesis in humans with nonalcoholic fatty liver disease. Cell Metab. 2011;14:804–10.
Francque S, Verrijken A, Caron S, Prawitt J, Paumelle R, Derudas B, et al. PPARalpha gene expression correlates with severity and histological treatment response in patients with non-alcoholic steatohepatitis. J Hepatol. 2015;63:164–73.
Videla LA, Pettinelli P. Misregulation of PPAR functioning and its pathogenic consequences associated with nonalcoholic fatty liver disease in human obesity. PPAR Res. 2012;2012:107434.
d’Avignon DA, Puchalska P, Ercal B, Chang Y, Martin SE, Graham MJ, et al. Hepatic ketogenic insufficiency reprograms hepatic glycogen metabolism and the lipidome. JCI Insight. 2018;3:e99762.
Puchalska P, Martin SE, Huang X, Lengfeld JE, Daniel B, Graham MJ, et al. Hepatocyte-macrophage acetoacetate shuttle protects against tissue fibrosis. Cell Metab. 2019;29:383–98. e387
Chen Z, Tian R, She Z, Cai J, Li H. Role of oxidative stress in the pathogenesis of nonalcoholic fatty liver disease. Free Radic Biol Med. 2020;152:116–41.
Rolo AP, Teodoro JS, Palmeira CM. Role of oxidative stress in the pathogenesis of nonalcoholic steatohepatitis. Free Radic Biol Med. 2012;52:59–69.
Loskovich MV, Grivennikova VG, Cecchini G, Vinogradov AD. Inhibitory effect of palmitate on the mitochondrial NADH:ubiquinone oxidoreductase (complex I) as related to the active-de-active enzyme transition. Biochem J. 2005;387:677–83.
Cocco T, Di Paola M, Papa S, Lorusso M. Arachidonic acid interaction with the mitochondrial electron transport chain promotes reactive oxygen species generation. Free Radic Biol Med. 1999;27:51–9.
Dongiovanni P, Fracanzani AL, Fargion S, Valenti L. Iron in fatty liver and in the metabolic syndrome: a promising therapeutic target. J Hepatol. 2011;55:920–32.
Rinella ME, Siddiqui MS, Gardikiotes K, Gottstein J, Elias M, Green RM. Dysregulation of the unfolded protein response in db/db mice with diet-induced steatohepatitis. Hepatology. 2011;54:1600–9.
Xiao D, He H, Huang W, Oo TL, Wang A, He LF. Analysis of mitochondrial markers of programmed cell death. Methods Mol Biol. 2018;1743:65–71.
Que X, Hung MY, Yeang C, Gonen A, Prohaska TA, Sun X, et al. Oxidized phospholipids are proinflammatory and proatherogenic in hypercholesterolaemic mice. Nature. 2018;558:301–6.
Fu S, Yang L, Li P, Hofmann O, Dicker L, Hide W, et al. Aberrant lipid metabolism disrupts calcium homeostasis causing liver endoplasmic reticulum stress in obesity. Nature. 2011;473:528–31.
Wan X, Xu C, Yu C, Li Y. Role of NLRP3 inflammasome in the progression of NAFLD to NASH. Can J Gastroenterol Hepatol. 2016;2016:6489012.
Brune B, Dehne N, Grossmann N, Jung M, Namgaladze D, Schmid T, et al. Redox control of inflammation in macrophages. Antioxid Redox Signal. 2013;19:595–637.
Tsuchida T, Friedman SL. Mechanisms of hepatic stellate cell activation. Nat Rev Gastroenterol Hepatol. 2017;14:397–411.
Luedde T, Kaplowitz N, Schwabe RF. Cell death and cell death responses in liver disease: Mechanisms and clinical relevance. Gastroenterology. 2014;147:765–83. e764
Schuster S, Feldstein AE. NASH: Novel therapeutic strategies targeting ASK1 in NASH. Nat Rev Gastroenterol Hepatol. 2017;14:329–30.
Loomba R, Lawitz E, Mantry PS, Jayakumar S, Caldwell SH, Arnold H, et al. The ASK1 inhibitor selonsertib in patients with nonalcoholic steatohepatitis: A randomized, phase 2 trial. Hepatology. 2018;67:549–59.
Pena-Blanco A, Garcia-Saez AJ. Bax, Bak and beyond—mitochondrial performance in apoptosis. FEBS J. 2018;285:416–31.
Kumar S, Verma AK, Rani R, Sharma A, Wang J, Shah SA, et al. Hepatic deficiency of augmenter of liver regeneration predisposes to nonalcoholic steatohepatitis and fibrosis. Hepatology. 2020;72:1586–604.
Zhao P, Sun X, Chaggan C, Liao Z, In Wong K, He F, et al. An AMPK-caspase-6 axis controls liver damage in nonalcoholic steatohepatitis. Science. 2020;367:652–60.
Hirsova P, Gores GJ. Death receptor-mediated cell death and proinflammatory signaling in nonalcoholic steatohepatitis. Cell Mol Gastroenterol Hepatol. 2015;1:17–27.
Cha JY, Kim DH, Chun KH. The role of hepatic macrophages in nonalcoholic fatty liver disease and nonalcoholic steatohepatitis. Lab Anim Res. 2018;34:133–9.
Wang Y, Li N, Zhang X, Horng T. Mitochondrial metabolism regulates macrophage biology. J Biol Chem. 2021;297:100904.
Noe JT, Mitchell RA. Tricarboxylic acid cycle metabolites in the control of macrophage activation and effector phenotypes. J Leukoc Biol. 2019;106:359–67.
Mills EL, Kelly B, Logan A, Costa ASH, Varma M, Bryant CE. et al. Succinate dehydrogenase supports metabolic repurposing of mitochondria to drive inflammatory macrophages. Cell. 2016;167:457–70 e413.
Lampropoulou V, Sergushichev A, Bambouskova M, Nair S, Vincent EE, Loginicheva E, et al. Itaconate links inhibition of succinate dehydrogenase with macrophage metabolic remodeling and regulation of inflammation. Cell Metab. 2016;24:158–66.
Wu G, Zhu Q, Zeng J, Gu X, Miao Y, Xu W, et al. Extracellular mitochondrial DNA promote NLRP3 inflammasome activation and induce acute lung injury through TLR9 and NF-kappaB. J Thorac Dis. 2019;11:4816–28.
Pan J, Ou Z, Cai C, Li P, Gong J, Ruan XZ, et al. Fatty acid activates NLRP3 inflammasomes in mouse Kupffer cells through mitochondrial DNA release. Cell Immunol. 2018;332:111–20.
Friedman SL. Hepatic stellate cells: Protean, multifunctional, and enigmatic cells of the liver. Physiological Rev. 2008;88:125–72.
Krysko DV, Agostinis P, Krysko O, Garg AD, Bachert C, Lambrecht BN, et al. Emerging role of damage-associated molecular patterns derived from mitochondria in inflammation. Trends Immunol. 2011;32:157–64.
An P, Wei LL, Zhao S, Sverdlov DY, Vaid KA, Miyamoto M, et al. Hepatocyte mitochondria-derived danger signals directly activate hepatic stellate cells and drive progression of liver fibrosis. Nat Commun. 2020;11:2362.
Hernández A, Reyes D, Geng Y, Arab JP, Cabrera D, Sepulveda R, et al. Extracellular vesicles derived from fat-laden hepatocytes undergoing chemical hypoxia promote a pro-fibrotic phenotype in hepatic stellate cells. Biochim Biophys Acta (BBA)—Mol Basis Dis. 2020;1866:165857.
Gajendiran P, Vega LI, Itoh K, Sesaki H, Vakili MR, Lavasanifar A, et al. Elevated mitochondrial activity distinguishes fibrogenic hepatic stellate cells and sensitizes for selective inhibition by mitotropic doxorubicin. J Cell Mol Med. 2018;22:2210–9.
Sancho P, Mainez J, Crosas-Molist E, Roncero C, Fernández-Rodriguez CM, Pinedo F, et al. NADPH oxidase NOX4 mediates stellate cell activation and hepatocyte cell death during liver fibrosis development. PLoS ONE. 2012;7:e45285.
Paik YH, Iwaisako K, Seki E, Inokuchi S, Schnabl B, Österreicher CH, et al. The nicotinamide adenine dinucleotide phosphate oxidase (NOX) homologues NOX1 and NOX2/gp91phox mediate hepatic fibrosis in mice. Hepatology. 2011;53:1730–41.
Andueza A, Garde N, García-Garzón A, Ansorena E, López-Zabalza MJ, Iraburu MJ, et al. NADPH oxidase 5 promotes proliferation and fibrosis in human hepatic stellate cells. Free Radic Biol Med. 2018;126:15–26.
Wree A, Eguchi A, McGeough MD, Pena CA, Johnson CD, Canbay A, et al. NLRP3 inflammasome activation results in hepatocyte pyroptosis, liver inflammation, and fibrosis in mice. Hepatology. 2014;59:898–910.
Zhao Y, Wang Z, Feng D, Zhao H, Lin M, Hu Y, et al. p66Shc contributes to liver fibrosis through the regulation of mitochondrial reactive oxygen species. Theranostics. 2019;9:1510–22.
Ai WL, Dong LY, Wang J, Li ZW, Wang X, Gao J, et al. Deficiency in augmenter of liver regeneration accelerates liver fibrosis by promoting migration of hepatic stellate cell. Biochim Biophys Acta Mol Basis Dis. 2018;1864:3780–91.
Zhang L, Zhang Y, Chang X, Zhang X. Imbalance in mitochondrial dynamics induced by low PGC-1α expression contributes to hepatocyte EMT and liver fibrosis. Cell Death Dis. 2020;11:226.
Foo NP, Lin SH, Lee YH, Wu MJ, Wang YJ. α-Lipoic acid inhibits liver fibrosis through the attenuation of ROS-triggered signaling in hepatic stellate cells activated by PDGF and TGF-β. Toxicology. 2011;282:39–46.
Boeckmans J, Natale A, Rombaut M, Buyl K, Rogiers V, De Kock J et al. Anti-NASH drug development hitches a lift on PPAR agonism. Cells. 2019;9:37.
Fernandez-Miranda C, Perez-Carreras M, Colina F, Lopez-Alonso G, Vargas C, Solis-Herruzo JA. A pilot trial of fenofibrate for the treatment of non-alcoholic fatty liver disease. Dig Liver Dis. 2008;40:200–5.
Ratziu V, Harrison SA, Francque S, Bedossa P, Lehert P, Serfaty L, et al. Elafibranor, an agonist of the peroxisome proliferator-activated receptor-alpha and -delta, induces resolution of nonalcoholic steatohepatitis without fibrosis worsening. Gastroenterology. 2016;150:1147–59. e1145
Sanyal AJ, Chalasani N, Kowdley KV, McCullough A, Diehl AM, Bass NM, et al. Pioglitazone, vitamin E, or placebo for nonalcoholic steatohepatitis. N Engl J Med. 2010;362:1675–85.
Francque SM, Bedossa P, Ratziu V, Anstee QM, Bugianesi E, Sanyal AJ, et al. A randomized, controlled trial of the Pan-PPAR agonist lanifibranor in NASH. N Engl J Med. 2021;385:1547–58.
Zarei M, Pizarro-Delgado J, Barroso E, Palomer X, Vazquez-Carrera M. Targeting FGF21 for the treatment of nonalcoholic steatohepatitis. Trends Pharm Sci. 2020;41:199–208.
Kremoser C. FXR agonists for NASH: How are they different and what difference do they make? J Hepatol. 2021;75:12–15.
Pineda Torra I, Claudel T, Duval C, Kosykh V, Fruchart JC, Staels B. Bile acids induce the expression of the human peroxisome proliferator-activated receptor alpha gene via activation of the farnesoid X receptor. Mol Endocrinol. 2003;17:259–72.
Sinha RA, Bruinstroop E, Singh BK, Yen PM. Nonalcoholic fatty liver disease and hypercholesterolemia: Roles of thyroid hormones, metabolites, and agonists. Thyroid. 2019;29:1173–91.
Guo Z, Li M, Han B, Qi X. Association of non-alcoholic fatty liver disease with thyroid function: A systematic review and meta-analysis. Dig Liver Dis. 2018;50:1153–62.
Luong XG, Stevens SK, Jekle A, Lin TI, Gupta K, Misner D, et al. Regulation of gene transcription by thyroid hormone receptor beta agonists in clinical development for the treatment of non-alcoholic steatohepatitis (NASH). PLoS ONE. 2020;15:e0240338.
Boland ML, Laker RC, Mather K, Nawrocki A, Oldham S, Boland BB, et al. Resolution of NASH and hepatic fibrosis by the GLP-1R/GcgR dual-agonist Cotadutide via modulating mitochondrial function and lipogenesis. Nat Metab. 2020;2:413–31.
Herzig S, Shaw RJ. AMPK: guardian of metabolism and mitochondrial homeostasis. Nat Rev Mol Cell Biol. 2018;19:121–35.
Shirwany NA, Zou MH. AMPK in cardiovascular health and disease. Acta Pharmacol Sin. 2010;31:1075–84.
Cusi K, Alkhouri N, Harrison SA, Fouqueray P, Moller DE, Hallakou-Bozec S, et al. Efficacy and safety of PXL770, a direct AMP kinase activator, for the treatment of non-alcoholic fatty liver disease (STAMP-NAFLD): A randomised, double-blind, placebo-controlled, phase 2a study. Lancet Gastroenterol Hepatol. 2021;6:889–902.
Sokol RJ, McKim JM Jr., Goff MC, Ruyle SZ, Devereaux MW, Han D, et al. Vitamin E reduces oxidant injury to mitochondria and the hepatotoxicity of taurochenodeoxycholic acid in the rat. Gastroenterology. 1998;114:164–74.
Parola M, Muraca R, Dianzani I, Barrera G, Leonarduzzi G, Bendinelli P, et al. Vitamin E dietary supplementation inhibits transforming growth factor beta 1 gene expression in the rat liver. FEBS Lett. 1992;308:267–70.
Ivancovsky-Wajcman D, Fliss-Isakov N, Salomone F, Webb M, Shibolet O, Kariv R, et al. Dietary vitamin E and C intake is inversely associated with the severity of nonalcoholic fatty liver disease. Dig Liver Dis. 2019;51:1698–705.
Bril F, Biernacki DM, Kalavalapalli S, Lomonaco R, Subbarayan SK, Lai J, et al. Role of vitamin E for nonalcoholic steatohepatitis in patients with type 2 diabetes: A randomized controlled trial. Diabetes Care. 2019;42:1481–8.
Khoshbaten M, Aliasgarzadeh A, Masnadi K, Tarzamani MK, Farhang S, Babaei H, et al. N-acetylcysteine improves liver function in patients with non-alcoholic fatty liver disease. Hepat Mon. 2010;10:12–16.
Abdelmalek MF, Sanderson SO, Angulo P, Soldevila-Pico C, Liu C, Peter J, et al. Betaine for nonalcoholic fatty liver disease: Results of a randomized placebo-controlled trial. Hepatology. 2009;50:1818–26.
Merat S, Aduli M, Kazemi R, Sotoudeh M, Sedighi N, Sohrabi M, et al. Liver histology changes in nonalcoholic steatohepatitis after one year of treatment with probucol. Dig Dis Sci. 2008;53:2246–50.
Busiello RA, Savarese S, Lombardi A. Mitochondrial uncoupling proteins and energy metabolism. Front Physiol. 2015;6:36.
Goedeke L, Shulman GI. Therapeutic potential of mitochondrial uncouplers for the treatment of metabolic associated fatty liver disease and NASH. Mol Metab. 2021;46:101178.
Tainter ML, Cutting WC, Stockton AB. Use of dinitrophenol in nutritional disorders: A critical survey of clinical results. Am J Public Health Nations Health. 1934;24:1045–53.
Grundlingh J, Dargan PI, El-Zanfaly M, Wood DM. 2,4-dinitrophenol (DNP): A weight loss agent with significant acute toxicity and risk of death. J Med Toxicol. 2011;7:205–12.
Perry RJ, Kim T, Zhang XM, Lee HY, Pesta D, Popov VB, et al. Reversal of hypertriglyceridemia, fatty liver disease, and insulin resistance by a liver-targeted mitochondrial uncoupler. Cell Metab. 2013;18:740–8.
Goedeke L, Peng L, Montalvo-Romeral V, Butrico GM, Dufour S, Zhang XM, et al. Controlled-release mitochondrial protonophore (CRMP) reverses dyslipidemia and hepatic steatosis in dysmetabolic nonhuman primates. Sci Transl Med. 2019;11:eaay0284.
Wei G, Song X, Fu Y, Gong T, Zhang Q. Sustained-release mitochondrial protonophore reverses nonalcoholic fatty liver disease in rats. Int J Pharm. 2017;530:230–8.
Jiang H, Jin J, Duan Y, Xie Z, Li Y, Gao A, et al. Mitochondrial uncoupling coordinated with PDH activation safely ameliorates hyperglycemia via promoting glucose oxidation. Diabetes. 2019;68:2197–209.
Jian C, Fu J, Cheng X, Shen LJ, Ji YX, Wang X, et al. Low-dose sorafenib acts as a mitochondrial uncoupler and ameliorates nonalcoholic steatohepatitis. Cell Metab. 2020;31:1206.
Alexopoulos SJ, Chen SY, Brandon AE, Salamoun JM, Byrne FL, Garcia CJ, et al. Mitochondrial uncoupler BAM15 reverses diet-induced obesity and insulin resistance in mice. Nat Commun. 2020;11:2397.
Salamoun JM, Garcia CJ, Hargett SR, Murray JH, Chen SY, Beretta M, et al. 6-Amino[1,2,5]oxadiazolo[3,4-b]pyrazin-5-ol derivatives as efficacious mitochondrial uncouplers in STAM mouse model of nonalcoholic steatohepatitis. J Med Chem. 2020;63:6203–24.
Bargaje R, Gupta S, Sarkeshik A, Park R, Xu T, Sarkar M, et al. Identification of novel targets for miR-29a using miRNA proteomics. PLoS ONE. 2012;7:e43243.
Zhang L, Zhang J, Tong Q, Wang G, Dong H, Wang Z, et al. Reduction of miR-29a-3p induced cardiac ischemia reperfusion injury in mice via targeting Bax. Exp Ther Med. 2019;18:1729–37.
Jan MI, Khan RA, Malik A, Ali T, Bilal M, Bo L, et al. Data of expression status of miR-29a and its putative target mitochondrial apoptosis regulatory gene DRP1 upon miR-15a and miR-214 inhibition. Data Brief. 2018;16:1000–4.
Yang YL, Wang PW, Wang FS, Lin HY, Huang YH. miR-29a modulates GSK3beta/SIRT1-linked mitochondrial proteostatic stress to ameliorate mouse non-alcoholic steatohepatitis. Int J Mol Sci. 2020;21:6884.
Castro RE, Ferreira DM, Afonso MB, Borralho PM, Machado MV, Cortez-Pinto H, et al. miR-34a/SIRT1/p53 is suppressed by ursodeoxycholic acid in the rat liver and activated by disease severity in human non-alcoholic fatty liver disease. J Hepatol. 2013;58:119–25.
Ding J, Li M, Wan X, Jin X, Chen S, Yu C, et al. Effect of miR-34a in regulating steatosis by targeting PPARalpha expression in nonalcoholic fatty liver disease. Sci Rep. 2015;5:13729.
Simao AL, Afonso MB, Rodrigues PM, Gama-Carvalho M, Machado MV, Cortez-Pinto H, et al. Skeletal muscle miR-34a/SIRT1:AMPK axis is activated in experimental and human non-alcoholic steatohepatitis. J Mol Med. 2019;97:1113–26.
Chen F, Feng L, Zheng YL, Lu J, Fan SH, Shan Q, et al. 2, 2’, 4, 4’-tetrabromodiphenyl ether (BDE-47) induces mitochondrial dysfunction and related liver injury via eliciting miR-34a-5p-mediated mitophagy impairment. Environ Pollut. 2020;258:113693.
Wang C, Li Q, Wang W, Guo L, Guo C, Sun Y, et al. GLP-1 contributes to increases in PGC-1alpha expression by downregulating miR-23a to reduce apoptosis. Biochem Biophys Res Commun. 2015;466:33–9.
Iliopoulos D, Drosatos K, Hiyama Y, Goldberg IJ, Zannis VI. MicroRNA-370 controls the expression of microRNA-122 and Cpt1alpha and affects lipid metabolism. J Lipid Res. 2010;51:1513–23.
Esau C, Davis S, Murray SF, Yu XX, Pandey SK, Pear M, et al. miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab. 2006;3:87–98.
Loyer X, Paradis V, Henique C, Vion AC, Colnot N, Guerin CL, et al. Liver microRNA-21 is overexpressed in non-alcoholic steatohepatitis and contributes to the disease in experimental models by inhibiting PPARalpha expression. Gut. 2016;65:1882–94.
Li K, Zhang J, Ji C, Wang L. MiR-144-3p and its target gene beta-amyloid precursor protein regulate 1-methyl-4-phenyl-1,2-3,6-tetrahydropyridine-induced mitochondrial dysfunction. Mol Cells. 2016;39:543–9.
Li R, Yan G, Li Q, Sun H, Hu Y, Sun J, et al. MicroRNA-145 protects cardiomyocytes against hydrogen peroxide H2O2-induced apoptosis through targeting the mitochondria apoptotic pathway. PLoS ONE. 2012;7:e44907.
Tomasetti M, Nocchi L, Staffolani S, Manzella N, Amati M, Goodwin J, et al. MicroRNA-126 suppresses mesothelioma malignancy by targeting IRS1 and interfering with the mitochondrial function. Antioxid Redox Signal. 2014;21:2109–25.
Trajkovski M, Hausser J, Soutschek J, Bhat B, Akin A, Zavolan M, et al. MicroRNAs 103 and 107 regulate insulin sensitivity. Nature. 2011;474:649–53.
Ogawa T, Enomoto M, Fujii H, Sekiya Y, Yoshizato K, Ikeda K, et al. MicroRNA-221/222 upregulation indicates the activation of stellate cells and the progression of liver fibrosis. Gut. 2012;61:1600–9.
Zhang Y, Huypens P, Adamson AW, Chang JS, Henagan TM, Boudreau A, et al. Alternative mRNA splicing produces a novel biologically active short isoform of PGC-1alpha. J Biol Chem. 2009;284:32813–26.
Kim J, Park MS, Ha K, Park C, Lee J, Mynatt RL, et al. NT-PGC-1alpha deficiency decreases mitochondrial FA oxidation in brown adipose tissue and alters substrate utilization in vivo. J Lipid Res. 2018;59:1660–70.
Zhu PP, Patterson A, Stadler J, Seeburg DP, Sheng M, Blackstone C. Intra- and intermolecular domain interactions of the C-terminal GTPase effector domain of the multimeric dynamin-like GTPase Drp1. J Biol Chem. 2004;279:35967–74.
Macdonald PJ, Francy CA, Stepanyants N, Lehman L, Baglio A, Mears JA, et al. Distinct splice variants of dynamin-related protein 1 differentially utilize mitochondrial fission factor as an effector of cooperative GTPase activity. J Biol Chem. 2016;291:493–507.
Cereghetti GM, Stangherlin A, Martins de Brito O, Chang CR, Blackstone C, Bernardi P, et al. Dephosphorylation by calcineurin regulates translocation of Drp1 to mitochondria. Proc Natl Acad Sci USA. 2008;105:15803–8.
Karbowski M, Neutzner A, Youle RJ. The mitochondrial E3 ubiquitin ligase MARCH5 is required for Drp1 dependent mitochondrial division. J Cell Biol. 2007;178:71–84.
Kim HK, Pham MHC, Ko KS, Rhee BD, Han J. Alternative splicing isoforms in health and disease. Pflug Arch. 2018;470:995–1016.
Dhillon S. Risdiplam: First approval. Drugs. 2020;80:1853–8.
Ratni H, Ebeling M, Baird J, Bendels S, Bylund J, Chen KS, et al. Discovery of risdiplam, a selective survival of motor neuron-2 (SMN2) gene splicing modifier for the treatment of spinal muscular atrophy (SMA). J Med Chem. 2018;61:6501–17.
Farrell G, Schattenberg JM, Leclercq I, Yeh MM, Goldin R, Teoh N, et al. Mouse models of nonalcoholic steatohepatitis: Toward optimization of their relevance to human nonalcoholic steatohepatitis. Hepatology. 2019;69:2241–57.
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (No. 92057116 and 81673493 to Jing-ya Li); the National Science and Technology Major Project (No. 2018ZX09711002-018 to Jing-ya Li); the Natural Science Foundation of Shanghai’s 2021 “Science and Technology Innovation Action Plan” (No. 21ZR1475300 to Zhi-fu Xie); the China Postdoctoral Science Foundation (No. 2021M703349 to Yu-feng Li).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Li, Yf., Xie, Zf., Song, Q. et al. Mitochondria homeostasis: Biology and involvement in hepatic steatosis to NASH. Acta Pharmacol Sin 43, 1141–1155 (2022). https://doi.org/10.1038/s41401-022-00864-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41401-022-00864-z
Keywords
This article is cited by
-
Weight regain, but not weight loss exacerbates hepatic fibrosis during multiple weight cycling events in male mice
European Journal of Nutrition (2024)
-
All about NASH: disease biology, targets, and opportunities on the road to NASH drugs
Acta Pharmacologica Sinica (2022)